Basic methods in genetics PCR Polymerase Chain Reaction

Basic methods in genetics • PCR; Polymerase Chain Reaction • Restriction enzyme digestions • Gel electrophoresis

PCR; Polymerase Chain Reaction • • Amplification of specific DNA sequences Invented by Kary Mullis in 1983 Revolutionized the world of molecular biology Mimics cell’s own DNA replication machinery

• Ingredients in PCR: • DNA as a template • Thermo stable DNA polymerase enzyme • Deoxynucleoside triphosphates (d. NTPs) • Synthetic oligonucleotide primers The flanking sequence of the target locus needs to be known!

• Three major steps in PCR: • Denaturation: Strand separation at 95°C • Annealing: Hybridization of primers at 45 -60°C • Extension: DNA synthesis at 72°C • Three steps are repeated for 25 to 40 times • Heat activation at 95°C for “hot start enzymes”. • Final elongation step at 72°C
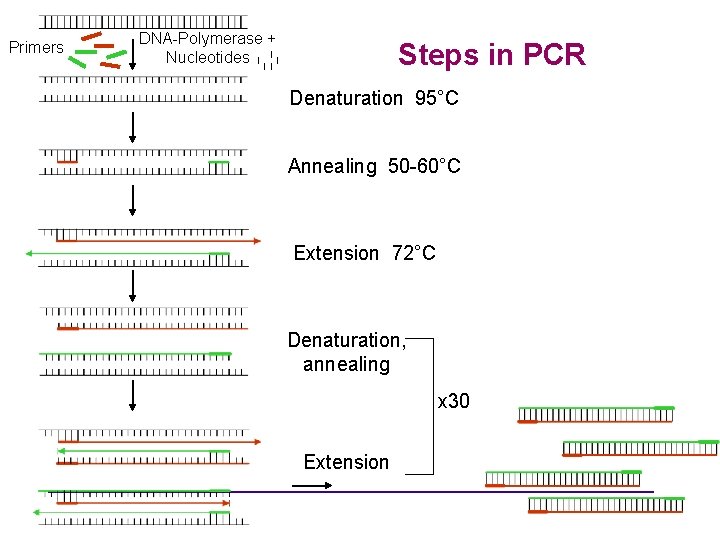
Primers DNA-Polymerase + Nucleotides Steps in PCR Denaturation 95°C Annealing 50 -60°C Extension 72°C Denaturation, annealing x 30 Extension

Technical problems • • Contamination Sensitivity to the levels of divalent cations Quality of template DNA Limited size of amplified product: • 300 bp-1000 bp are most efficient • possible to amplify fragments of several kb

Primer design • • Must be very specific No primer-primer interactions No “hairpin” formation No self-annealing • DNA sequence from the database (or from sequencing) • ENSEMBL: http: //www. ensembl. org/ • BLAST : http: //www. ncbi. nlm. nih. gov/BLAST/ • Primer 3 program: http: //www-genome. wi. mit. edu/cgibin/primer 3_www. cgi

What are enzymes? • Proteins that speed up chemical reactions in the body= protein catalyst essential to sustain life • Substrate = The molecule with which an enzyme interacts • Enzyme function is highly dependent on environmental characteristics such as temperature and p. H.

Restriction enzymes Nucleases: • exonucleases: remove nucleotides from the end of DNA or RNA • endonucleases: make cuts at internal phosphodiester bonds Restriction endonucleases: • Discovered in the late 1960 s • Found and purified from bacteria • Three types: • Type I and III: do not recognize a specific sequence to cut • Type II: cut specific recognized sequence

Type II enzymes: • Over 2500 different enzymes have been isolated • More than 500 enzymes are commercially available • Cut often sequence at palindromic hexanucleotide sequences • e. g. Eco. RI : GAATTC • CTTAAG • Most enzymes cut within the recognition sequence • Leaves “sticky” or “blunt” ends
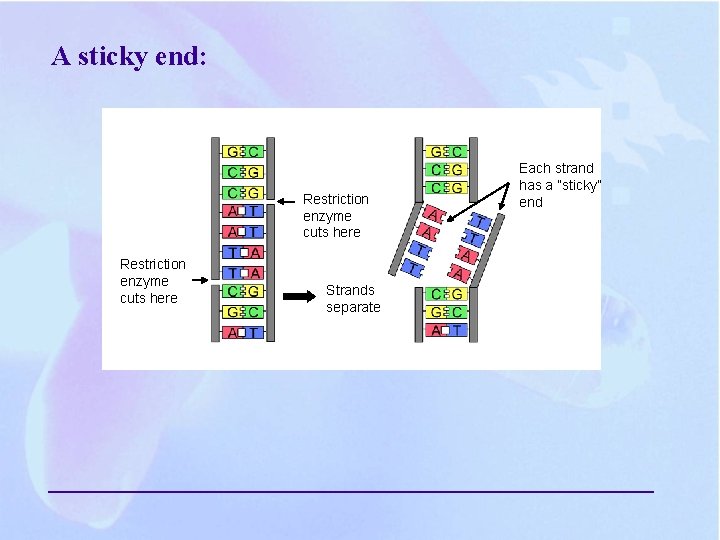
A sticky end: Restriction enzyme cuts here Strands separate Each strand has a ”sticky” end
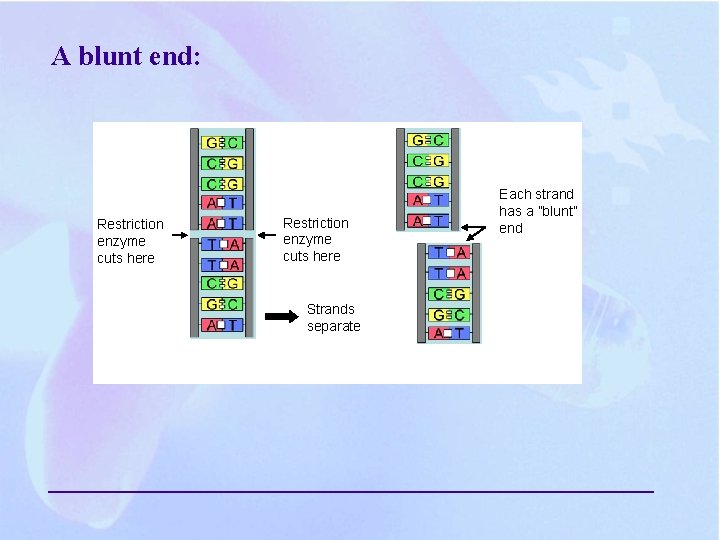
A blunt end: Restriction enzyme cuts here Strands separate Each strand has a ”blunt” end

The designing of digestion reactions: • Sequence of the target DNA must be known • Webcutter 2. 0: http: //www. firstmarket. com/cutter/cut 2. html

Gel electrophoresis • A method that separates macromolecules on the basis of: • size • electric charge • other physical properties • Gel acts as a support medium • Electric field is generated across the gel • DNA is negatively charged → migration towards the positive pole • Small molecules move faster than big molecules • Ethidium bromide staining • Intercalates between bases of DNA • Can be visualized under UV-light

Requirements in gel electrophoresis: • • An electrophoresis chamber and power supply Gel casting tray Sample comb Agarose Electrophoresis buffer Loading buffer DNA ladder (= size standard) Ethidium bromide • Transilluminator
- Slides: 19