BiClustering Data Mining Clustering Kmeans clustering minimizes Where

Bi-Clustering
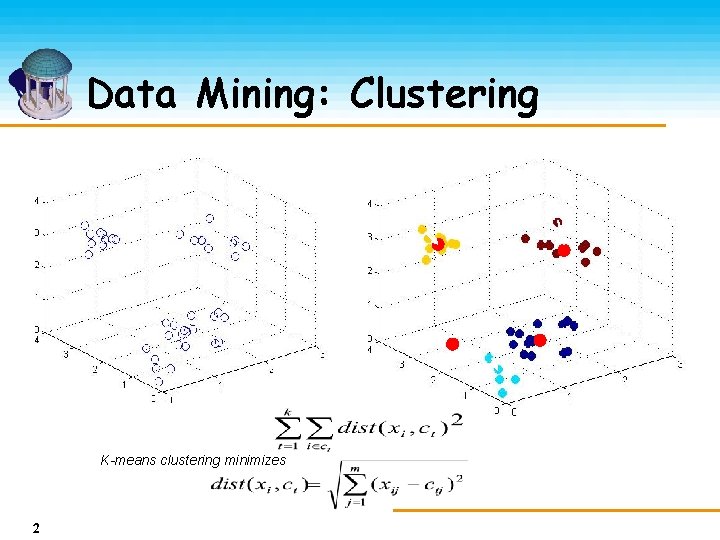
Data Mining: Clustering K-means clustering minimizes Where 2
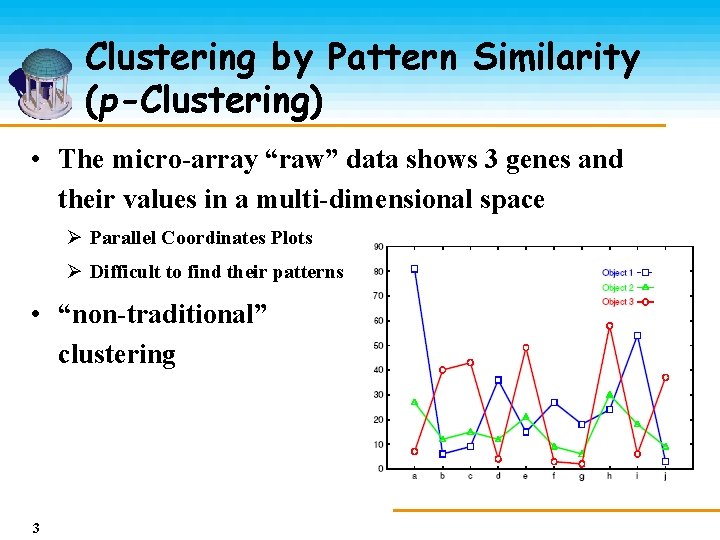
Clustering by Pattern Similarity (p-Clustering) • The micro-array “raw” data shows 3 genes and their values in a multi-dimensional space Ø Parallel Coordinates Plots Ø Difficult to find their patterns • “non-traditional” clustering 3
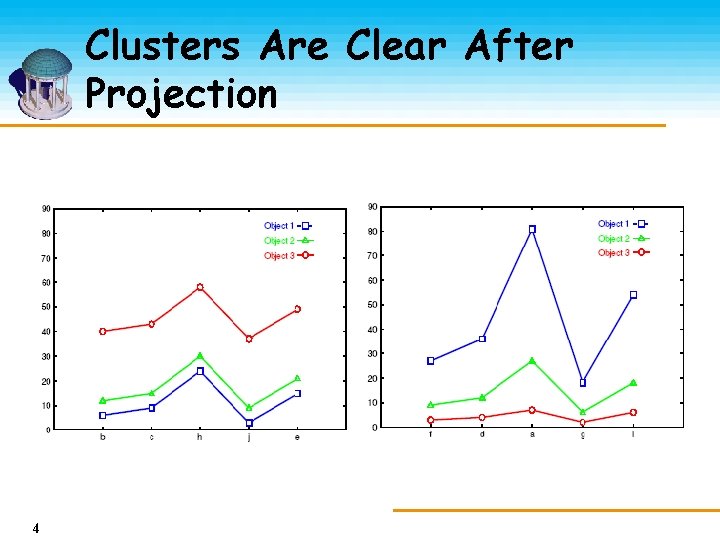
Clusters Are Clear After Projection 4
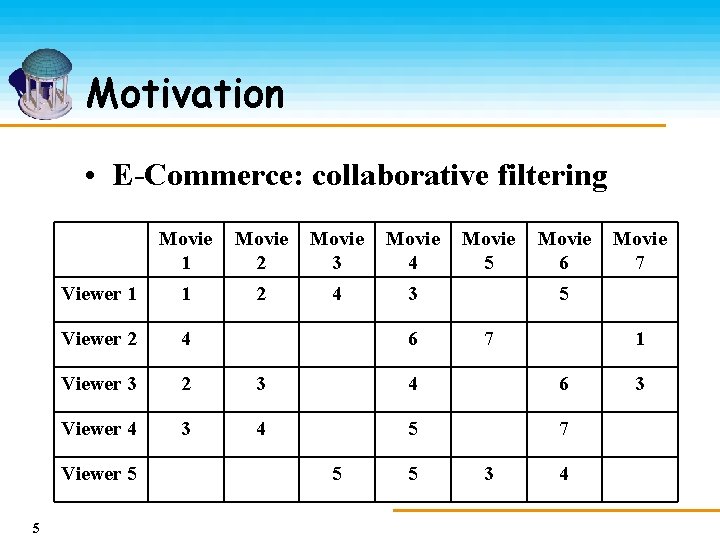
Motivation • E-Commerce: collaborative filtering Movie 1 Movie 2 Movie 3 Movie 4 Viewer 1 1 2 4 3 Viewer 2 4 Viewer 3 2 3 4 6 Viewer 4 3 4 5 7 Viewer 5 5 6 5 5 Movie 6 Movie 7 5 7 3 1 4 3
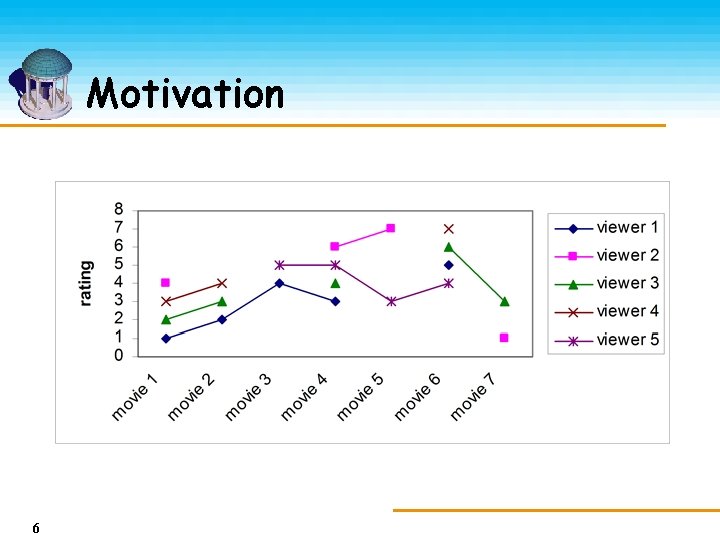
Motivation 6
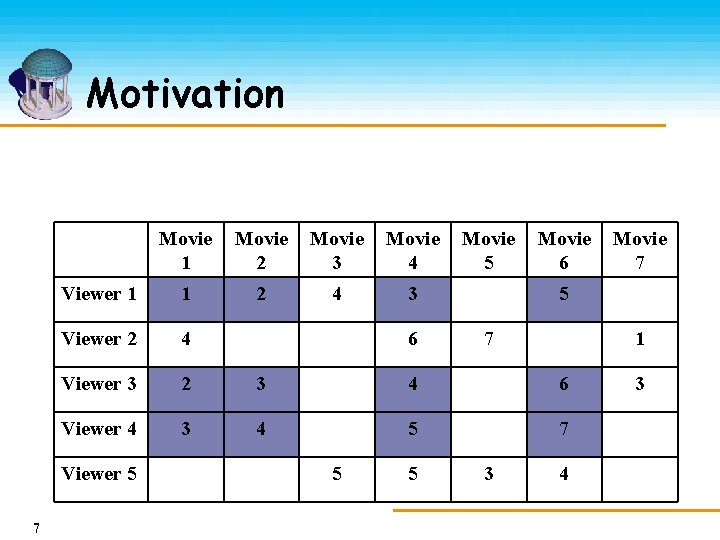
Motivation Movie 1 Movie 2 Movie 3 Movie 4 Viewer 1 1 2 4 3 Viewer 2 4 Viewer 3 2 3 4 6 Viewer 4 3 4 5 7 Viewer 5 7 6 5 5 Movie 6 Movie 7 5 7 3 1 4 3
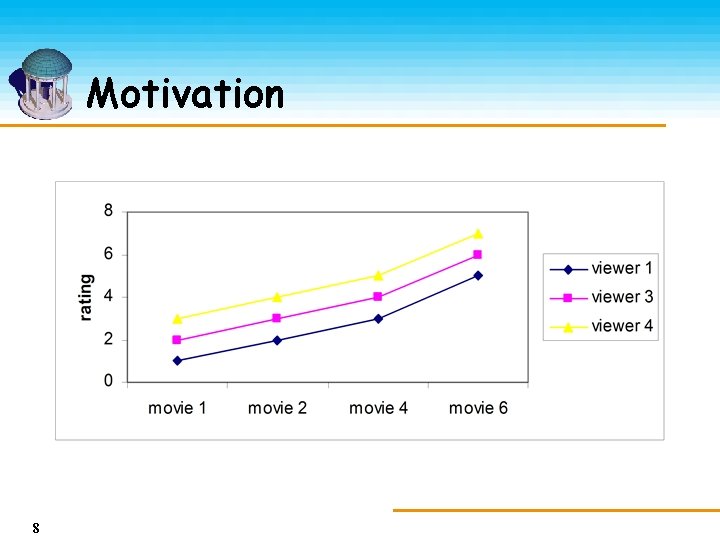
Motivation 8
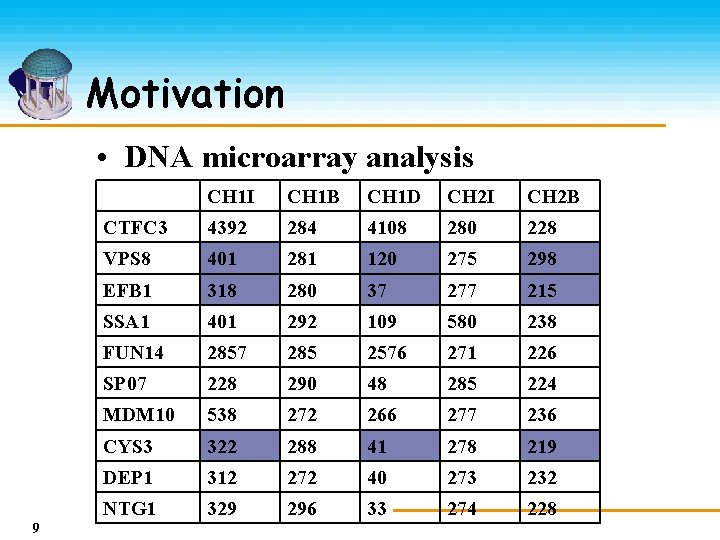
Motivation • DNA microarray analysis 9 CH 1 I CH 1 B CH 1 D CH 2 I CH 2 B CTFC 3 4392 284 4108 280 228 VPS 8 401 281 120 275 298 EFB 1 318 280 37 277 215 SSA 1 401 292 109 580 238 FUN 14 2857 285 2576 271 226 SP 07 228 290 48 285 224 MDM 10 538 272 266 277 236 CYS 3 322 288 41 278 219 DEP 1 312 272 40 273 232 NTG 1 329 296 33 274 228
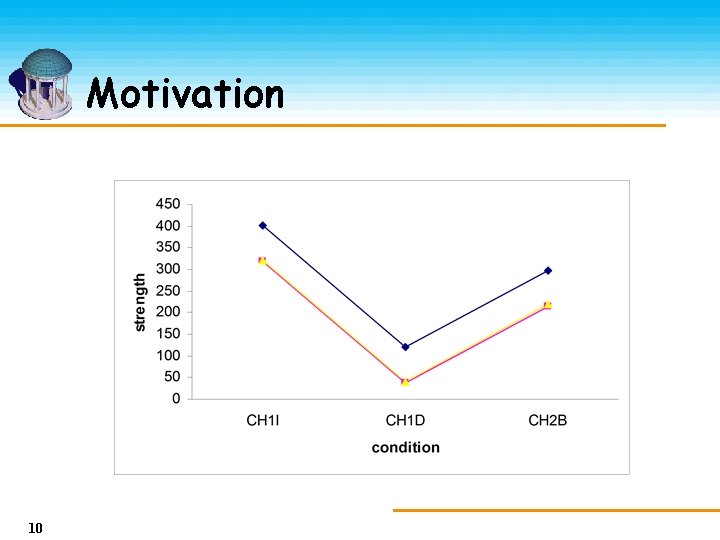
Motivation 10

Motivation • Strong coherence exhibits by the selected objects on the selected attributes. Ø They are not necessarily close to each other but rather bear a constant shift. Ø Object/attribute bias • bi-cluster 11

Challenges • The set of objects and the set of attributes are usually unknown. • Different objects/attributes may possess different biases and such biases Ø may be local to the set of selected objects/attributes Ø are usually unknown in advance • May have many unspecified entries 12

Previous Work • Subspace clustering Ø Identifying a set of objects and a set of attributes such that the set of objects are physically close to each other on the subspace formed by the set of attributes. • Collaborative filtering: Pearson R Ø Only considers global offset of each object/attribute. 13

bi-cluster • Consists of a (sub)set of objects and a (sub)set of attributes Ø Corresponds to a submatrix Ø Occupancy threshold v. Each object/attribute has to be filled by a certain percentage. Ø Volume: number of specified entries in the submatrix Ø Base: average value of each object/attribute (in the bi-cluster) 14

bi-cluster CH 1 I CH 1 B CH 1 D CH 2 I CH 2 B Obj base CTFC 3 VPS 8 401 120 298 273 EFB 1 318 37 215 190 322 41 219 194 347 66 244 219 SSA 1 FUN 14 SP 07 MDM 10 CYS 3 DEP 1 NTG 1 Attr base 15
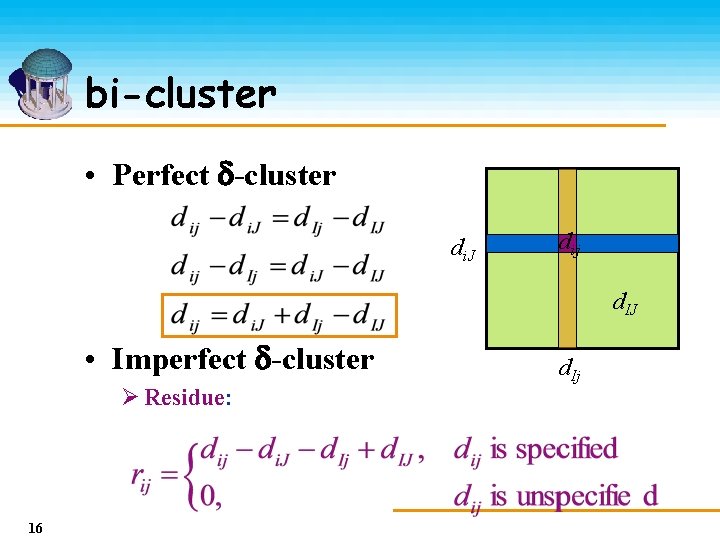
bi-cluster • Perfect -cluster di. J dij d. IJ • Imperfect -cluster Ø Residue: 16 d. Ij

bi-cluster • The smaller the average residue, the stronger the coherence. • Objective: identify -clusters with residue smaller than a given threshold 17

Cheng-Church Algorithm • Find one bi-cluster. • Replace the data in the first bi-cluster with random data • Find the second bi-cluster, and go on. • The quality of the bi-cluster degrades (smaller volume, higher residue) due to the insertion of random data. 18
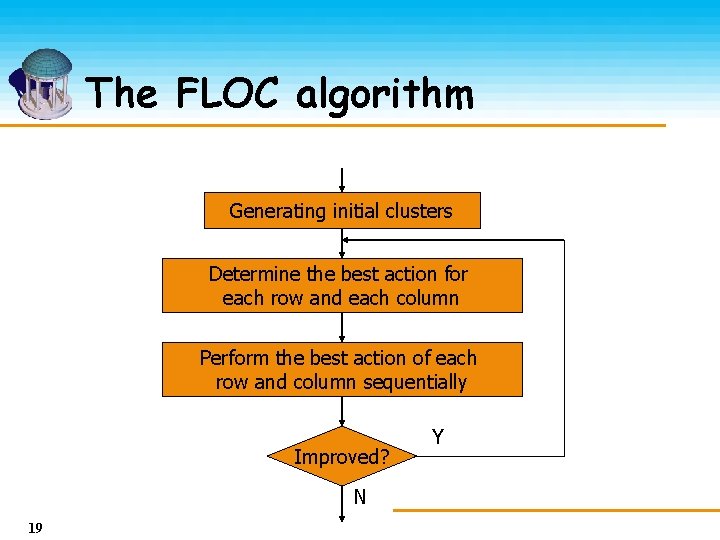
The FLOC algorithm Generating initial clusters Determine the best action for each row and each column Perform the best action of each row and column sequentially Improved? N 19 Y
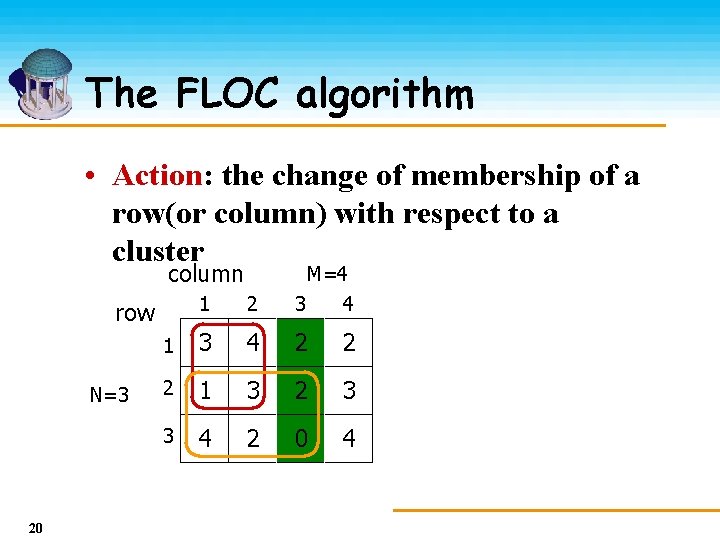
The FLOC algorithm • Action: the change of membership of a row(or column) with respect to a cluster column 1 2 M=4 3 4 1 3 4 2 2 2 1 3 2 3 3 4 2 0 4 row N=3 20 M+N actions are Performed at each iteration

The FLOC algorithm • Gain of an action: the residue reduction incurred by performing the action • Order of action: Ø Fixed order Ø Random order Ø Weighted random order • Complexity: O((M+N)MNkp) 21

The FLOC algorithm • Additional features Ø Maximum allowed overlap among clusters Ø Minimum coverage of clusters Ø Minimum volume of each cluster • Can be enforced by “temporarily blocking” certain action during the mining process if such action would violate some constraint. 22

Performance • Microarray data: 2884 genes, 17 conditions Ø 100 bi-clusters with smallest residue were returned. Ø Average residue = 10. 34 v. The average residue of clusters found via the state of the art method in computational biology field is 12. 54 Ø The average volume is 25% bigger Ø The response time is an order of magnitude faster 23

Conclusion Remark • The model of bi-cluster is proposed to capture coherent objects with incomplete data set. Ø base Ø residue • Many additional features can be accommodated (nearly for free). 24
- Slides: 24