Basic metrics of food webs A pitcher plant
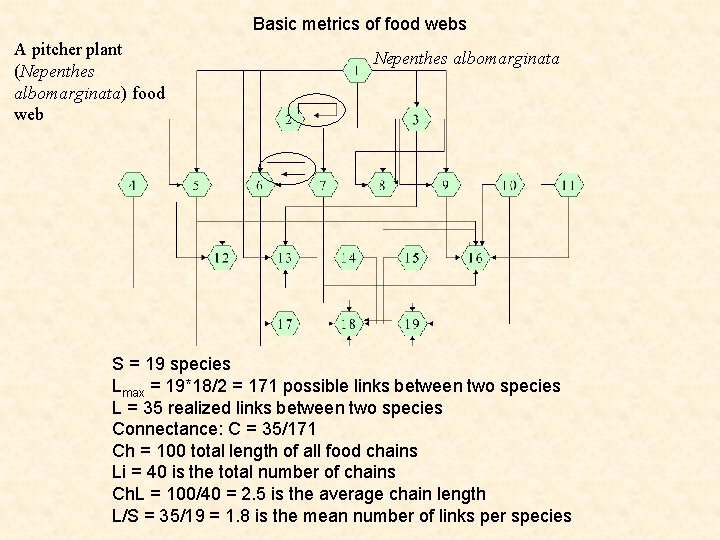
Basic metrics of food webs A pitcher plant (Nepenthes albomarginata) food web Nepenthes albomarginata S = 19 species Lmax = 19*18/2 = 171 possible links between two species L = 35 realized links between two species Connectance: C = 35/171 Ch = 100 total length of all food chains Li = 40 is the total number of chains Ch. L = 100/40 = 2. 5 is the average chain length L/S = 35/19 = 1. 8 is the mean number of links per species
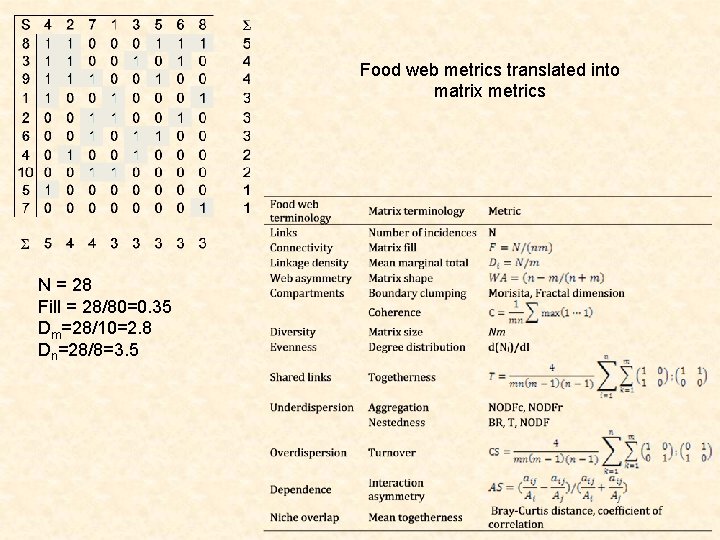
Food web metrics translated into matrix metrics N = 28 Fill = 28/80=0. 35 Dm=28/10=2. 8 Dn=28/8=3. 5
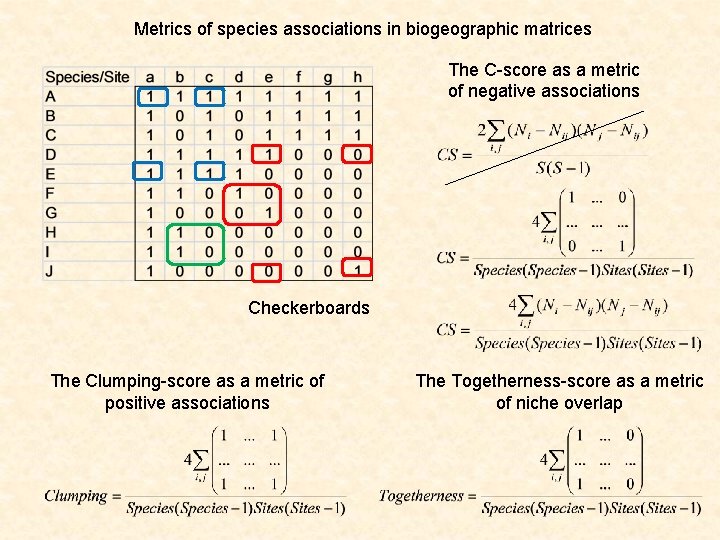
Metrics of species associations in biogeographic matrices The C-score as a metric of negative associations Checkerboards The Clumping-score as a metric of positive associations The Togetherness-score as a metric of niche overlap
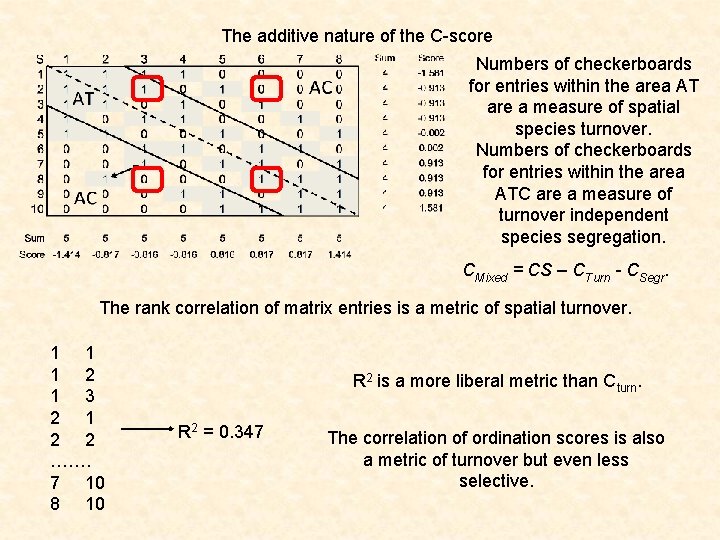
The additive nature of the C-score Numbers of checkerboards for entries within the area AT are a measure of spatial species turnover. Numbers of checkerboards for entries within the area ATC are a measure of turnover independent species segregation. CMixed = CS – CTurn - CSegr. The rank correlation of matrix entries is a metric of spatial turnover. 1 1 1 2 1 3 2 1 2 2 ……. 7 10 8 10 R 2 is a more liberal metric than Cturn. R 2 = 0. 347 The correlation of ordination scores is also a metric of turnover but even less selective.
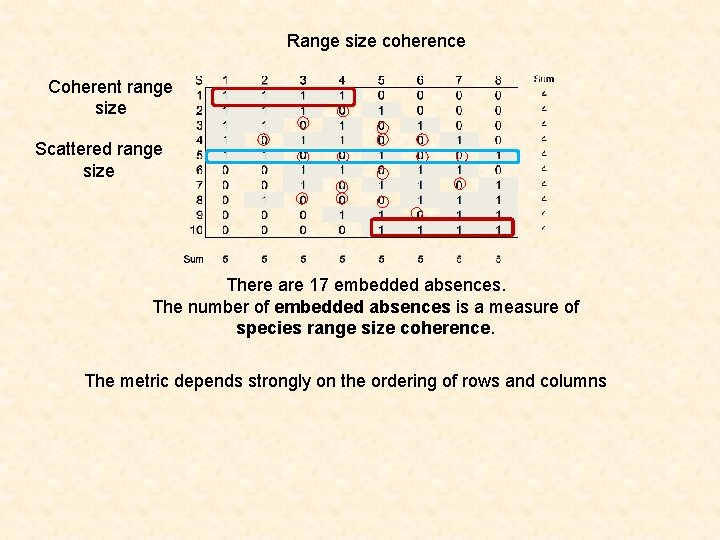
Range size coherence Coherent range size Scattered range size There are 17 embedded absences. The number of embedded absences is a measure of species range size coherence. The metric depends strongly on the ordering of rows and columns

The measurement of nestedness The distance concept of nestedness. Sort the matrix rows and olumns according to some gradient. Define an isocline that divides the matrix into a perfectly filled an empty part. The normalized squared sum of relative distances of unexpected absences and unexpected presences is now a metric of nestednessis.
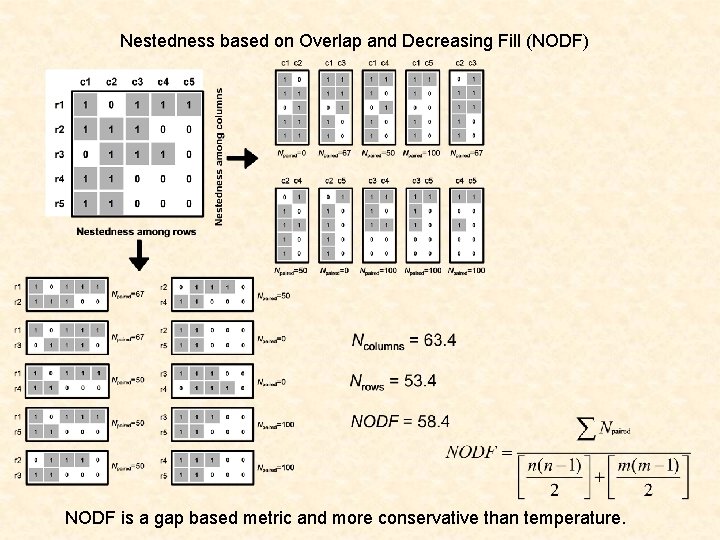
Nestedness based on Overlap and Decreasing Fill (NODF) NODF is a gap based metric and more conservative than temperature.

The disorder measure of Brualdi and Sanderson Ho many cells must be filled or emptied to achieve a perfectly ordered matrix. The Brualdi Sanderson measure is a count of this number Discrepancy is a gap counting metric.
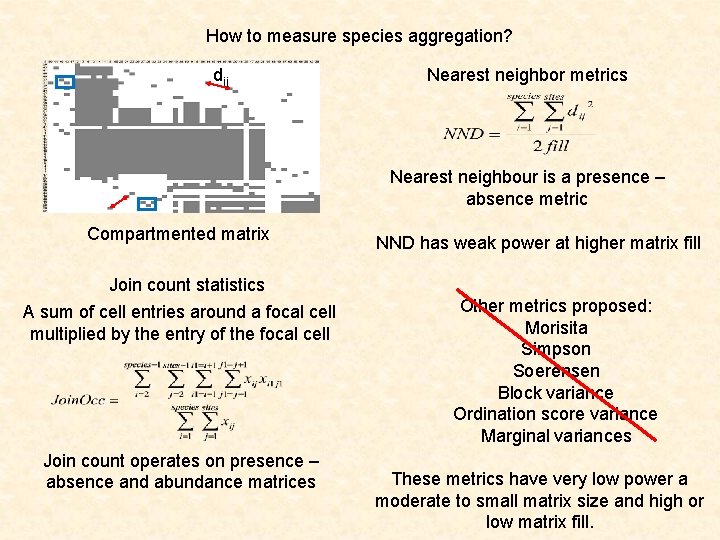
How to measure species aggregation? dij Nearest neighbor metrics Nearest neighbour is a presence – absence metric Compartmented matrix NND has weak power at higher matrix fill Join count statistics A sum of cell entries around a focal cell multiplied by the entry of the focal cell Join count operates on presence – absence and abundance matrices Other metrics proposed: Morisita Simpson Soerensen Block variance Ordination score variance Marginal variances These metrics have very low power a moderate to small matrix size and high or low matrix fill.

Abundance based metrics The C-score extension The metric CA is a count of the number of abundance checkerboards in the matrix. Other 2 x 2 submatrices catch matrix properties that have not well defined ecological meaning.

Nestedness in abundance matrices The metric is a sum of all pairs in the matrix (first sorted accoding to species richness then sorted according to weights), where the weight in the row/column of lower species richness is smaller than the weight in the row/column of higher species richness

A complete table of methods for co-occurrence analysis Segregated Independent of matrix sorting C-score Segregated Dependent of matrix sorting Whole matrix Aggregated nested Clumping score CS/Clumping Nest. Pairs Togetherness Species only Aggregated Simpson dissimilarity Simpson similarity Soerensen dissimilarity Morisita Chao Other joint occurrence/absence metrics Whole matrix Aggregated Segregated Aggregated nested NND Block Join-coint Other distance based metrics NODF BR T Species only PA null models PA A A and PA All Data type PA null models PA PA A and PA No fixed - fixed All A null models All PA No fixed - fixed Data type PA null models PA A and PA PA PA All All Segregated Aggregated Data type PA null models Embedded absences r 2 CTurn CSegr PA PA PA All All All For seriation Morisita A null models All Data type A null models All All A null models

Pattern detection in large matrices Pajek: software for social network analysis WAND: ecological network analysis These programs use cluster analysis and ordination to sort the matrix according to numbers of occurrences. Didstance metrics are then used to identify compartments. Klique. Finder: software for compartment analysis They generate hypotheses about matrix structure. They do not fully allow for statistical inference.
- Slides: 14