BIDS Heu Di Conv f MRIprep IntroductionTutorial BIDS

BIDS, Heu. Di. Conv, f. MRIprep Introduction/Tutorial

BIDS- Brain Imaging Data Structure • Data acquired through neuroimaging experiments can be organized in many ways because of differences in scanner software, DICOM and NIFTI tools, and custom data organizing scripts within different labs. • BIDS provides a simple solution to this by introducing a standard for neuroimaging data organization. • The goal is to define a powerful, flexible, and consistent framework for integrating the diverse types of experimental data routinely acquired in neuroimaging studies. • https: //openfmri. org/data-organization/ - BIDS overview • http: //bids. neuroimaging. io/bids_spec. pdf - BIDS manual • https: //academic. oup. com/gigascience/article/5/suppl_1/3/2965208 - paper that provides some more information about BIDS and applying it to other programs

BIDS apps of interest • Heu. Di. Conv- Heuristics DICOM Converter • f. MRIprep- f. MRI pre-processing • MRIQC- MRI quality control

Heu. Di. Conv- Heuristics DICOM Converter • DICOM Digital Imaging and Communications in Medicine • A Standard for storing and transmitting medical images enabling the integration of medical imaging devices (such as scanners). • Heu. Di. Conv is a flexible DICOM converter for organizing brain imaging data into structured directory layouts. • https: //www. dicomstandard. org/ - DICOM overview • https: //github. com/nipy/heudiconv - Git. Hub Heu. Di. Conv overview

f. MRIprep- f. MRI pre-processing • f. MRIprep is a f. MRI pre-processing pipeline that is designed to provide an easily accessible, state-of-the-art interface that is robust to differences in scan acquisition protocols and that requires minimal user input, while providing easily interpretable and comprehensive error and output reporting. • Allows us to take f. MRI data from raw to fully preprocessed form. • https: //github. com/poldracklab/fmriprep - Git. Hub f. MRIprep overview • http: //fmriprep. readthedocs. io/en/latest/ - f. MRIprep website overview

Before running f. MRIprep: • https: //github. com/poldracklab/fmriprep/blob/master/docs/installati on. rst - list of instructions to install f. MRIprep on your computer. • Note that we will be running f. MRIprep in a Docker Container through the Command Prompt on our lab desktop computers. • https: //osf. io/29 j 8 m/wiki/home/ - additional lab notes if you run into issues installing anything from the first link.

2 ways to run f. MRIprep • For an individual participant: • Faster • More simple • Good if you want to quickly check a portion of data • For a group of participants: • More thorough • Better if all of your data is ready to be processed

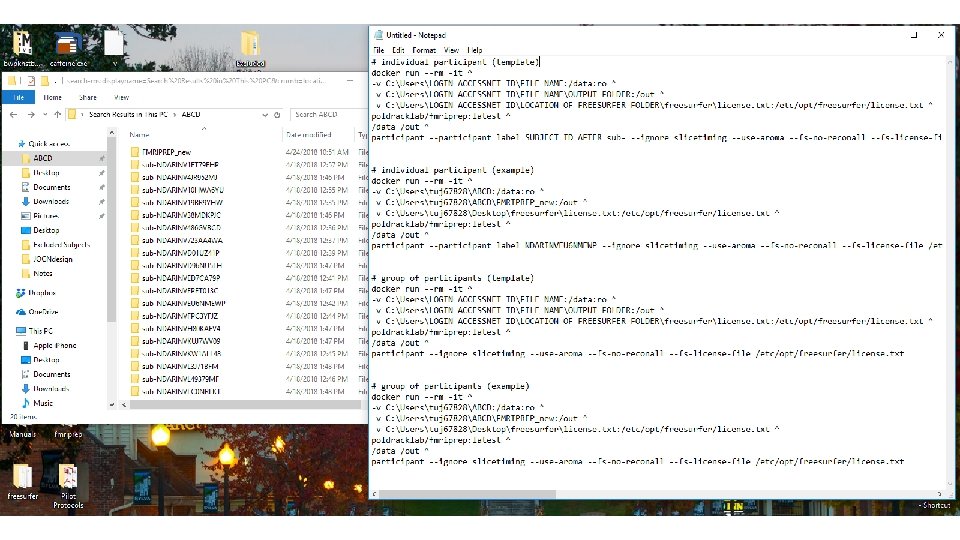
f. MRIprep for an individual participant:

f. MRIprep for a group of participants:

MRIQC- MRI Quality Control • Extracts no-reference image quality metrics from structural (T 1 w and T 2 w) and functional MRI data. • Essential for excluding problematic acquisitions and avoiding bias in subsequent image processing and analysis. • https: //github. com/poldracklab/mriqc - Git. Hub MRIQC overview • https: //mriqc. readthedocs. io/en/latest/ - MRIQC website overview • https: //www. ncbi. nlm. nih. gov/pmc/articles/PMC 5612458/ - paper that provides more information about MRIQC and why it is useful • https: //mriqc. readthedocs. io/en/latest/running. html - documentation to help you run MRIQC

Before running MRIQC: • https: //mriqc. readthedocs. io/en/latest/install. html - list of instructions to install MRIQC to your computer. • Note that we will be running MRIQC in a Docker Container through the Command Prompt on our lab desktop computers. • https: //osf. io/29 j 8 m/wiki/home/ - additional lab notes if you run into issues installing anything from the first link.

MRIQC code
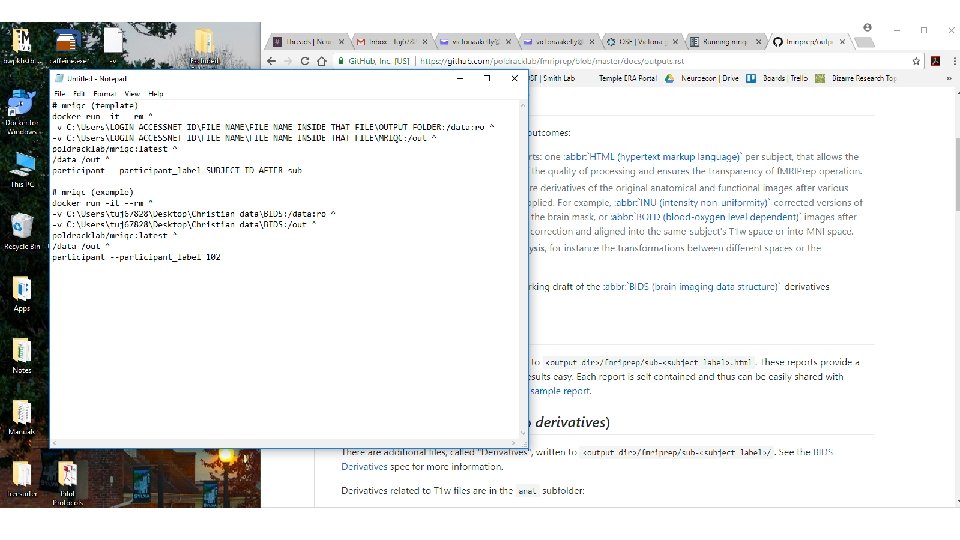

What do we do once we’ve converted the data?
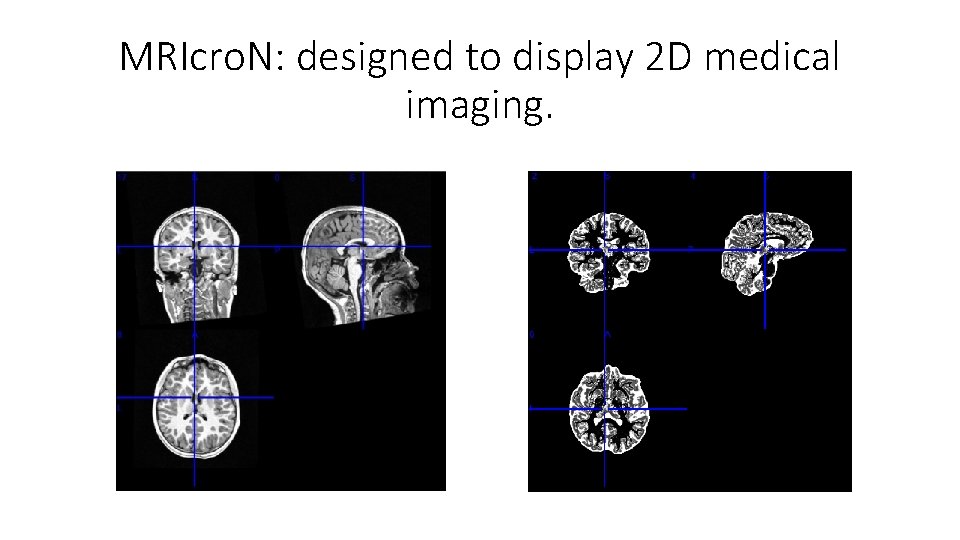
MRIcro. N: designed to display 2 D medical imaging.
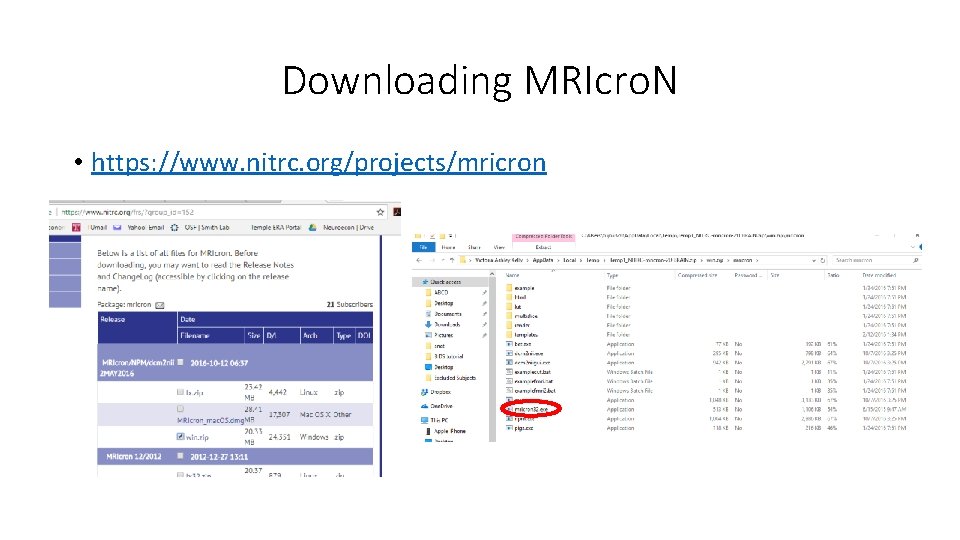
Downloading MRIcro. N • https: //www. nitrc. org/projects/mricron
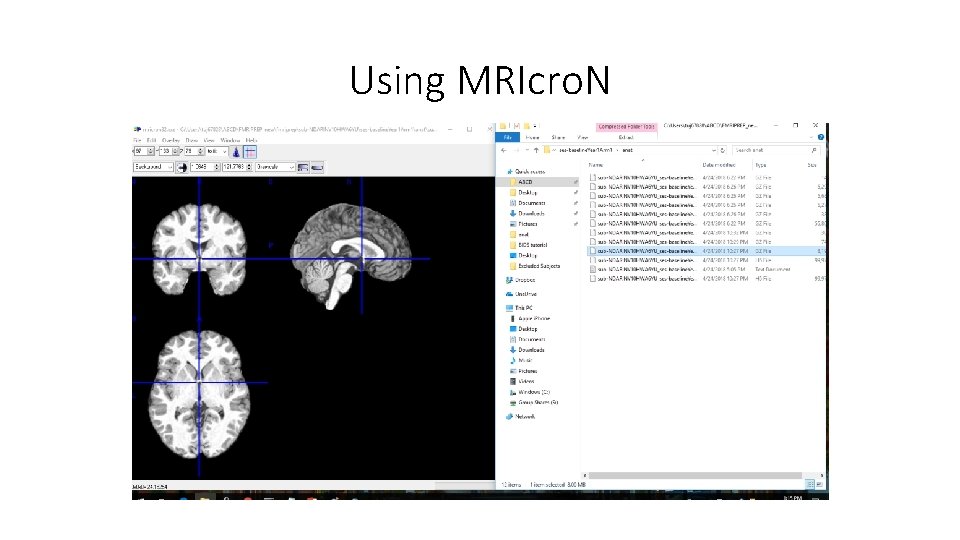
Using MRIcro. N
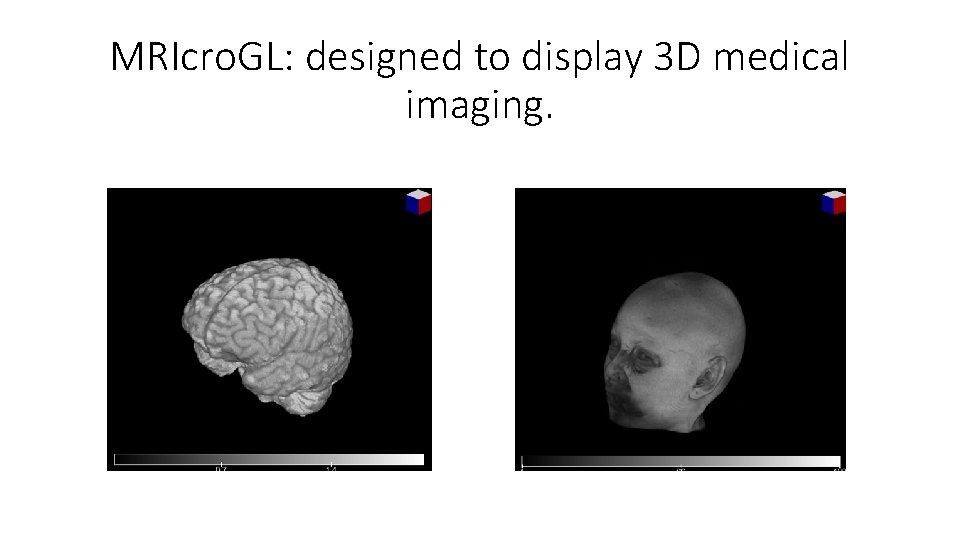
MRIcro. GL: designed to display 3 D medical imaging.
Downloading MRIcro. GL • https: //www. nitrc. org/frs/? group_id=889
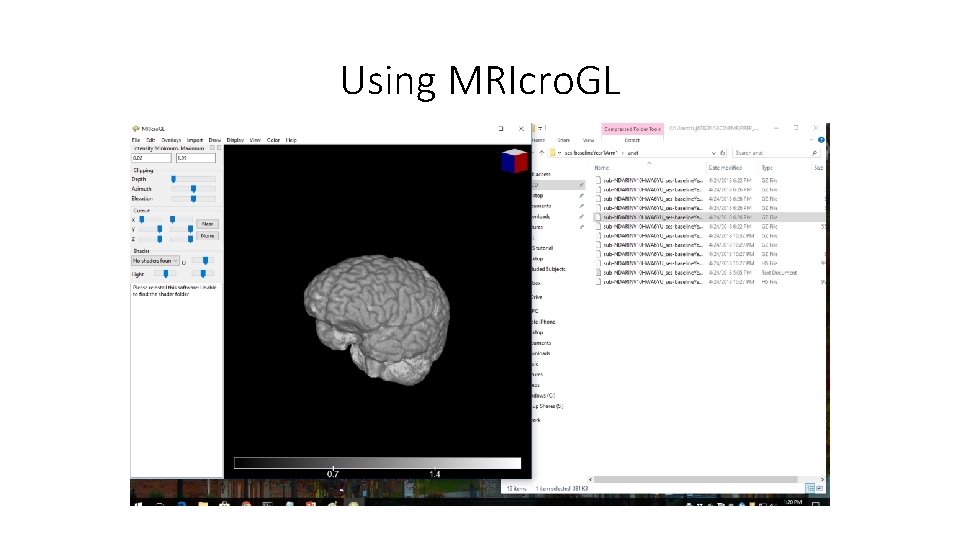
Using MRIcro. GL
- Slides: 21