Bee Space Informatics Research Cheng Xiang Cheng Zhai

Bee. Space Informatics Research Cheng. Xiang (“Cheng”) Zhai Department of Computer Science Institute for Genomic Biology Statistics Graduate School of Library & Information Science University of Illinois at Urbana-Champaign Bee. Space Workshop, May 22, 2009
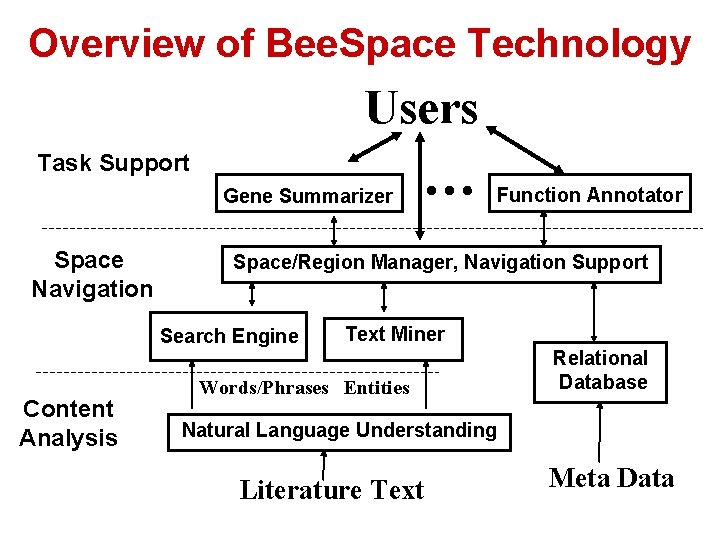
Overview of Bee. Space Technology Users Task Support Gene Summarizer Space Navigation Function Annotator Space/Region Manager, Navigation Support Search Engine Content Analysis … Text Miner Words/Phrases Entities Relational Database Natural Language Understanding Literature Text Meta Data

Part 1: Content Analysis
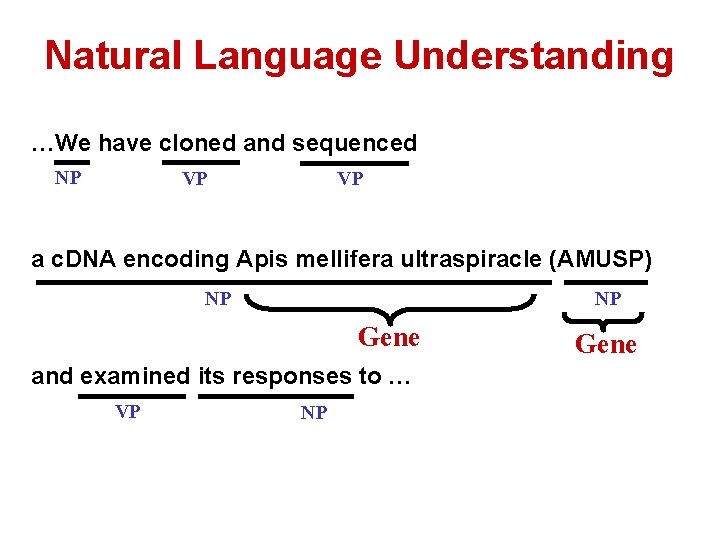
Natural Language Understanding …We have cloned and sequenced NP VP VP a c. DNA encoding Apis mellifera ultraspiracle (AMUSP) NP NP Gene and examined its responses to … VP NP Gene

Sample Technique 1: Automatic Gene Recognition • Syntactic clues: – Capitalization (especially acronyms) – Numbers (gene families) – Punctuation: -, /, : , etc. • Contextual clues: – Local: surrounding words such as “gene”, “encoding”, “regulation”, “expressed”, etc. – Global: same noun phrase occurs several times in the same article

Maximum Entropy Model for Gene Tagging • • Given an observation (a token or a noun phrase), together with its context, denoted as x Predict y {gene, non-gene} Maximum entropy model: P(y|x) = K exp( ifi(x, y)) Typical f: – y = gene & candidate phrase starts with a capital letter – y = gene & candidate phrase contains digits • Estimate i with training data

Domain overfitting problem • • When a learning based gene tagger is applied to a domain different from the training domain(s), the performance tends to decrease significantly. The same problem occurs in other types of text, e. g. , named entities in news articles. Training domain Test domain F 1 mouse 0. 541 fly mouse 0. 281 Reuters 0. 908 Reuters WSJ 0. 643

Observation I • Overemphasis on domain-specific features in the trained model wingless daughterless eyeless apexless … fly “suffix –less” weighted high in the model trained from fly data

Observation II • Generalizable features: generalize well in all domains – …decapentaplegic and wingless are expressed in analogous patterns in each primordium of… (fly) – …that CD 38 is expressed by both neurons and glial cells…that PABPC 5 is expressed in fetal brain and in a range of adult tissues. (mouse)

Observation II • Generalizable features: generalize well in all domains – …decapentaplegic and wingless are expressed in analogous patterns in each primordium of… (fly) – …that CD 38 is expressed by both neurons and glial cells…that PABPC 5 is expressed in fetal brain and in a range of adult tissues. (mouse) “wi+2 = expressed” is generalizable
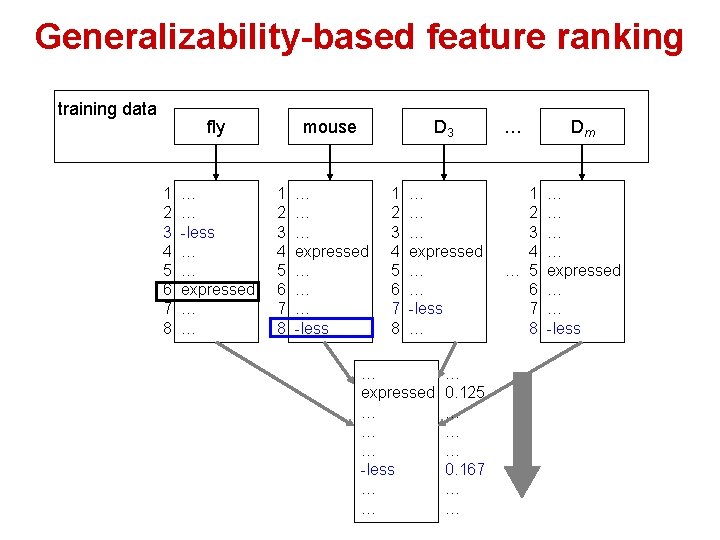
Generalizability-based feature ranking training data fly 1 2 3 4 5 6 7 8 … … -less … … expressed … … mouse 1 2 3 4 5 6 7 8 D 3 … … … expressed … … … -less 1 2 3 4 5 6 7 8 … … … expressed … … -less … … expressed … … … -less … … … 0. 125 … … … 0. 167 … … … 1 2 3 4 … 5 6 7 8 Dm … … expressed … … -less
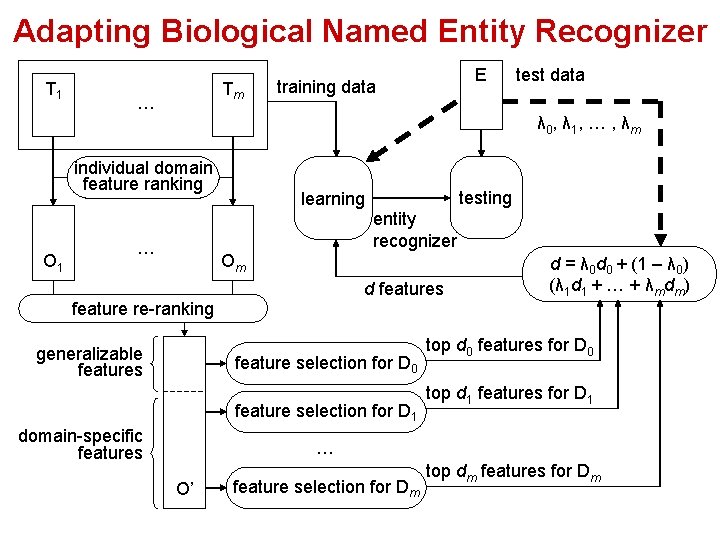
Adapting Biological Named Entity Recognizer T 1 Tm … E training data test data λ 0, λ 1, … , λm individual domain feature ranking O 1 testing learning entity recognizer … Om d features d = λ 0 d 0 + (1 – λ 0) (λ 1 d 1 + … + λmdm) feature re-ranking generalizable features feature selection for D 0 feature selection for D 1 domain-specific features top d 0 features for D 0 top d 1 features for D 1 … O’ feature selection for Dm top dm features for Dm
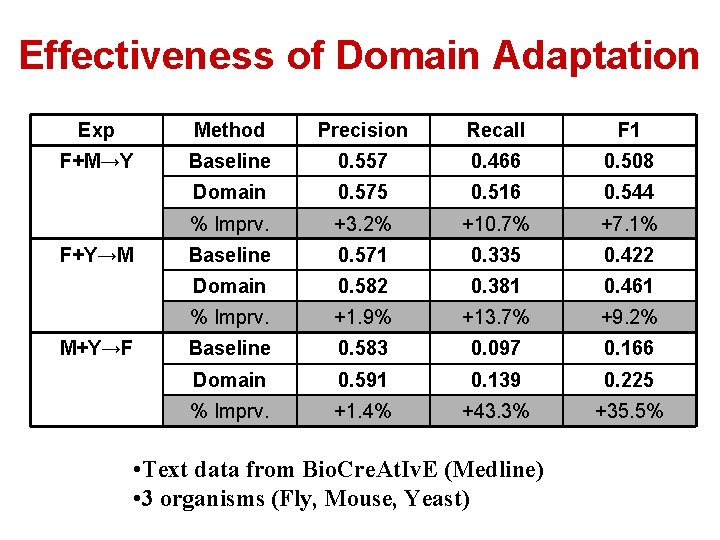
Effectiveness of Domain Adaptation Exp Method Precision Recall F 1 F+M→Y Baseline 0. 557 0. 466 0. 508 Domain 0. 575 0. 516 0. 544 % Imprv. +3. 2% +10. 7% +7. 1% Baseline 0. 571 0. 335 0. 422 Domain 0. 582 0. 381 0. 461 % Imprv. +1. 9% +13. 7% +9. 2% Baseline 0. 583 0. 097 0. 166 Domain 0. 591 0. 139 0. 225 % Imprv. +1. 4% +43. 3% +35. 5% F+Y→M M+Y→F • Text data from Bio. Cre. At. Iv. E (Medline) • 3 organisms (Fly, Mouse, Yeast)

Gene Recognition in V 3 • A variation of the basic maximum entropy – Classes: {Begin, Inside, Outside} – Features: syntactical features, POS tags, class labels of previous two tokens – Post-processing to exploit global features • Leverage existing toolkit: BMR

Part 2: Navigation Support
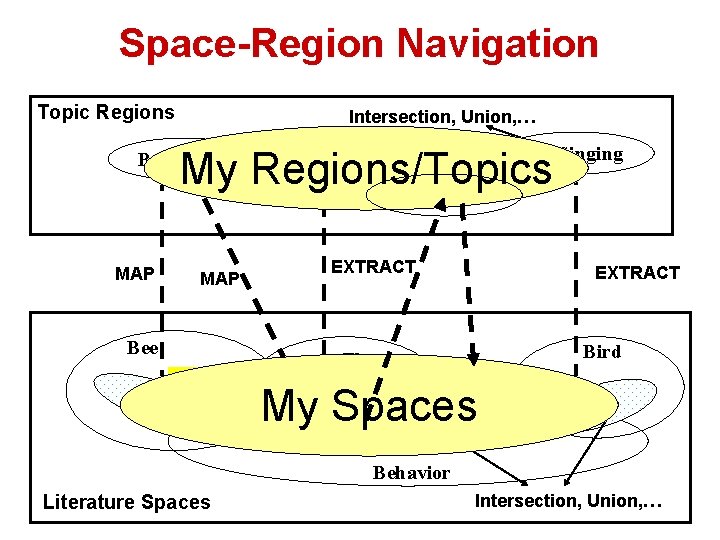
Space-Region Navigation Topic Regions … My Regions/Topics Intersection, Union, … Fly Rover Bee Forager MAP EXTRACT MAP Bee Bird Singing EXTRACT … My Spaces Fly Bird SWITCHING Behavior Literature Spaces Intersection, Union, …

MAP: Topic/Region Space • • MAP: Use the topic/region description as a query to search a given space Retrieval algorithm: – Query word distribution: p(w| Q) – Document word distribution: p(w| D) – Score a document based on similarity of Q and D • Leverage existing retrieval toolkits: Lemur/Indri

EXTRACT: Space Topic/Region • Assume k topics, each being represented by a word distribution • Use a k-component mixture model to fit the documents in a given space (EM algorithm) • The estimated k component word distributions are taken as k topic regions Likelihood: Maximum likelihood estimator: Bayesian estimator:
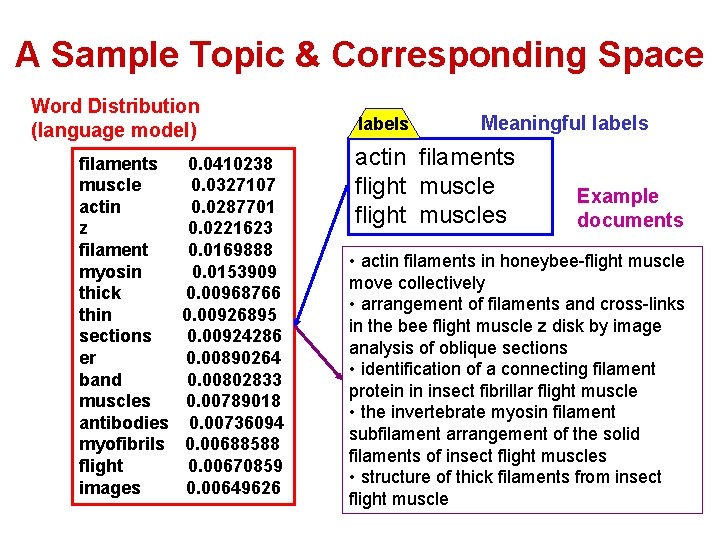
A Sample Topic & Corresponding Space Word Distribution (language model) filaments muscle actin z filament myosin thick thin sections er band muscles antibodies myofibrils flight images 0. 0410238 0. 0327107 0. 0287701 0. 0221623 0. 0169888 0. 0153909 0. 00968766 0. 00926895 0. 00924286 0. 00890264 0. 00802833 0. 00789018 0. 00736094 0. 00688588 0. 00670859 0. 00649626 labels Meaningful labels actin filaments flight muscles Example documents • actin filaments in honeybee-flight muscle move collectively • arrangement of filaments and cross-links in the bee flight muscle z disk by image analysis of oblique sections • identification of a connecting filament protein in insect fibrillar flight muscle • the invertebrate myosin filament subfilament arrangement of the solid filaments of insect flight muscles • structure of thick filaments from insect flight muscle

Incorporating Topic Priors • Either topic extraction or clustering: – User exploration: usually has preference. – E. g. , want one topic/cluster is about foraging behavior • Use prior to guild topic extraction – Prior as a simple language model – E. g. forage 0. 2; foraging 0. 3; food 0. 05; etc.
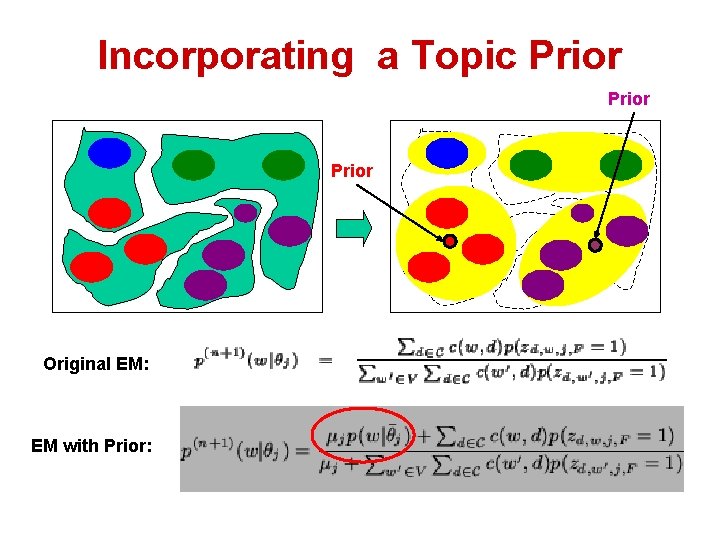
Incorporating a Topic Prior Original EM: EM with Prior:
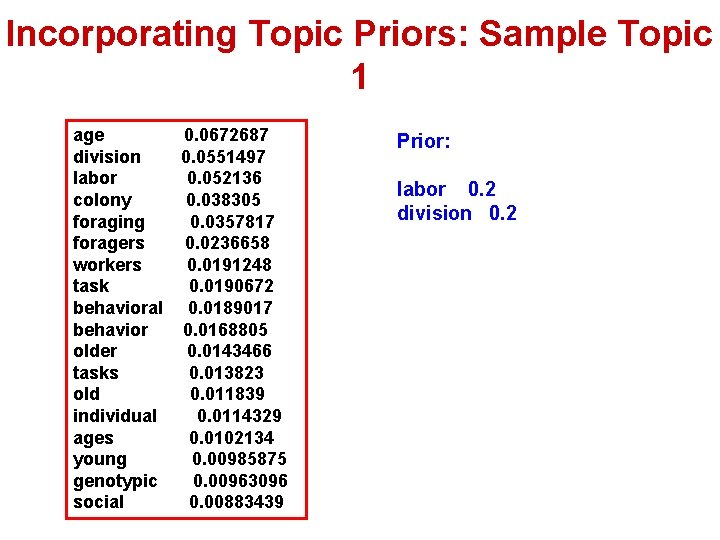
Incorporating Topic Priors: Sample Topic 1 age 0. 0672687 division 0. 0551497 labor 0. 052136 colony 0. 038305 foraging 0. 0357817 foragers 0. 0236658 workers 0. 0191248 task 0. 0190672 behavioral 0. 0189017 behavior 0. 0168805 older 0. 0143466 tasks 0. 013823 old 0. 011839 individual 0. 0114329 ages 0. 0102134 young 0. 00985875 genotypic 0. 00963096 social 0. 00883439 Prior: labor 0. 2 division 0. 2
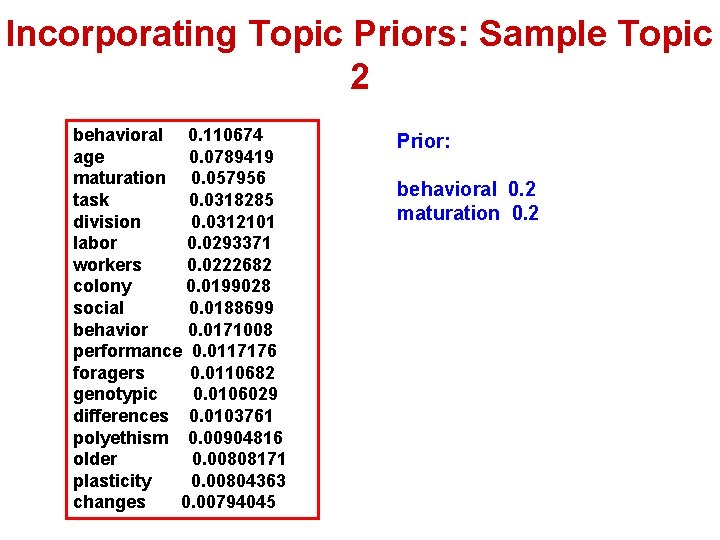
Incorporating Topic Priors: Sample Topic 2 behavioral 0. 110674 age 0. 0789419 maturation 0. 057956 task 0. 0318285 division 0. 0312101 labor 0. 0293371 workers 0. 0222682 colony 0. 0199028 social 0. 0188699 behavior 0. 0171008 performance 0. 0117176 foragers 0. 0110682 genotypic 0. 0106029 differences 0. 0103761 polyethism 0. 00904816 older 0. 00808171 plasticity 0. 00804363 changes 0. 00794045 Prior: behavioral 0. 2 maturation 0. 2
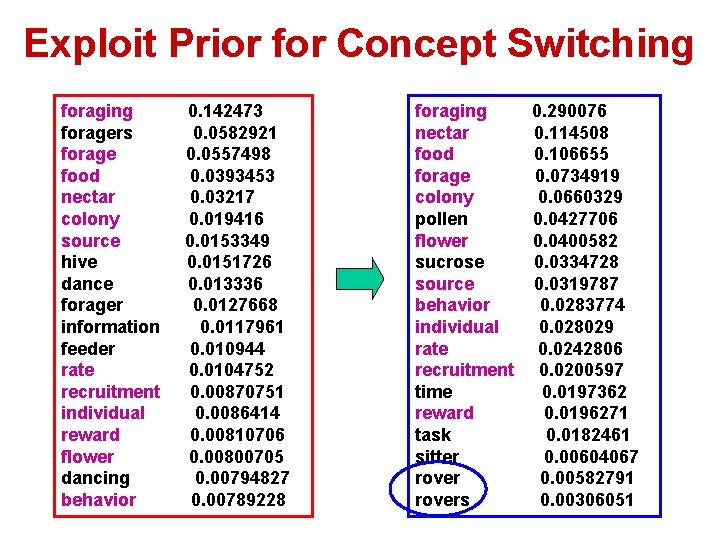
Exploit Prior for Concept Switching foragers forage food nectar colony source hive dance forager information feeder rate recruitment individual reward flower dancing behavior 0. 142473 0. 0582921 0. 0557498 0. 0393453 0. 03217 0. 019416 0. 0153349 0. 0151726 0. 013336 0. 0127668 0. 0117961 0. 010944 0. 0104752 0. 00870751 0. 0086414 0. 00810706 0. 00800705 0. 00794827 0. 00789228 foraging 0. 290076 nectar 0. 114508 food 0. 106655 forage 0. 0734919 colony 0. 0660329 pollen 0. 0427706 flower 0. 0400582 sucrose 0. 0334728 source 0. 0319787 behavior 0. 0283774 individual 0. 028029 rate 0. 0242806 recruitment 0. 0200597 time 0. 0197362 reward 0. 0196271 task 0. 0182461 sitter 0. 00604067 rover 0. 00582791 rovers 0. 00306051

Part 3: Task Support

Gene Summarization • Task: Automatically generate a text summary for a given gene • Challenge: Need to summarize different aspects of a gene • Standard summarization methods would generate an unstructured summary • Solution: A new method for generating semistructured summaries

An Ideal Gene Summary • http: //flybase. bio. indiana. edu/. bin/fbidq. html? FBgn 0000017 GP EL SI GI MP WFPI
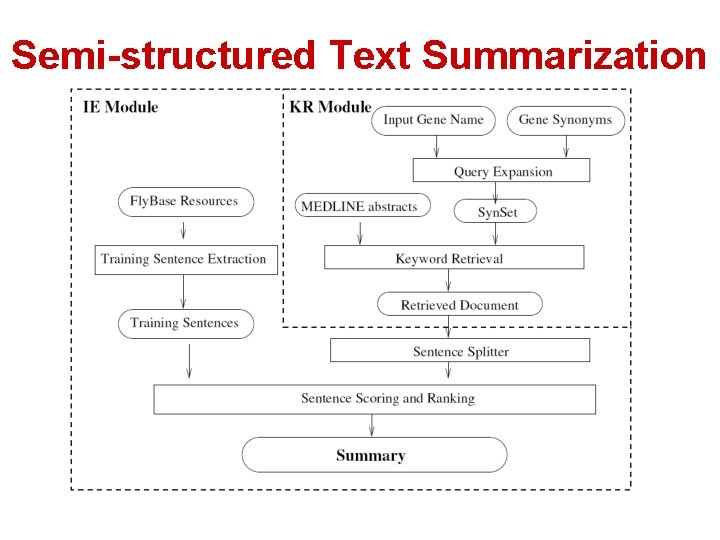
Semi-structured Text Summarization

Summary example (Abl)

A General Entity Summarizer • • • Task: Given any entity and k aspects to summarize, generate a semi-structured summary Assumption: Training sentences available for each aspect Method: – Train a recognizer for each aspect – Given an entity, retrieve sentences relevant to the entity – Classify each sentence into one of the k aspects – Choose the best sentences in each category

Summary • • All the methods we developed are – General – Scalable The problems are hard, but good progress has been made in all the directions – The V 3 system has only incorporated the basic research results – More advanced technologies are available for immediate implementation • • • Better tokenization for retrieval Domain adaptation techniques Automatic topic labeling General entity summarizer More research to be done in – – Entity & relation extraction Graph mining/question answering Domain adaptation Active learning
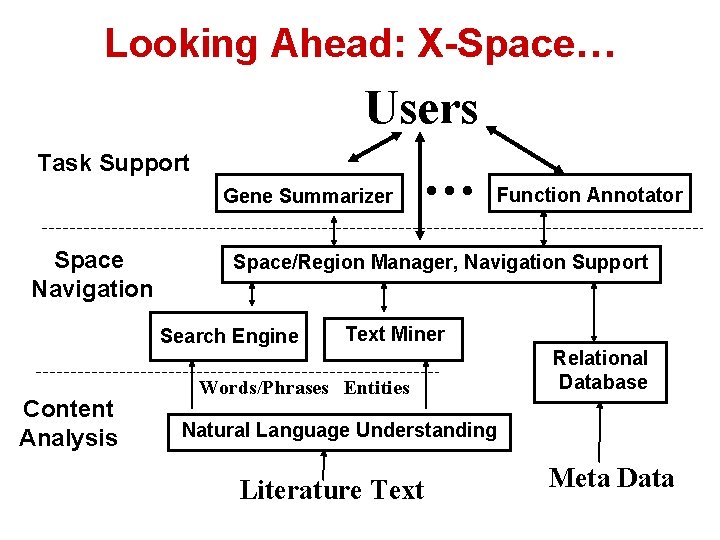
Looking Ahead: X-Space… Users Task Support Gene Summarizer Space Navigation Function Annotator Space/Region Manager, Navigation Support Search Engine Content Analysis … Text Miner Words/Phrases Entities Relational Database Natural Language Understanding Literature Text Meta Data

Thank You! Questions?
- Slides: 33