BEGR 424Bio 324 Molecular Biology William Terzaghi Spring

BEGR 424/Bio 324 Molecular Biology William Terzaghi Spring, 2013

BEGR 424/BIO 324 - Resource and Policy Information Instructor: Dr. William Terzaghi Office: SLC 363 Office hours: MWF 10: 00 -12: 00, or by appointment Phone: (570) 408 -4762 Email: terzaghi@wilkes. edu

BEGR 424/BIO 324 - Resource and Policy Information Instructor: Dr. William Terzaghi Office: SLC 363 Office hours: MWF 10: 00 -12: 00, or by appointment Phone: (570) 408 -4762 Email: terzaghi@wilkes. edu Course webpage: http: //staffweb. wilkes. edu/william. terzaghi/BIO 324. html

General considerations What do you hope to learn?

General considerations What do you hope to learn? Graduate courses 1. learning about current literature

General considerations What do you hope to learn? Graduate courses 1. learning about current literature • Learning how to give presentations

General considerations What do you hope to learn? Graduate courses 1. learning about current literature 2. Learning current techniques

General considerations What do you hope to learn? Graduate courses 1. learning about current literature 2. Learning current techniques • Using them!

Plan A • Provide a genuine experience in using cell and molecular biology to learn about a fundamental problem in biology. • Rather than following a set series of lectures, study a problem and see where it leads us. • Lectures & presentations will relate to current status • Some class time will be spent in lab & vice-versa • we may need to come in at other times as well

Plan A 1. Pick a problem 2. Design some experiments

Plan A 1. Pick a problem 2. Design some experiments 3. See where they lead us

Plan A 1. Pick a problem 2. Design some experiments 3. See where they lead us Grading? Combination of papers and presentations

Plan A Grading? Combination of papers and presentations • First presentation: 10 points • Research presentation: 10 points • Final presentation: 15 points • Assignments: 5 points each • Poster: 10 points • Intermediate report 10 points • Final report: 30 points

1. 2. 3. 4. 5. 6. Plan A Topics? Bypassing Calvin cycle Making vectors for Dr. Harms Making vectors for Dr. Lucent Cloning & sequencing antisense RNA Studying nc. RNA Something else?

Plan A Assignments? 1. identify a gene and design primers 2. presentation on new sequencing tech 3. designing a protocol to verify your clone 4. presentations on gene regulation 5. presentation on applying mol bio Other work 1. draft of report on cloning & sequencing 2. poster for symposium 3. final gene report 4. draft of formal report 5. formal report

Plan B Standard lecture course, except: 1. Last lectures will be chosen by you -> electives

Plan B Standard lecture course, except: 1. Last lectures will be chosen by you -> electives 2. Last 4 labs will be an independent research project

Plan B Standard lecture course, except: 1. Last lectures will be chosen by you -> electives 2. Last 4 labs will be an independent research project 3. 20% of grade will be “elective” • Paper • Talk • Research proposal • Poster • Exam

Date JAN 14 16 18 21 23 25 28 30 FEB 1 4 6 8 11 13 15 18 Plan B schedule- Spring 2013 TOPIC General Introduction Genome organization Cloning & libraries: why and how DNA fingerprinting DNA sequencing Genome projects Studying proteins Meiosis & recombination Recombination Cell cycle Mitosis Exam 1 DNA replication Transcription 1 Transcription 2 Transcription 3

20 22 25 27 MAR 1 4 6 8 11 13 15 18 20 22 25 27 29 Apr 1 m. RNA processing Post-transcriptional regulation Protein degradation Epigenetics Small RNA Spring Recess RNomics Proteomics Exam 2 Protein synthesis 1 Protein synthesis 2 Membrane structure/Protein targeting 1 Protein targeting 2 Organelle genomes Easter

APR 3 5 8 10 12 15 17 19 22 24 26 29 May 1 ? ? ? Mitochondrial genomes and RNA editing Nuclear: cytoplasmic genome interactions Elective Elective Elective Exam 3 Elective Last Class! Final examination

Lab Schedule Date Jan TOPIC 16 DNA extraction and analysis 23 BLAST, etc, primer design 30 PCR Feb 6 RNA extraction and analysis 13 RT-PCR 20 q. RT-PCR 27 cloning PCR fragments Mar 6 Spring Recess 13 DNA sequencing 20 Induced gene expression 27 Northern analysis Apr 3 Independent project 10 Independent project 17 Independent project 24 Independent project
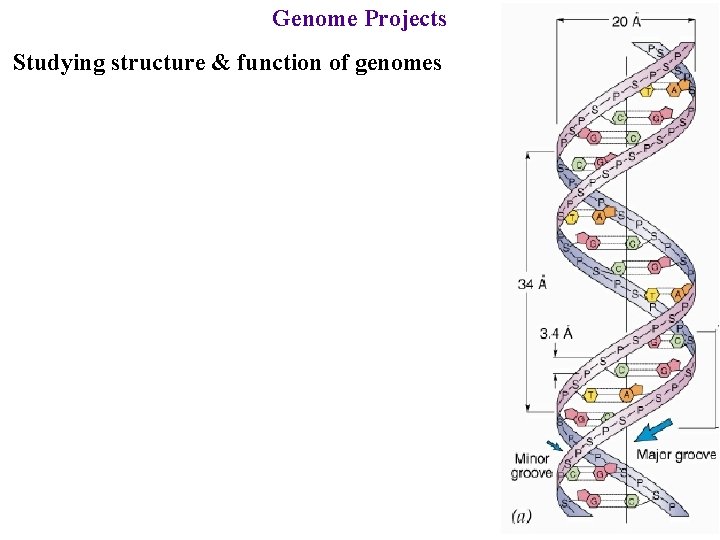
Genome Projects Studying structure & function of genomes
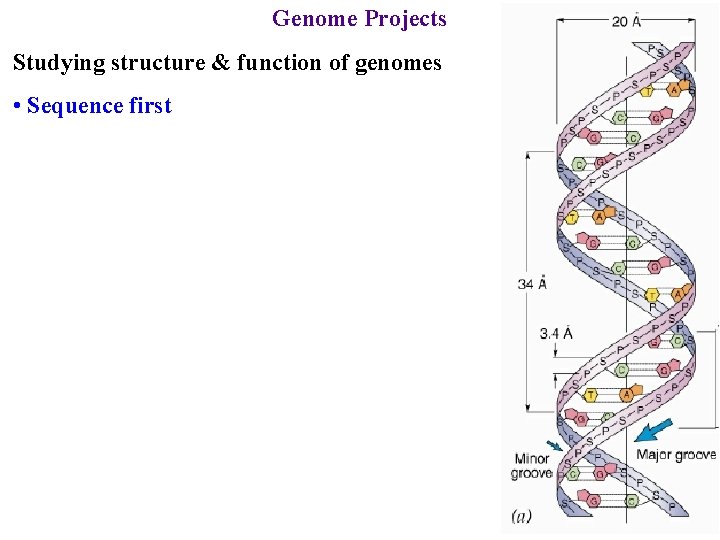
Genome Projects Studying structure & function of genomes • Sequence first
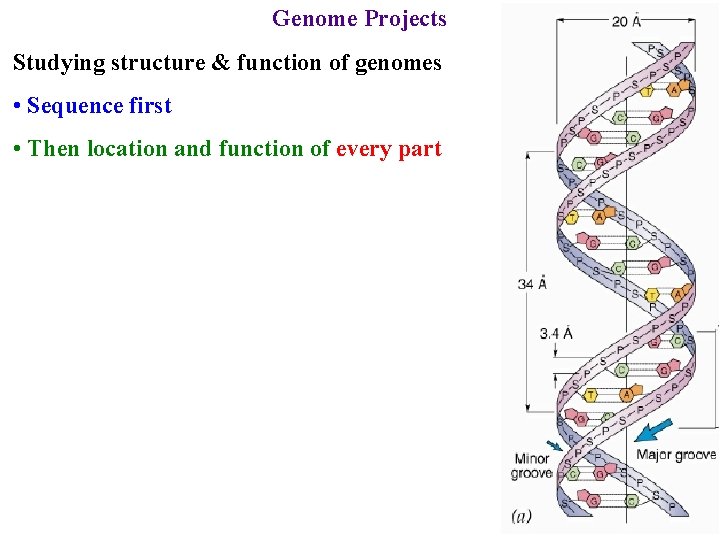
Genome Projects Studying structure & function of genomes • Sequence first • Then location and function of every part

Genome Projects How much DNA is there? SV 40 has 5000 base pairs E. coli has 5 x 106 Yeast has 2 x 107 Arabidopsis has 108 Rice has 5 x 108 Humans have 3 x 109 Soybeans have 3 x 109 Toads have 3 x 109 Salamanders have 8 x 1010 Lilies have 1011

Genome Projects C-value paradox: DNA content/haploid genome varies widely

Genome Projects C-value paradox: DNA content/haploid genome varies widely Some phyla show little variation: birds all have ~109 bp

Genome Projects C-value paradox: DNA content/haploid genome varies widely Some phyla show little variation: birds all have ~109 bp mammals all have ~ 3 x 109 bp

Genome Projects C-value paradox: DNA content/haploid genome varies widely Some phyla show little variation: birds all have ~109 bp mammals all have ~ 3 x 109 bp Other phyla are all over: insects and amphibians vary 100 x

Genome Projects C-value paradox: DNA content/haploid genome varies widely Some phyla show little variation: birds all have ~109 bp mammals all have ~ 3 x 109 bp Other phyla are all over: insects and amphibians vary 100 x flowering plants vary 1000 x

C-value paradox One cause = variations in chromosome numbers and ploidy 2 C chromosome numbers vary widely Haplopappus has 2

C-value paradox One cause = variations in chromosome numbers and ploidy 2 C chromosome numbers vary widely Haplopappus has 2 Arabidopsis has 10

C-value paradox One cause = variations in chromosome numbers and ploidy 2 C chromosome numbers vary widely Haplopappus has 2 Arabidopsis has 10 Rice has 24 Humans have 46 Tobacco (hexaploid) has 72 Kiwifruit (octaploid) have 196

C-value paradox Chromosome numbers vary So does chromosome size! Reason = variation in amounts of repetitive DNA

C-value paradox Chromosome numbers vary So does chromosome size! Reason = variation in amounts of repetitive DNA first demonstrated using Cot curves

Cot curves • denature (melt) DNA by heating
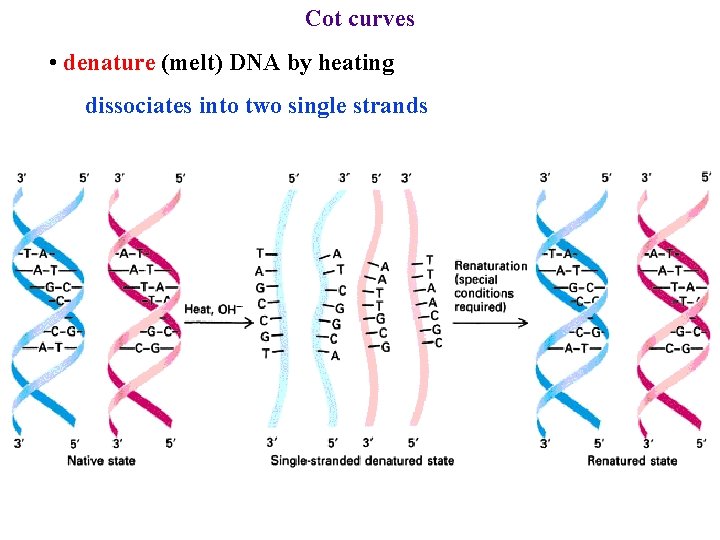
Cot curves • denature (melt) DNA by heating dissociates into two single strands
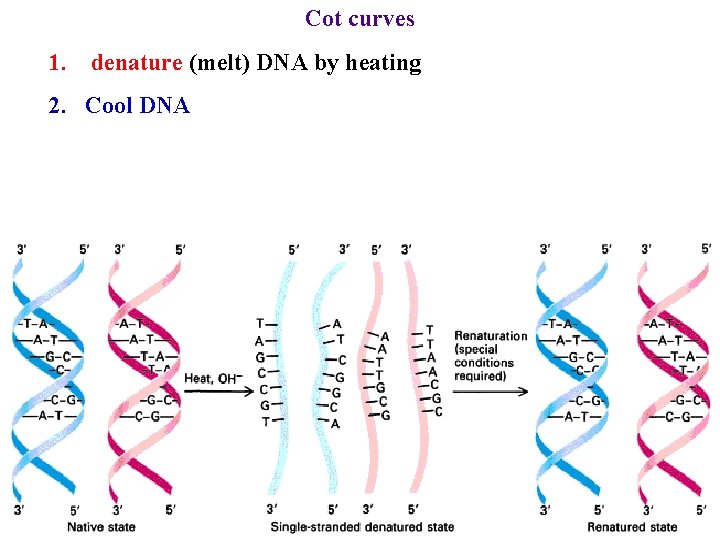
Cot curves 1. denature (melt) DNA by heating 2. Cool DNA
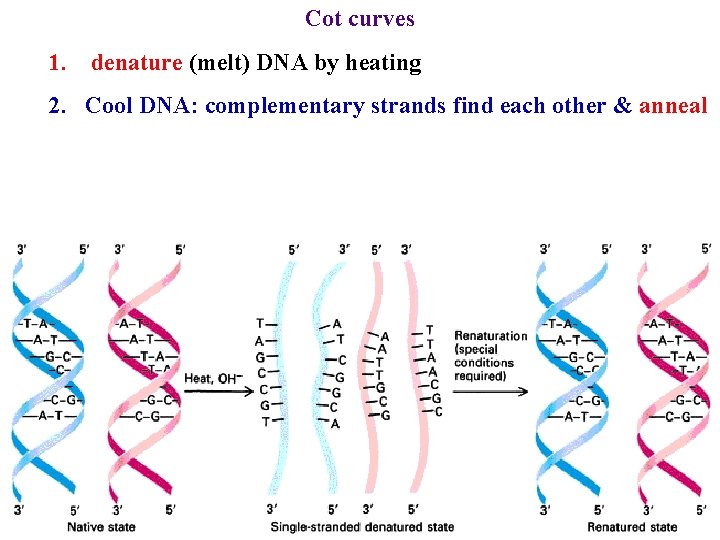
Cot curves 1. denature (melt) DNA by heating 2. Cool DNA: complementary strands find each other & anneal
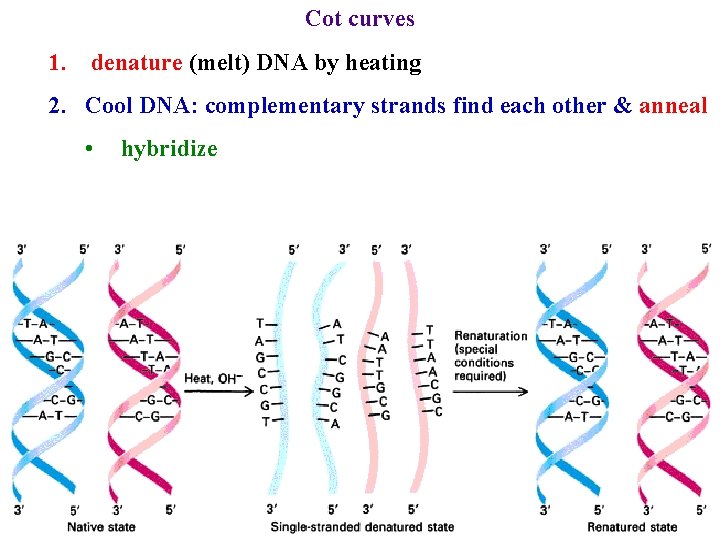
Cot curves 1. denature (melt) DNA by heating 2. Cool DNA: complementary strands find each other & anneal • hybridize
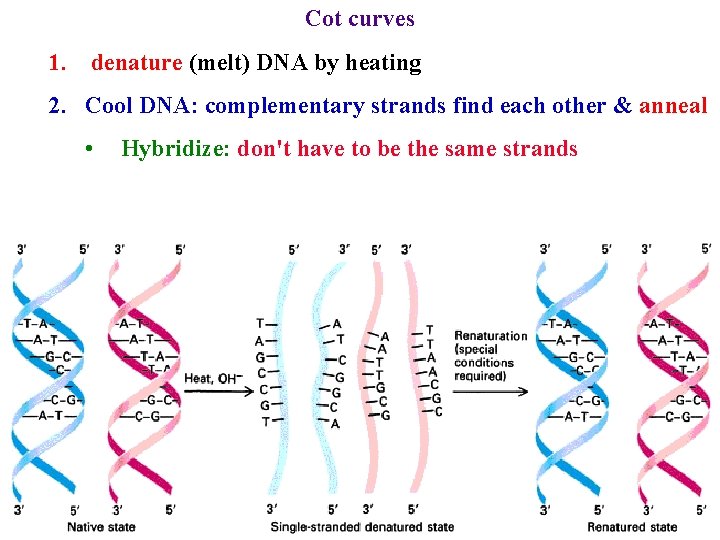
Cot curves 1. denature (melt) DNA by heating 2. Cool DNA: complementary strands find each other & anneal • Hybridize: don't have to be the same strands

Cot curves 1. denature (melt) DNA by heating 2. Cool DNA: complementary strands find each other & anneal • Hybridize: don't have to be the same strands 3. Rate depends on [complementary strands]
![Cot curves 1) denature DNA 2) cool DNA 3) at intervals measure [single-stranded DNA] Cot curves 1) denature DNA 2) cool DNA 3) at intervals measure [single-stranded DNA]](http://slidetodoc.com/presentation_image/42348b43ee52a803b52b874a28a47543/image-44.jpg)
Cot curves 1) denature DNA 2) cool DNA 3) at intervals measure [single-stranded DNA]
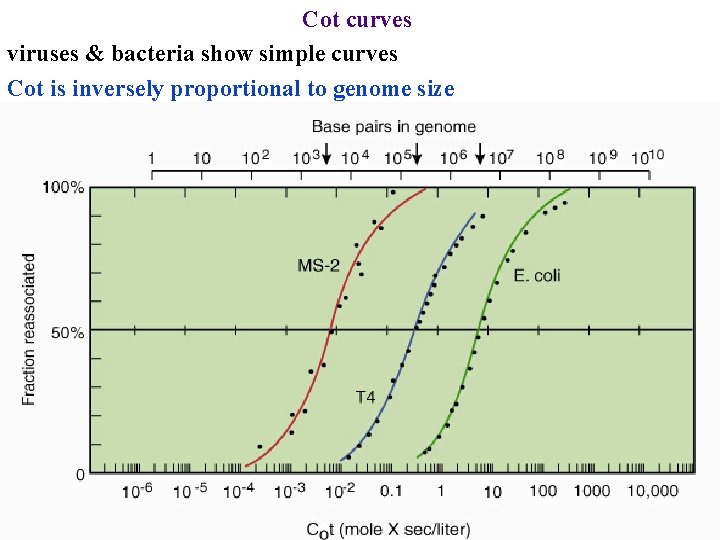
Cot curves viruses & bacteria show simple curves Cot is inversely proportional to genome size
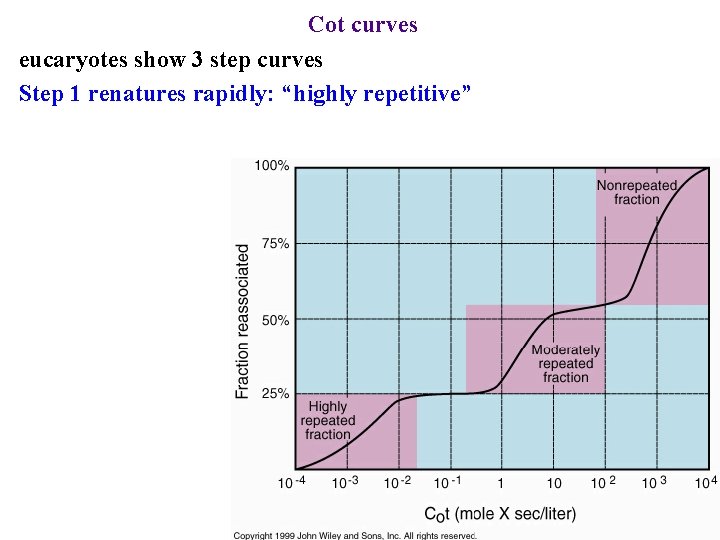
Cot curves eucaryotes show 3 step curves Step 1 renatures rapidly: “highly repetitive”
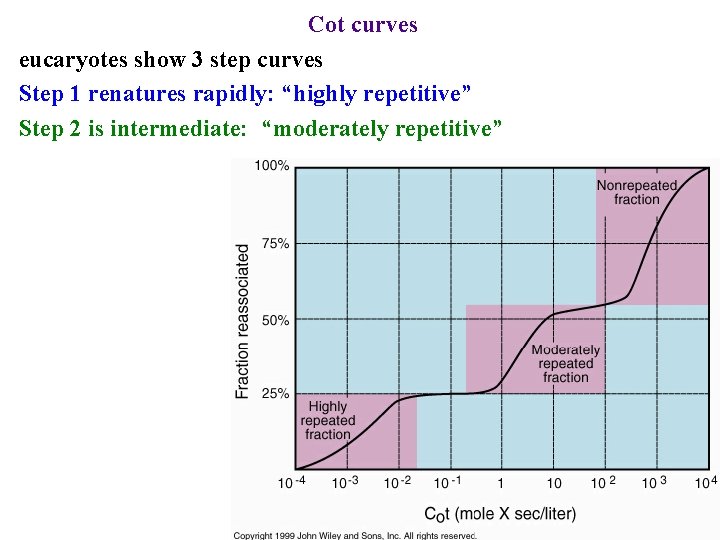
Cot curves eucaryotes show 3 step curves Step 1 renatures rapidly: “highly repetitive” Step 2 is intermediate: “moderately repetitive”
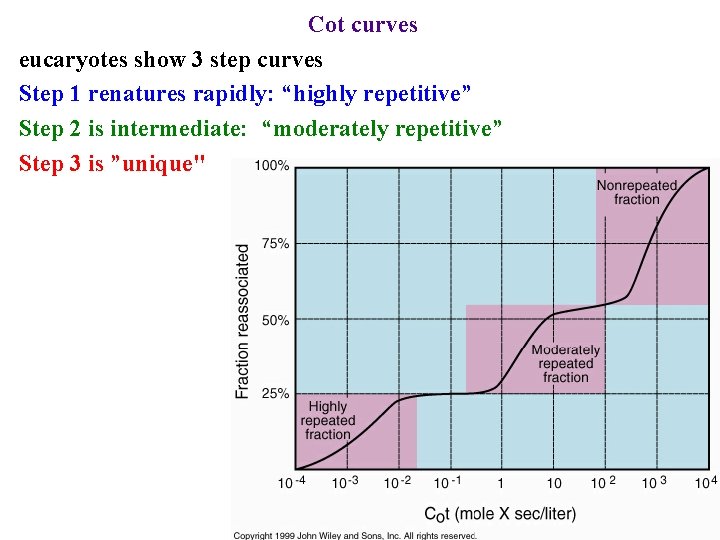
Cot curves eucaryotes show 3 step curves Step 1 renatures rapidly: “highly repetitive” Step 2 is intermediate: “moderately repetitive” Step 3 is ”unique"

Molecular cloning To identify the types of DNA sequences found within each class they must be cloned

Molecular cloning To identify the types of DNA sequences found within each class they must be cloned Force host to make millions of copies of a specific sequence

Molecular cloning To identify the types of DNA sequences found within each class they must be cloned Why? To obtain enough copies of a specific sequence to work with! typical genes are 1, 000 bp cf haploid human genome is 3, 000, 000 bp average gene is < 1/1, 000 of total genome
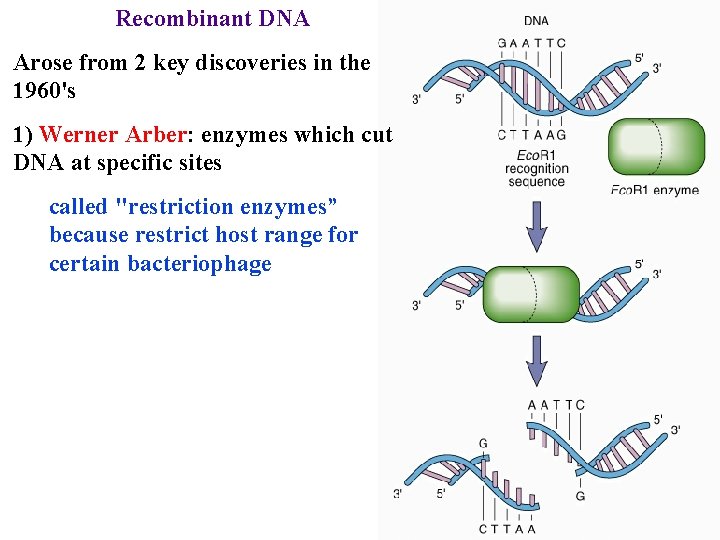
Recombinant DNA Arose from 2 key discoveries in the 1960's 1) Werner Arber: enzymes which cut DNA at specific sites called "restriction enzymes” because restrict host range for certain bacteriophage
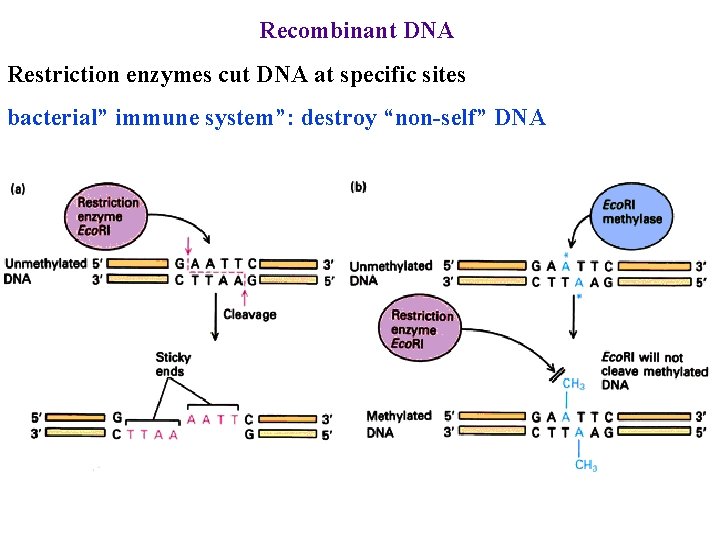
Recombinant DNA Restriction enzymes cut DNA at specific sites bacterial” immune system”: destroy “non-self” DNA
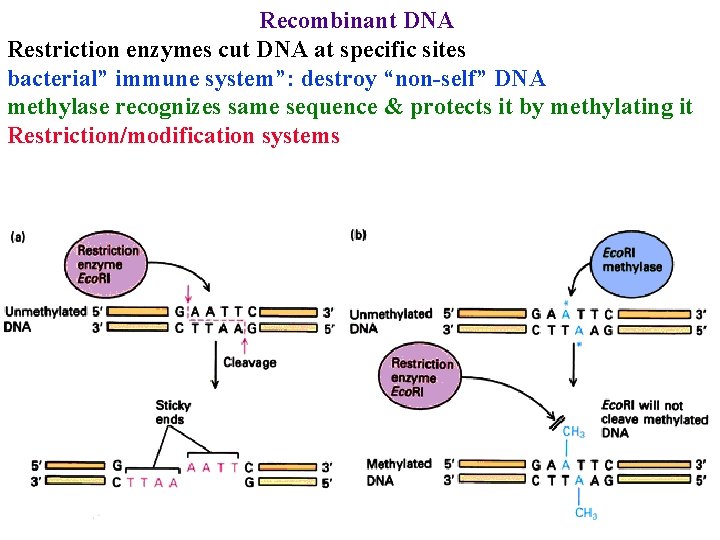
Recombinant DNA Restriction enzymes cut DNA at specific sites bacterial” immune system”: destroy “non-self” DNA methylase recognizes same sequence & protects it by methylating it Restriction/modification systems
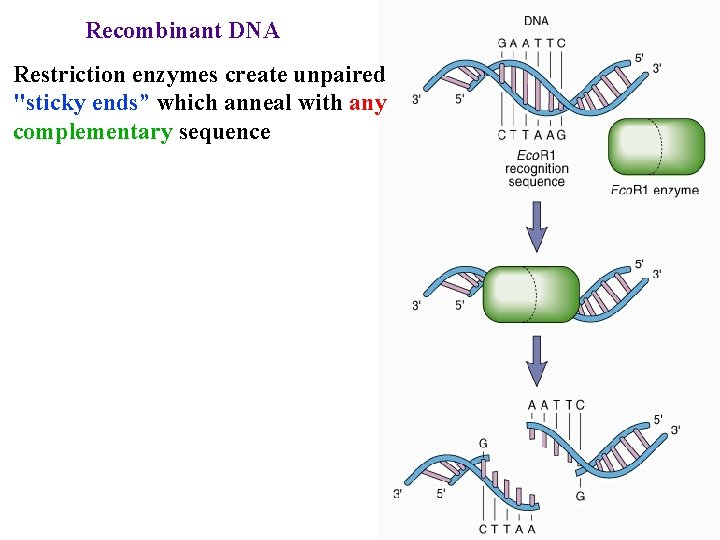
Recombinant DNA Restriction enzymes create unpaired "sticky ends” which anneal with any complementary sequence

Recombinant DNA Arose from 2 key discoveries in the 1960's 1) restriction enzymes 2) Weiss: DNA ligase -> enzyme which glues DNA strands together seals "nicks" in DNA backbone
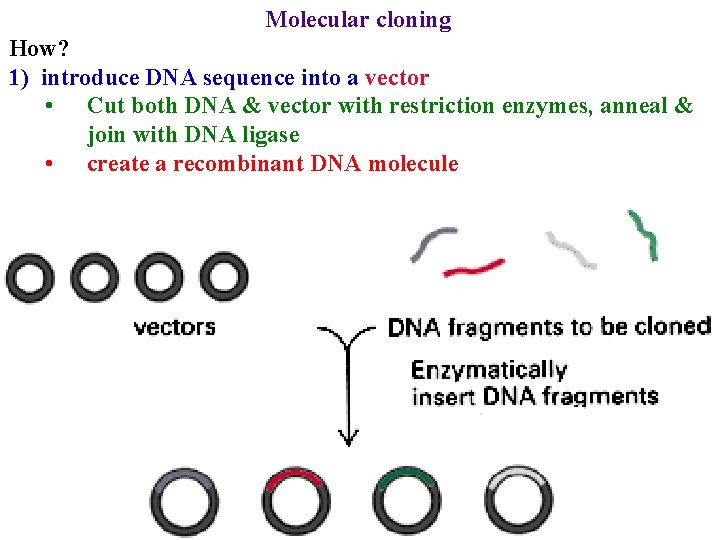
Molecular cloning How? 1) introduce DNA sequence into a vector • Cut both DNA & vector with restriction enzymes, anneal & join with DNA ligase • create a recombinant DNA molecule

Molecular cloning How? 1) create recombinant DNA 2) transform recombinant molecules into suitable host
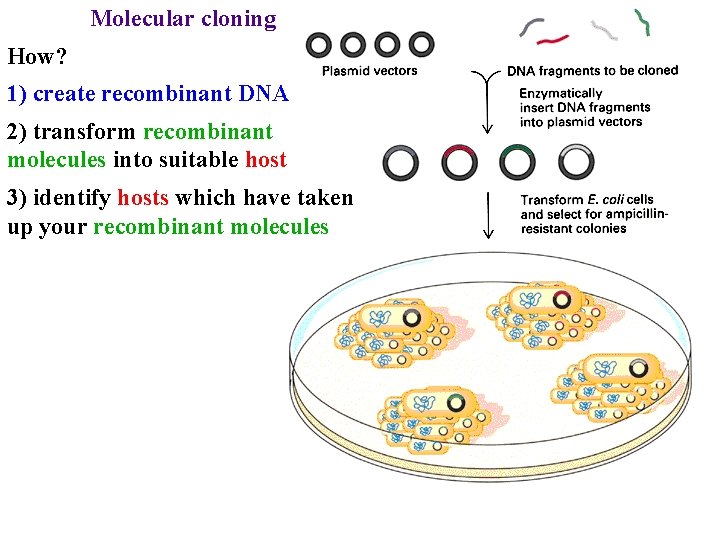
Molecular cloning How? 1) create recombinant DNA 2) transform recombinant molecules into suitable host 3) identify hosts which have taken up your recombinant molecules
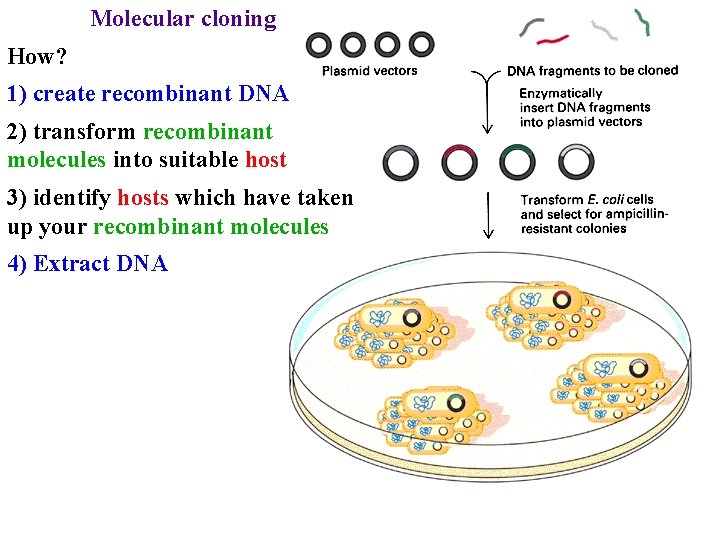
Molecular cloning How? 1) create recombinant DNA 2) transform recombinant molecules into suitable host 3) identify hosts which have taken up your recombinant molecules 4) Extract DNA
- Slides: 60