Big data challenges in personalized cancer medicine Bioinformatics

Big data challenges in personalized cancer medicine Bioinformatics activities in the Norwegian Cancer Genomics Consortium (NCGC) Sigve Nakken Postdoctoral fellow, Eivind Hovigs group Norwegian Cancer Genomics Consortium (NCGC) Department of Tumor Biology, ICR, OUS cancergenomics. no

N o r w e g i a n C a n cer genomics Consortium (NCGC) radium. no/myklebost • Founded by oncologists and cancer scientists across the country (Tromsø, Trondheim, Bergen, Oslo) • Contributing to and following the national priorization of ”Individualized cancer treatment based on the gene profile of the tumour” as the most important topic in cancer research • Has obtained grants of 75 Mkr (≈ 10 MUSD) from the Research Council • Industrial partners: OCC, Pub. Gene, Bergen. Bio • Project divided into work packages ü WP 4: Data handling and establishment of national infrastructure cancergenomics. no
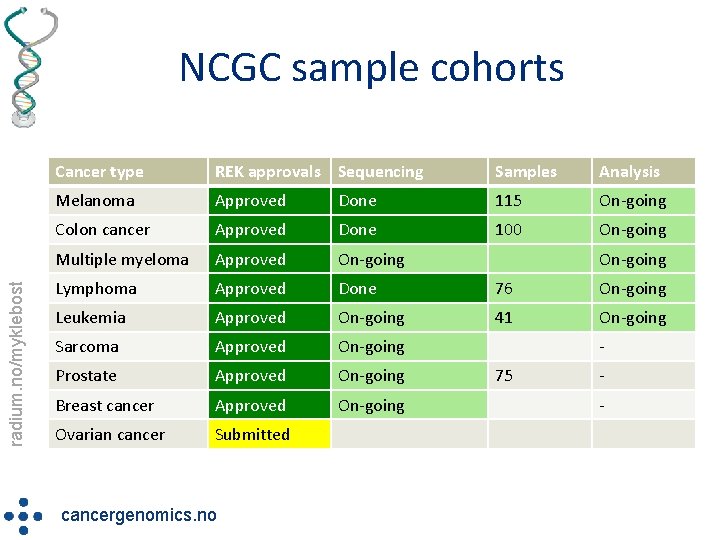
radium. no/myklebost NCGC sample cohorts Cancer type REK approvals Sequencing Samples Analysis Melanoma Approved Done 115 On-going Colon cancer Approved Done 100 On-going Multiple myeloma Approved On-going Lymphoma Approved Done 76 On-going Leukemia Approved On-going 41 On-going Sarcoma Approved On-going Prostate Approved On-going Breast cancer Approved On-going Ovarian cancer Submitted cancergenomics. no On-going 75 -

radium. no/myklebost NCGC cancer genome sequencing • Exome sequencing • Goal: identify & characterize the acquired genetic changes in the tumor sample by massively parallel deep sequencing üSNVs & Insertions/deletions üCopy number aberrations üStructural rearrangements cancergenomics. no
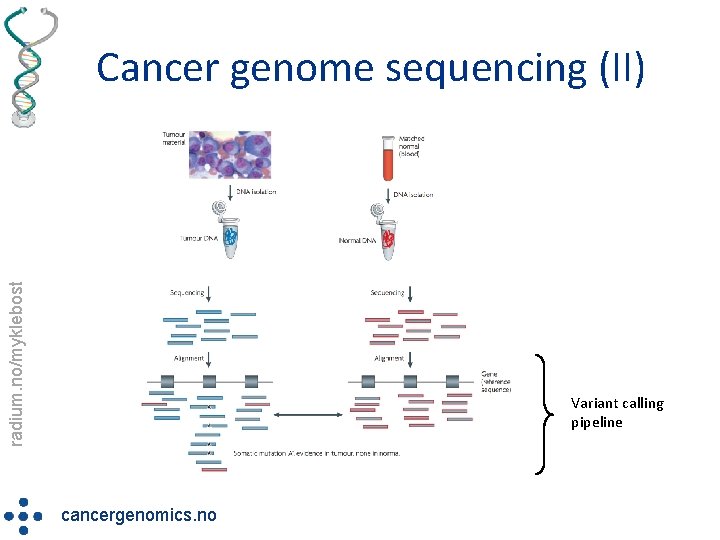
radium. no/myklebost Cancer genome sequencing (II) Variant calling pipeline cancergenomics. no

Cancer genome sequencing (III) radium. no/myklebost • How deep should I sequence my tumor sample? (to detect a mutant subpopulation at X percent? ) • Biological complexity ü Tumor purity ü Ploidy ü Local CNAs • Technical biases ü ü Uneven coverage (GC) PCR artefacts Sequencing quality/errors Oxidation (DNA extraction + library prep) • Other ü Tumor-control mismatch cancergenomics. no

Somatic variant calling radium. no/myklebost • Two key components ü Read alignment – mapping each read to its proper position in the genome ü Mutation calling – quantify the likelihood of a true somatic mutation cancergenomics. no • Best-practice workflows defined ü Still many different algorithms to choose from • Need for benchmark

ICGC mutation benchmark • Purpose: Assess concordance & accuracy of somatic SNV/indel calling among variant calling pipelines used in different research groups • Evaluate impact of different algorithms (aligner, caller etc. ) radium. no/myklebost • NCGC: optimize and verify running pipeline (“ICGC stamp”) • Participants were given raw sequence reads from a medulloblastoma (MB 99) genome (tumor + normal), ~40 X coverage ü task: submit somatic indels + snvs • Coordinated by CNAG, Barcelona (Ivo Gut’s lab) • Weekly global telephone conferences • BM 1. 2 cancergenomics. no
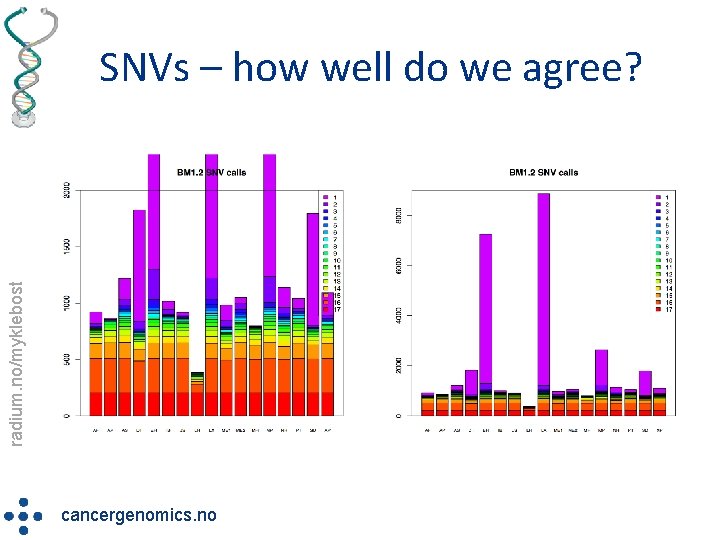
radium. no/myklebost SNVs – how well do we agree? cancergenomics. no
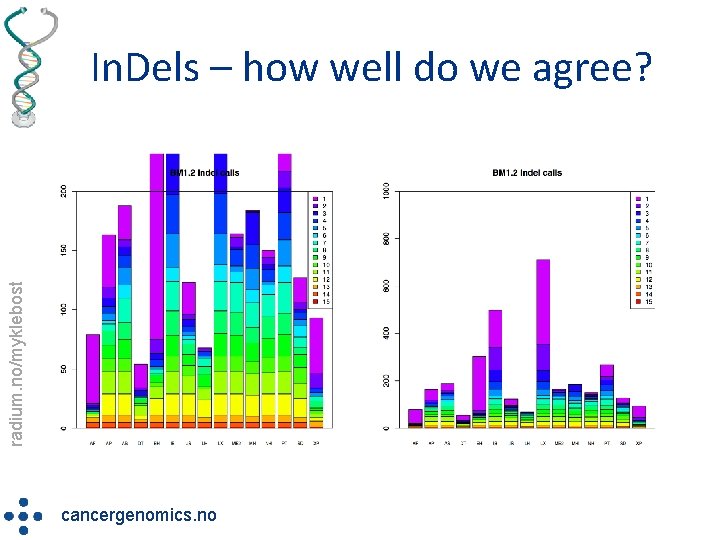
radium. no/myklebost In. Dels – how well do we agree? cancergenomics. no

Verification of calls – GOLD set 300 X sequencing of the same genome Six different pipelines called somatic SNVs and In. Dels SNVs with concordance of > 3 accepted SNVs with concordance < 3 and all indels reviewed manually radium. no/myklebost ü ü cancergenomics. no
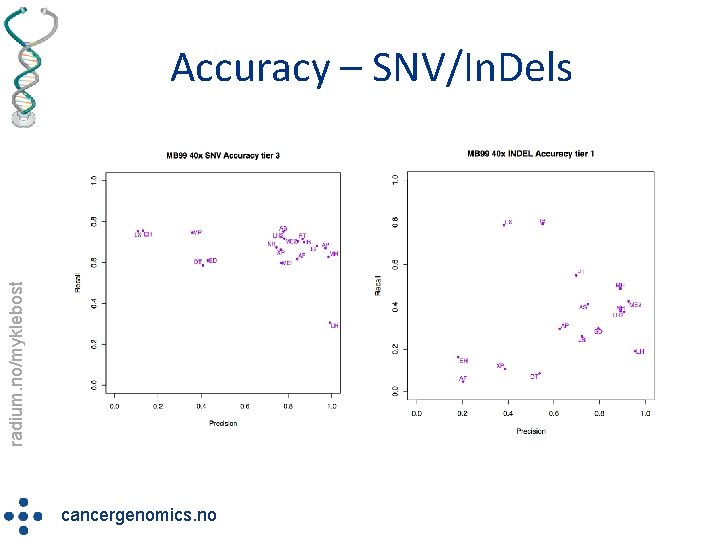
radium. no/myklebost Accuracy – SNV/In. Dels cancergenomics. no
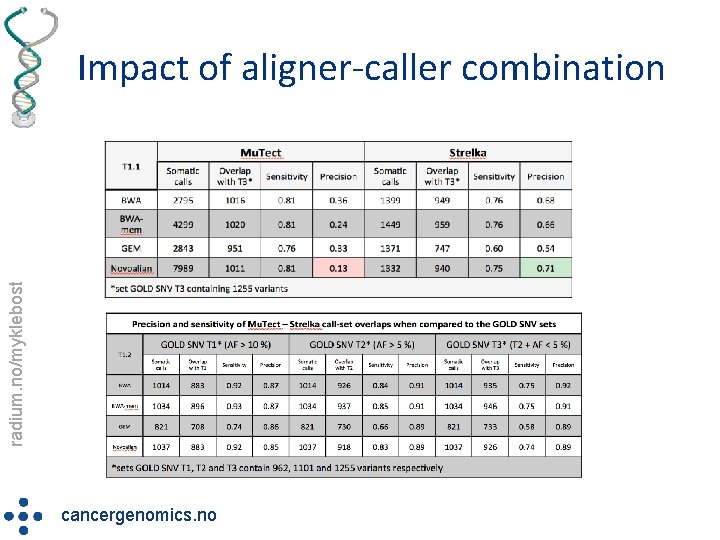
radium. no/myklebost Impact of aligner-caller combination cancergenomics. no

radium. no/myklebost Benchmark manuscript cancergenomics. no
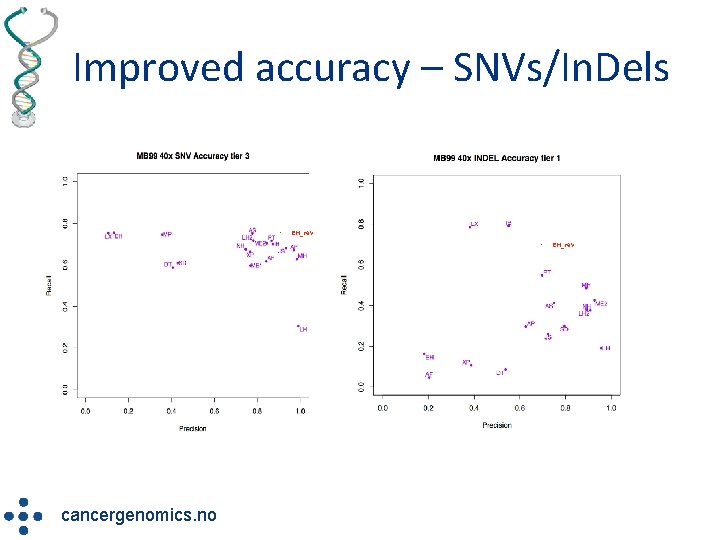
Improved accuracy – SNVs/In. Dels • EH_rev radium. no/myklebost • cancergenomics. no EH_rev

Interpretation of variants radium. no/myklebost • Which variants/genes are of functional relevance? ü Is my variant a frequent mutation? Which cancer types? ü Is my variant likely to alter the activity of the encoding protein? ü Is my variant known as a drug sensitivity marker? ü Which mutant genes are known drug targets? • Annotation pipeline cancergenomics. no Variant calling Functional annotation Prioritization

Variants – phenotypic effect? radium. no/myklebost • Computational prediction of damaging variants • Machine learning • Numerous algorithms ü SIFT, Poly. Phen 2, Mutation. Taster, Mutation. Assessor, Provean, FATHMM, etc. . • Challenge: many have been trained with Mendelian disease mutations ü Gain-of-function mutations hard to predict cancergenomics. no

Variants – clinical associations? radium. no/myklebost • Recent promising resources/data on clinically associated variants cancergenomics. no
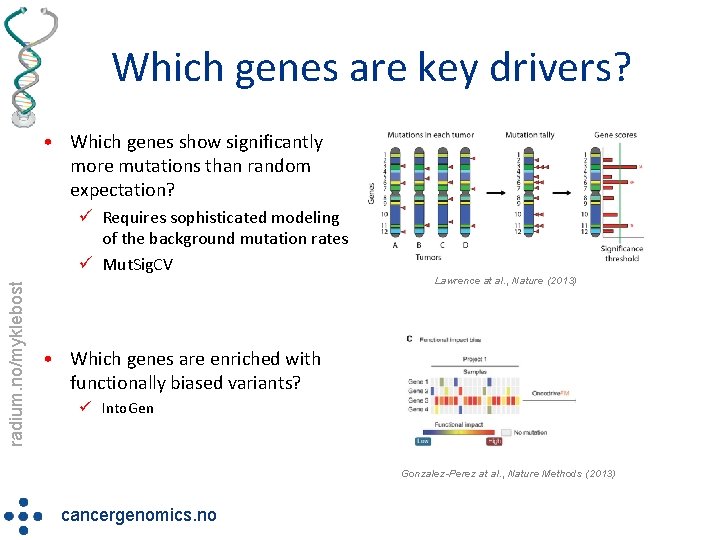
Which genes are key drivers? • Which genes show significantly more mutations than random expectation? radium. no/myklebost ü Requires sophisticated modeling of the background mutation rates ü Mut. Sig. CV Lawrence at al. , Nature (2013) • Which genes are enriched with functionally biased variants? ü Into. Gen Gonzalez-Perez at al. , Nature Methods (2013) cancergenomics. no

radium. no/myklebost NCGC – data trends cancergenomics. no
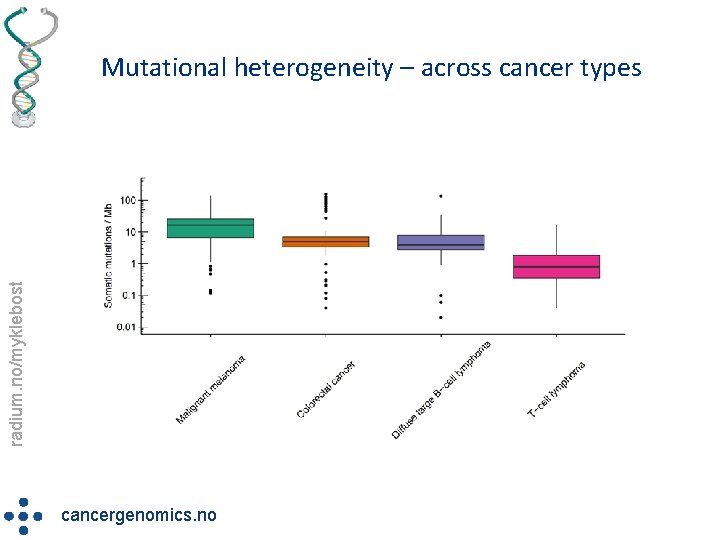
radium. no/myklebost Mutational heterogeneity – across cancer types cancergenomics. no
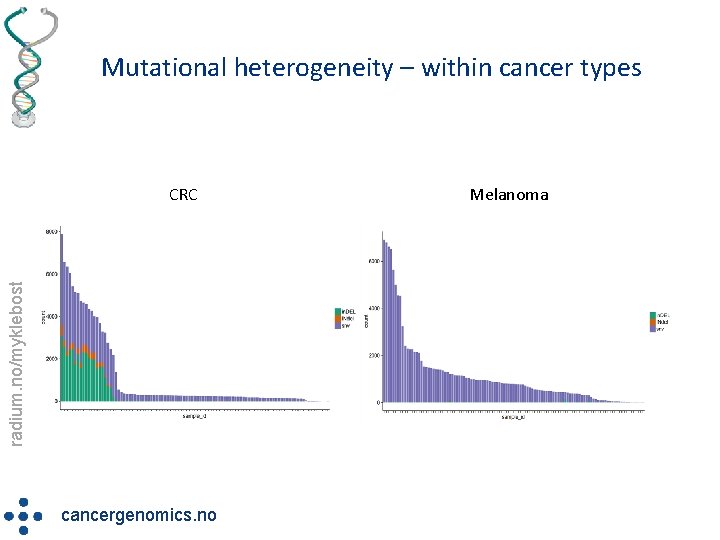
Mutational heterogeneity – within cancer types radium. no/myklebost CRC cancergenomics. no Melanoma

radium. no/myklebost Functional heterogeneity cancergenomics. no
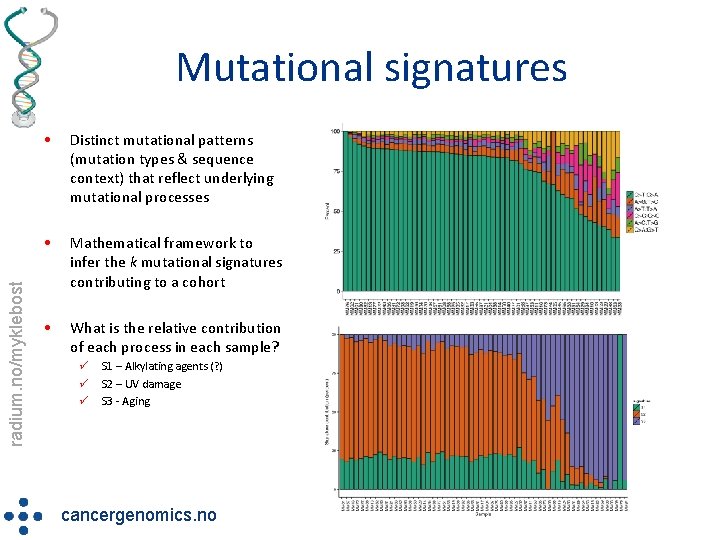
radium. no/myklebost Mutational signatures • Distinct mutational patterns (mutation types & sequence context) that reflect underlying mutational processes • Mathematical framework to infer the k mutational signatures contributing to a cohort • What is the relative contribution of each process in each sample? ü ü ü S 1 – Alkylating agents (? ) S 2 – UV damage S 3 - Aging cancergenomics. no

radium. no/myklebost In progress/future plans • Evaluation of more read aligners/variant callers • Integration of improved calling of copy number aberrations • Inference of clonal population structure • Report pr. tumor case – QC, mutated cancer genes, actionable targets etc. • Improved tools for visualization of results cancergenomics. no

radium. no/myklebost Other activities cancergenomics. no

Acknowledgements • NCGC radium. no/myklebost üPrincipal investigators üDepartment of Tumor Biology • Leonardo Meza-Zepeda, Susanne Lorenz, Ola Myklebost • Daniel Vodak, Ghislain Fournous, Lars Birger Aasheim, Eivind Hovig • ICGC Technical Validation group cancergenomics. no
- Slides: 27