Bayesian inference with Mr Bayes Molecular Phylogenetics exercise

Bayesian inference with Mr. Bayes Molecular Phylogenetics – exercise Ben Stöver WS 2012/2013

Bayesian inference with Mr. Bayes 1. Mr. Bayes 2 1. 1 Overview on Mr. Bayes • • • Implements the Metropolis-Hastings-Green-algorithm. Analyses can be started from the internal command line (just like in PAUP*). The help command displays a list of available commands. help <command> displays help to a certain commend. More information under http: //mrbayes. sourceforge. net/mb 3. 2_manual. pdf Command reference under http: //mrbayes. sourceforge. net/Help/ Ben Stöver 13. 02. 2012 Winter term 2012/2013 Fortgeschrittenenmodul Molecular Phylogenetics

Bayesian inference with Mr. Bayes 1. Mr. Bayes 3 1. 2 Important commands to start an analysis • execute <alignment. File>: Reads alignment data from a file (e. g. Nexus). • lset: Can be used to set the model parameters (e. g. nst=6 rates=invgamma sets the GTR+I+G model) • mcmc: Starts an MCMC analysis with the loaded alignment. • ngen: The number of generations for the chain to run. • samplefreq: The frequency of generations when a tree shall be written to the log file • printfreq: The frequency to print information on the current run to the command line. • diagnfreq: The frequency to analize the current process (e. g. the measure of the similarity of the current tree samples). • savebrlens: Specify yes here, if branch lengths shall be saved. • nchains: Specifies the number of parallel chains (default is 4). Ben Stöver 13. 02. 2012 Winter term 2012/2013 Fortgeschrittenenmodul Molecular Phylogenetics

Bayesian inference with Mr. Bayes 1. Mr. Bayes 4 1. 3 Important commands to finish an analysis • plot: • sump: manual) • sumt: Ben Stöver Plots the deveopment of an estimated parameter over time (The parameter to be displayed can be selected. ) Summarize samples of model parameters (section 2. 2. 8 in Summarize the tree samples 13. 02. 2012 Winter term 2012/2013 Fortgeschrittenenmodul Molecular Phylogenetics

Bayesian inference with Mr. Bayes 1. Mr. Bayes 5 1. 4 Running an analysis • During an analysis the current ln(linkelihood) of all chains are printed out. run 1 diagnosis output heated chains Ben Stöver 13. 02. 2012 run 2 cold chain Winter term 2012/2013 Fortgeschrittenenmodul Molecular Phylogenetics
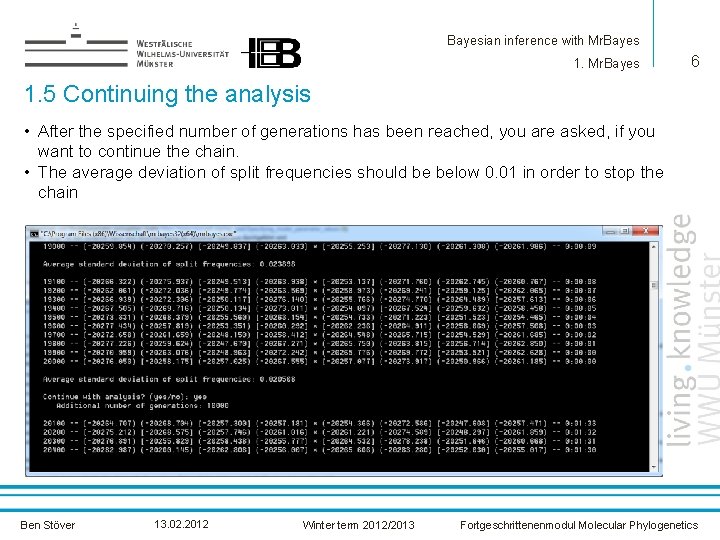
Bayesian inference with Mr. Bayes 1. Mr. Bayes 6 1. 5 Continuing the analysis • After the specified number of generations has been reached, you are asked, if you want to continue the chain. • The average deviation of split frequencies should be below 0. 01 in order to stop the chain Ben Stöver 13. 02. 2012 Winter term 2012/2013 Fortgeschrittenenmodul Molecular Phylogenetics
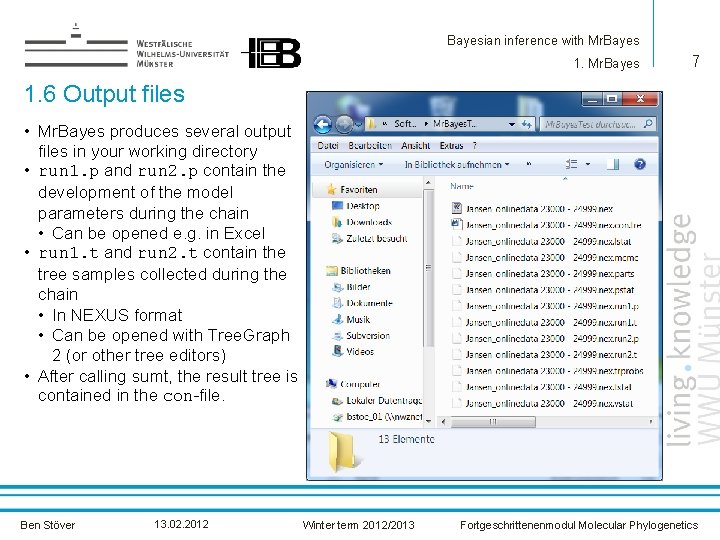
Bayesian inference with Mr. Bayes 1. Mr. Bayes 7 1. 6 Output files • Mr. Bayes produces several output files in your working directory • run 1. p and run 2. p contain the development of the model parameters during the chain • Can be opened e. g. in Excel • run 1. t and run 2. t contain the tree samples collected during the chain • In NEXUS format • Can be opened with Tree. Graph 2 (or other tree editors) • After calling sumt, the result tree is contained in the con-file. Ben Stöver 13. 02. 2012 Winter term 2012/2013 Fortgeschrittenenmodul Molecular Phylogenetics
- Slides: 7