Basics of f MRI Preprocessing Douglas N Greve

Basics of f. MRI Preprocessing Douglas N. Greve greve@nmr. mgh. harvard. edu 1
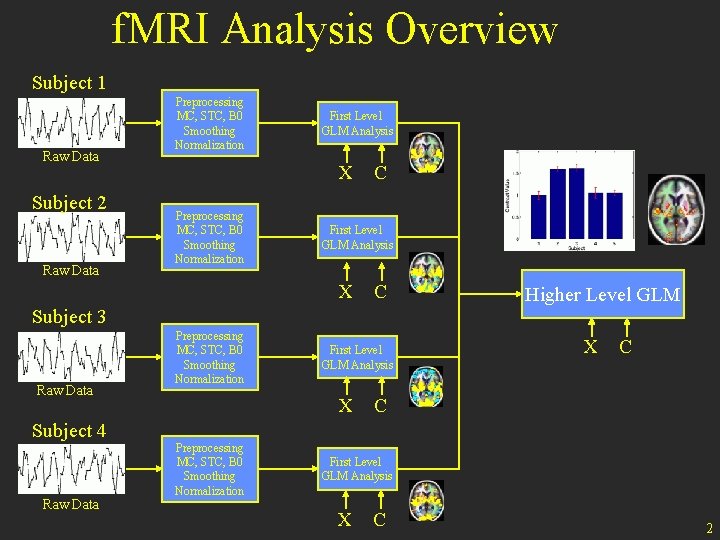
f. MRI Analysis Overview Subject 1 Raw Data Subject 2 Raw Data Preprocessing MC, STC, B 0 Smoothing Normalization First Level GLM Analysis X Preprocessing MC, STC, B 0 Smoothing Normalization C First Level GLM Analysis X C Subject 3 Raw Data Subject 4 Raw Data Preprocessing MC, STC, B 0 Smoothing Normalization First Level GLM Analysis X Preprocessing MC, STC, B 0 Smoothing Normalization Higher Level GLM X C C First Level GLM Analysis X C 2

Preprocessing • Assures that assumptions of the analysis are met • Time course comes from a single location • Uniformly spaced in time • Spatial “smoothness” • Analysis – separating signal from noise 3
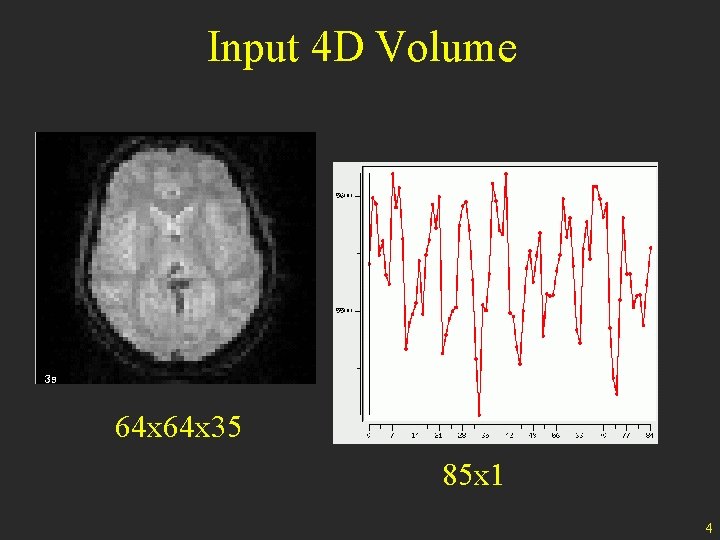
Input 4 D Volume 64 x 35 85 x 1 4

Preprocessing • 1. 2. 3. 4. 5. • Start with a 4 D data set Motion Correction Slice-Timing Correction B 0 Distortion Correction Spatial Normalization Spatial Smoothing End with a 4 D data set • • • Can be done in other orders Not everything is always done Note absence of temporal filtering 5

Motion • Analysis assumes that time course represents a value from a single location • Subjects move • Shifts can cause noise, uncertainty • Edge of the brain and tissue boundaries 6
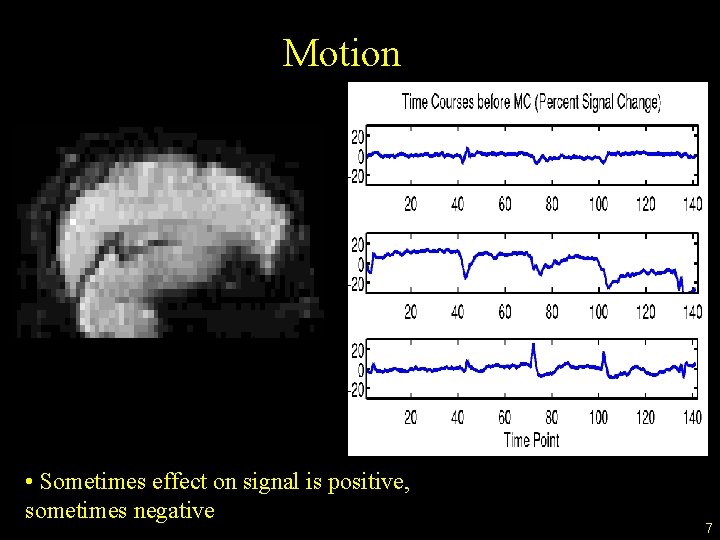
Motion • Sometimes effect on signal is positive, sometimes negative 7
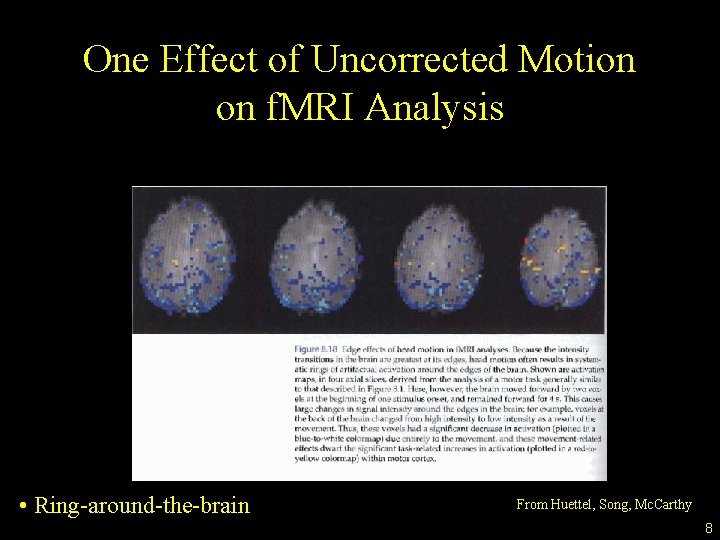
One Effect of Uncorrected Motion on f. MRI Analysis • Ring-around-the-brain From Huettel, Song, Mc. Carthy 8
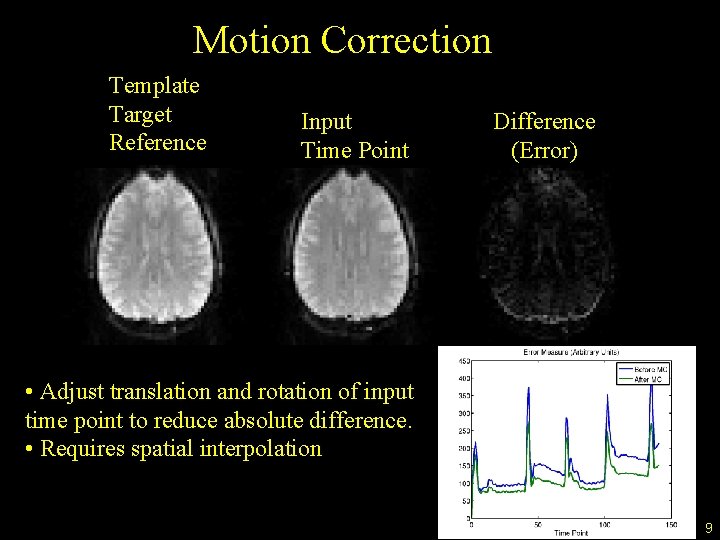
Motion Correction Template Target Reference Input Time Point Difference (Error) • Adjust translation and rotation of input time point to reduce absolute difference. • Requires spatial interpolation 9
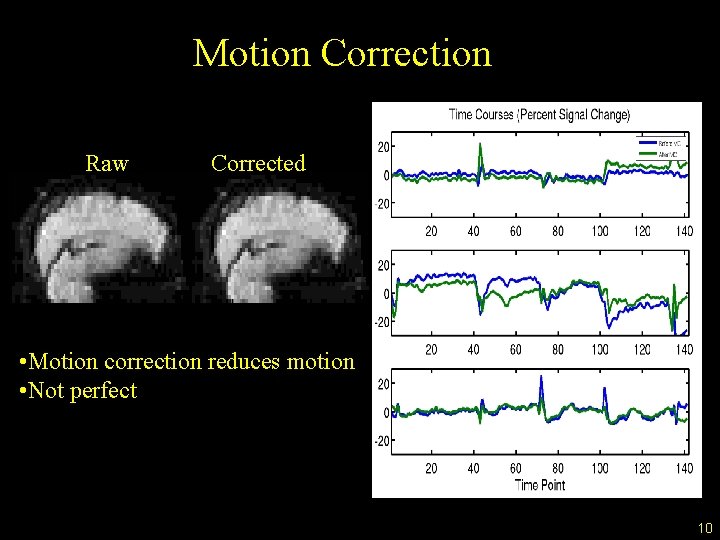
Motion Correction Raw Corrected • Motion correction reduces motion • Not perfect 10
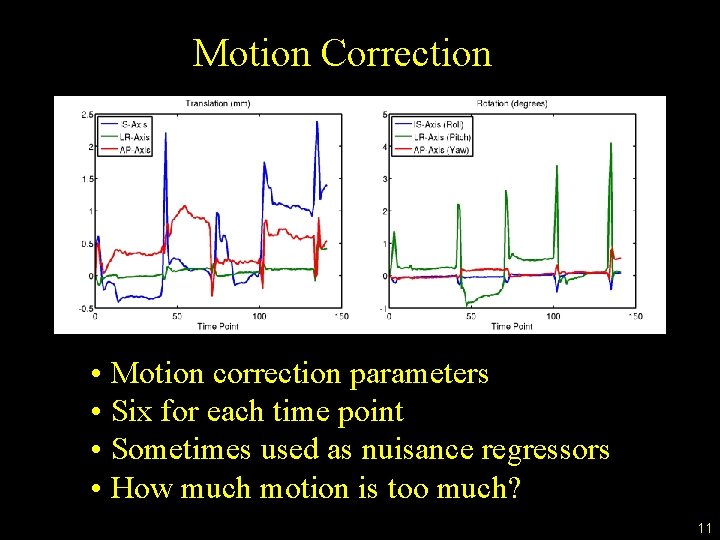
Motion Correction • Motion correction parameters • Six for each time point • Sometimes used as nuisance regressors • How much motion is too much? 11

Slice Timing Ascending Interleaved • Volume not acquired all at one time • Acquired slice-by-slice • Each slice has a different delay 12
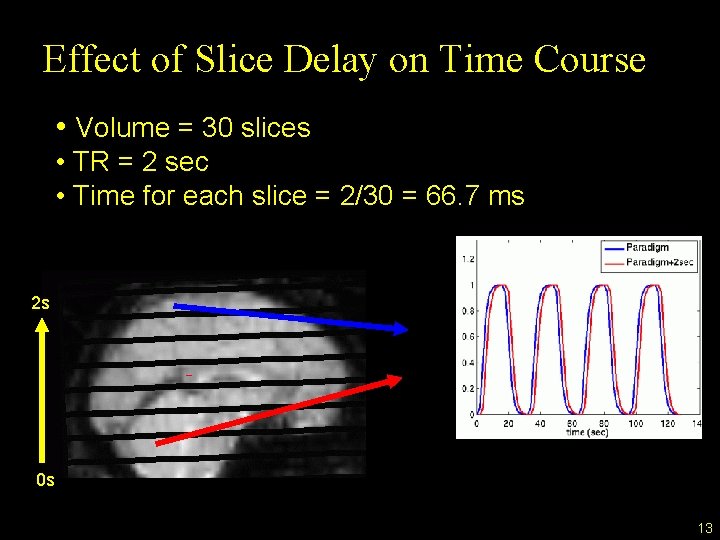
Effect of Slice Delay on Time Course • Volume = 30 slices • TR = 2 sec • Time for each slice = 2/30 = 66. 7 ms 2 s 0 s 13
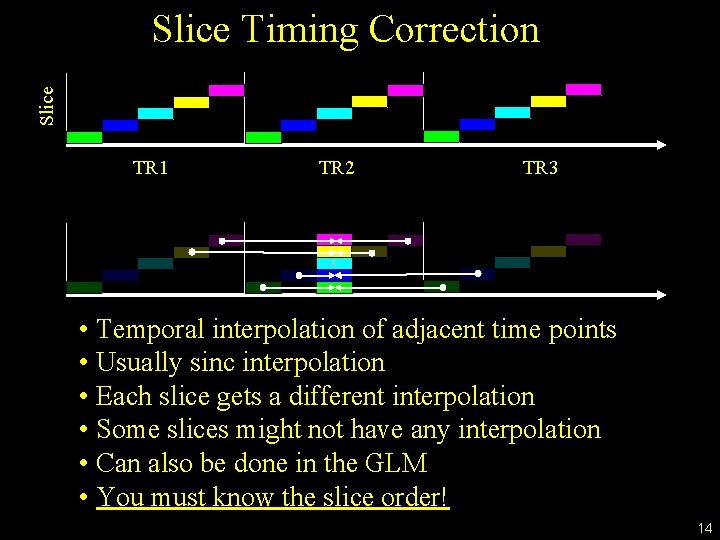
Slice Timing Correction TR 1 TR 2 TR 3 X • Temporal interpolation of adjacent time points • Usually sinc interpolation • Each slice gets a different interpolation • Some slices might not have any interpolation • Can also be done in the GLM • You must know the slice order! 14

Motion, Slice Timing, Spin History • Every other slice is bright • Interleaved slice acquisition • Happens with sequential, harder to see 15
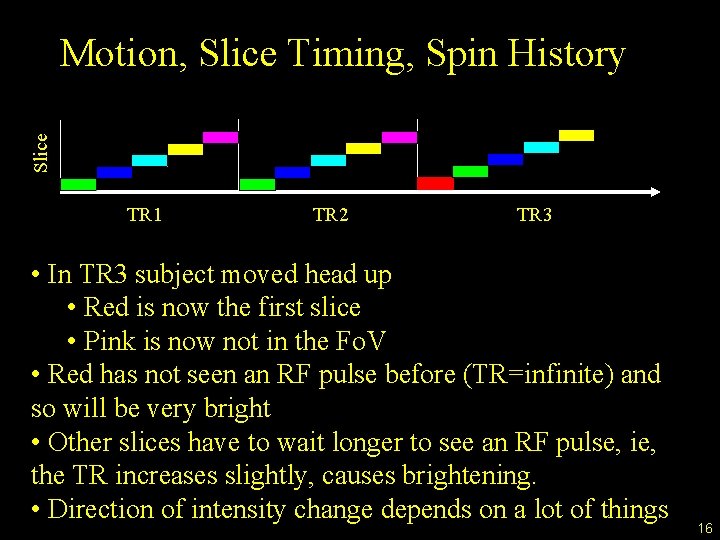
Slice Motion, Slice Timing, Spin History TR 1 TR 2 TR 3 • In TR 3 subject moved head up • Red is now the first slice • Pink is now not in the Fo. V • Red has not seen an RF pulse before (TR=infinite) and so will be very bright • Other slices have to wait longer to see an RF pulse, ie, the TR increases slightly, causes brightening. • Direction of intensity change depends on a lot of things 16
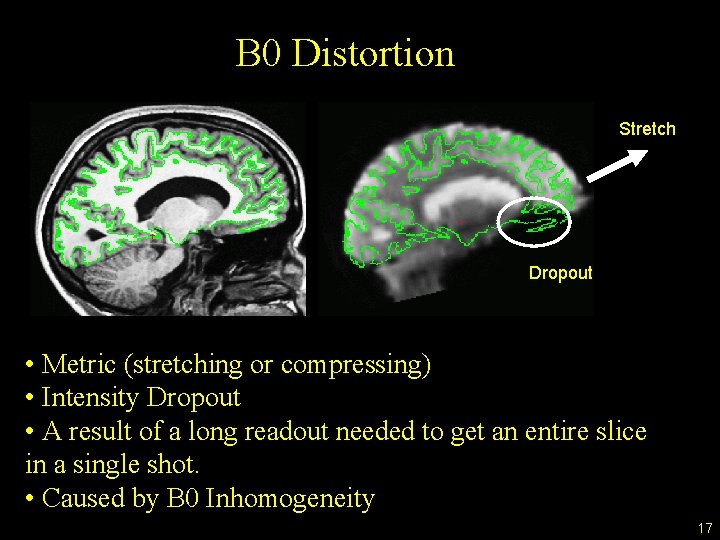
B 0 Distortion Stretch Dropout • Metric (stretching or compressing) • Intensity Dropout • A result of a long readout needed to get an entire slice in a single shot. • Caused by B 0 Inhomogeneity 17
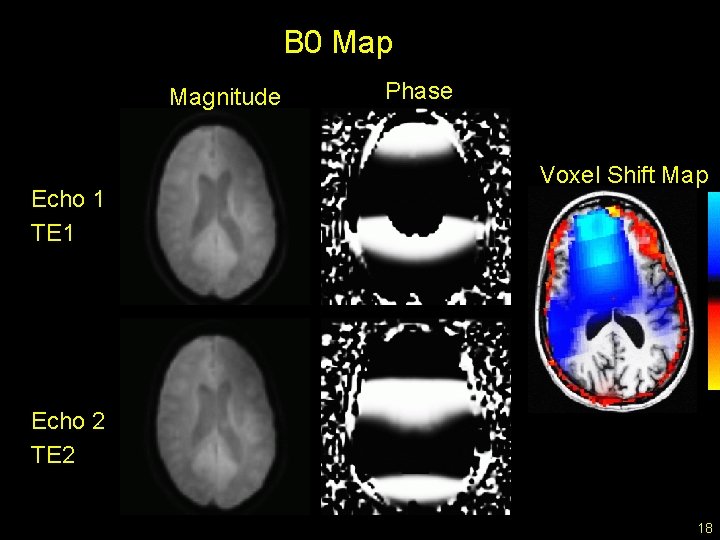
B 0 Map Magnitude Echo 1 TE 1 Phase Voxel Shift Map Echo 2 TE 2 18
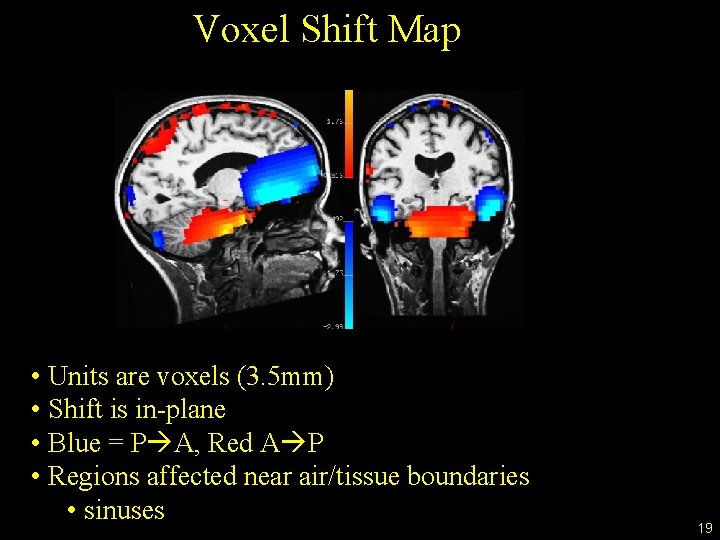
Voxel Shift Map • Units are voxels (3. 5 mm) • Shift is in-plane • Blue = P A, Red A P • Regions affected near air/tissue boundaries • sinuses 19
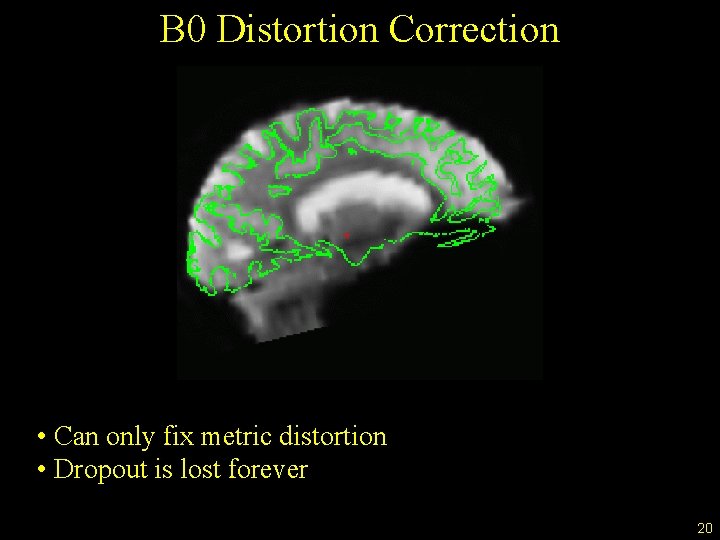
B 0 Distortion Correction • Can only fix metric distortion • Dropout is lost forever 20

B 0 Distortion Correction • Can only fix metric distortion • Dropout is lost forever • Interpolation • Need: • “Echo spacing” – readout time • Phase encode direction • More important for surface than for volume • Important when combining from different scanners 21

Spatial Normalization • Transform volume into another volume • Re-slicing, re-gridding • New volume is an “atlas” space • Align brains of different subjects so that a given voxel represents the “same” location. • Similar to motion correction • Preparation for comparing across subjects • Volume-based • Surface-based • Combined Volume-surface-based (CVS) 22
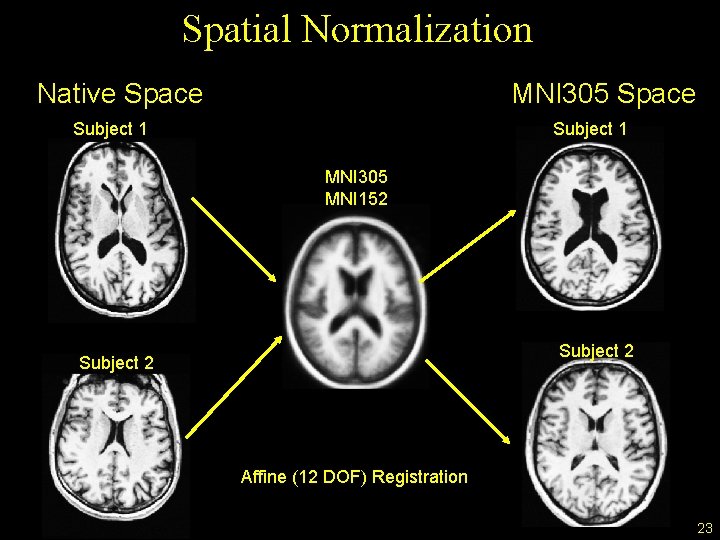
Spatial Normalization Native Space MNI 305 Space Subject 1 MNI 305 MNI 152 Subject 2 Affine (12 DOF) Registration 23
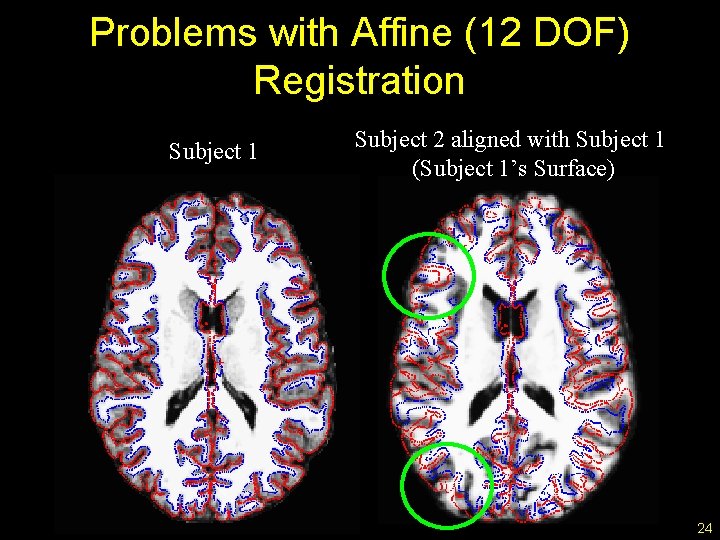
Problems with Affine (12 DOF) Registration Subject 1 Subject 2 aligned with Subject 1 (Subject 1’s Surface) 24
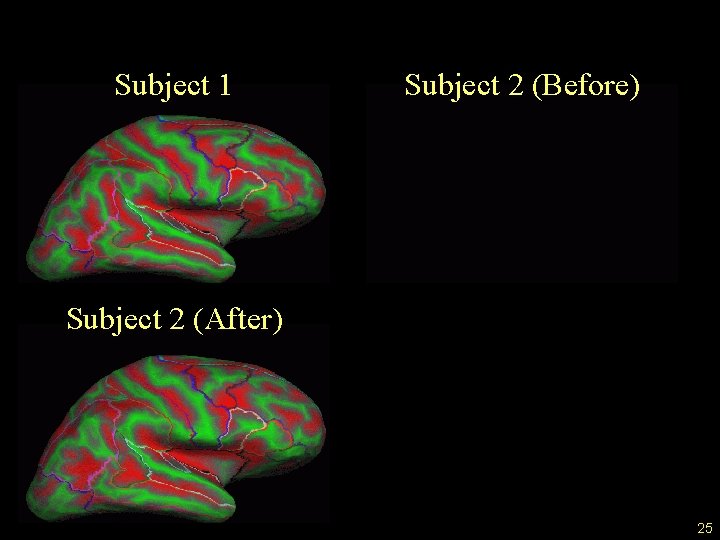
Surface Registration Subject 1 Subject 2 (Before) Subject 2 (After) 25
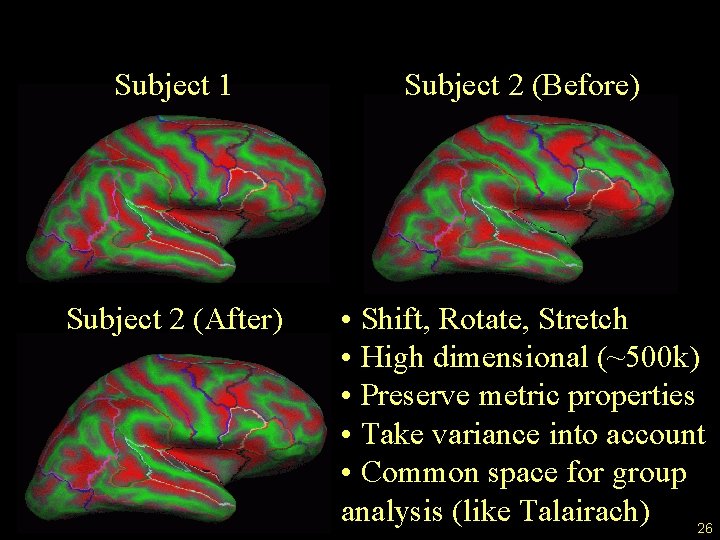
Surface Registration Subject 1 Subject 2 (Before) Subject 2 (After) • Shift, Rotate, Stretch • High dimensional (~500 k) • Preserve metric properties • Take variance into account • Common space for group analysis (like Talairach) 26

Spatial Smoothing • Replace voxel value with a weighted average of nearby voxels (spatial convolution) • Weighting is usually Gaussian • 3 D (volume) • 2 D (surface) • Do after all interpolation, before computing a standard deviation • Similarity to interpolation • Improve SNR • Improve Intersubject registration • Can have a dramatic effect on your results 27
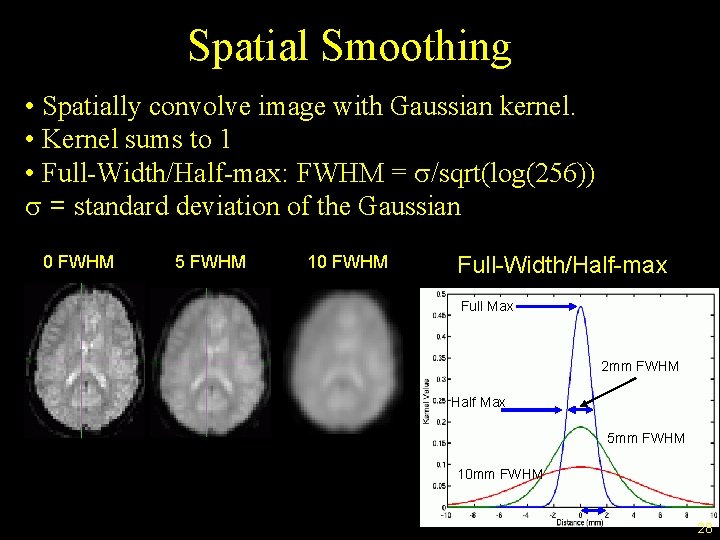
Spatial Smoothing • Spatially convolve image with Gaussian kernel. • Kernel sums to 1 • Full-Width/Half-max: FWHM = s/sqrt(log(256)) s = standard deviation of the Gaussian 0 FWHM 5 FWHM 10 FWHM Full-Width/Half-max Full Max 2 mm FWHM Half Max 5 mm FWHM 10 mm FWHM 28
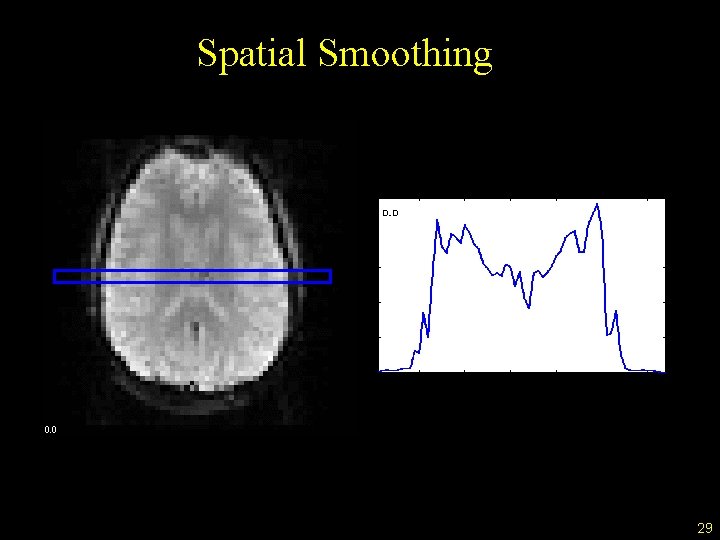
Spatial Smoothing 29
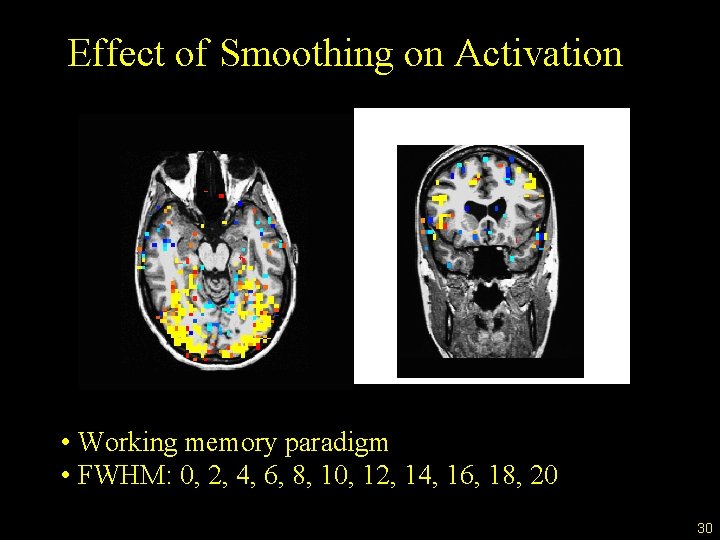
Effect of Smoothing on Activation • Working memory paradigm • FWHM: 0, 2, 4, 6, 8, 10, 12, 14, 16, 18, 20 30
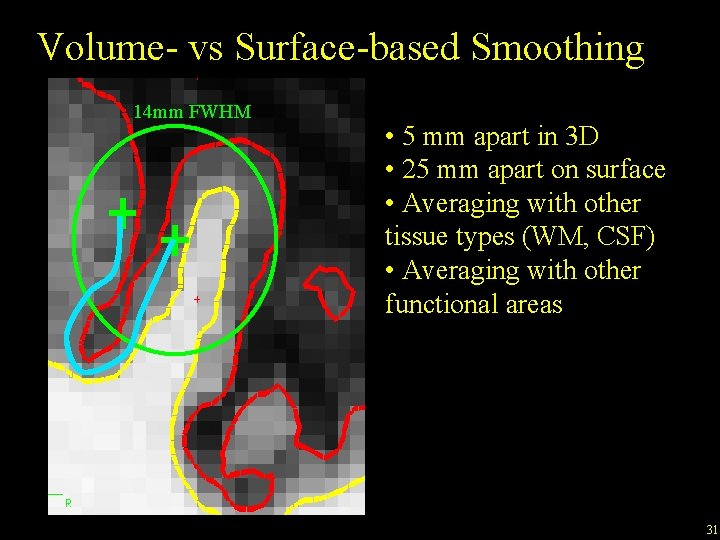
Volume- vs Surface-based Smoothing 14 mm FWHM • 5 mm apart in 3 D • 25 mm apart on surface • Averaging with other tissue types (WM, CSF) • Averaging with other functional areas 31

Choosing the FWHM • No hard and fast rules • Matched filter theorem – set equal to activation size • How big is that? • Changes with brain location • Changes with contrast • May change with subject, population, etc • Two voxels – to meet the assumptions of Gaussian Random Fields (GRF) for correction of multiple comparisons 32

Temporal filtering • Replace value at a given time point with a weighted average of its neighbors • Should/Is NOT be done as a preprocessing step – contrary to what many people think. • Belongs in analysis. 33
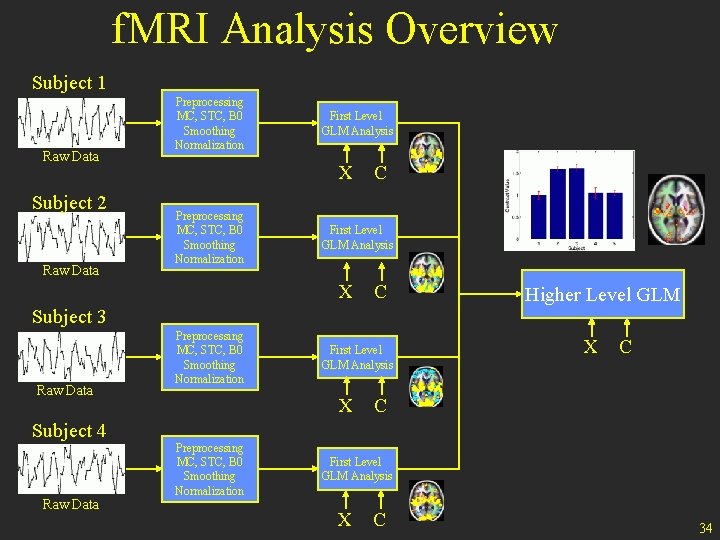
f. MRI Analysis Overview Subject 1 Raw Data Subject 2 Raw Data Preprocessing MC, STC, B 0 Smoothing Normalization First Level GLM Analysis X Preprocessing MC, STC, B 0 Smoothing Normalization C First Level GLM Analysis X C Subject 3 Raw Data Subject 4 Raw Data Preprocessing MC, STC, B 0 Smoothing Normalization First Level GLM Analysis X Preprocessing MC, STC, B 0 Smoothing Normalization Higher Level GLM X C C First Level GLM Analysis X C 34

Preprocessing • 1. 2. 3. 4. 5. • Start with a 4 D data set Motion Correction - Interpolation Slice-Timing Correction B 0 Distortion Correction - Interpolation Spatial Normalization - Interpolation Spatial Smoothing – Interpolation-like End with a 4 D data set • • Can be done in other orders Note absence of temporal filtering 35

36
- Slides: 36