Basic enzyme kinetics Aulanniam Biochemistry Laboratory Brawijaya University
Basic enzyme kinetics Aulanni’am Biochemistry Laboratory Brawijaya University Aulani " Biokimia Enzim Lanjut" Presentasi 5

Enzymes • Enzymes are catalysts, that speed up the rate of a reaction, without changing the extent of the reaction. • They are (in general) large proteins and are highly specific, i. e. , usually each enzyme speeds up only one single biochemical reaction. • They are highly regulated by a pile of things. Phosphorylation, calcium, ATP, their own products, etc, resulting in extremely complex webs of intracellular biochemical reactions. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Law of mass action Given a basic reaction A+B k 1 k-1 C we assume that the rate of forward reaction is linearly proportional to the concentrations of A and B, and the back reaction is linearly proportional to the concentration of C. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Equilibrium is reached when the net rate of reaction is zero. Thus or This equilibrium constant tells us the extent of the reaction, NOT its speed. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Basic problem of enzyme kinetics Suppose an enzyme were to react with a substrate, giving a product. S+E P+E If we simply applied the law of mass action to this reaction, the rate of reaction would be a linearly increasing function of [S]. As [S] gets very big, so would the reaction rate. This doesn’t happen. In reality, the reaction rate saturates. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Michaelis and Menten In 1913, Michaelis and Menten proposed the following mechanism for a saturating reaction rate S+E k 1 k-1 C k 2 P+E Complex. product • Easy to use mass action to derive the equations. • There are conservation constraints. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Equilibrium approximation And thus, since Thus reaction velocity Aulani " Biokimia Enzim Lanjut" Presentasi 5

Pseudo-steady state approximation And thus, since Thus reaction velocity Looks very similar to previous, but is actually quite different! Aulani " Biokimia Enzim Lanjut" Presentasi 5
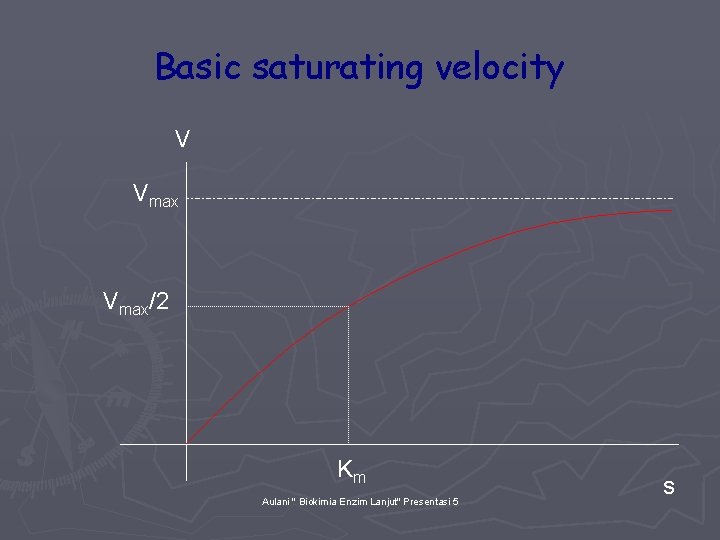
Basic saturating velocity V Vmax/2 Km Aulani " Biokimia Enzim Lanjut" Presentasi 5 s
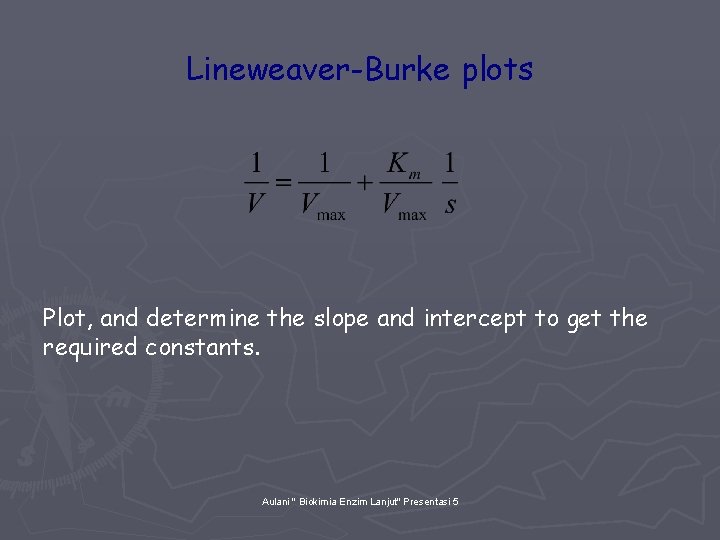
Lineweaver-Burke plots Plot, and determine the slope and intercept to get the required constants. Aulani " Biokimia Enzim Lanjut" Presentasi 5
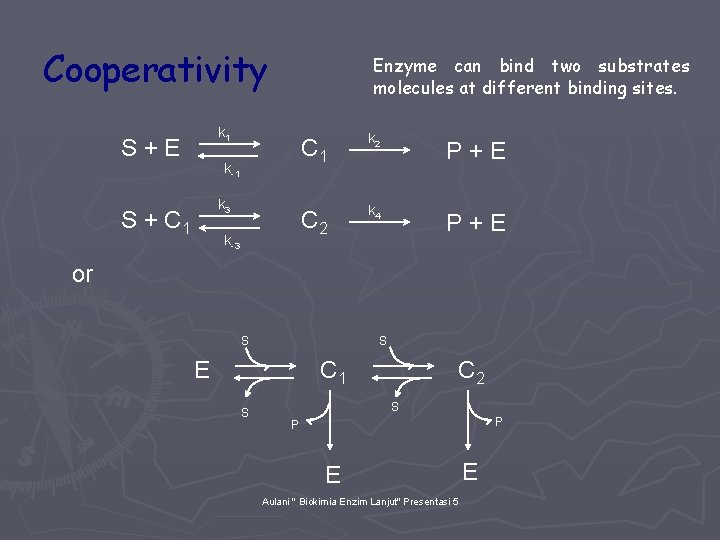
Cooperativity Enzyme can bind two substrates molecules at different binding sites. k 1 S+E k-1 k 3 S + C 1 k-3 C 1 k 2 P+E C 2 k 4 P+E or S S E C 1 S C 2 S P P E Aulani " Biokimia Enzim Lanjut" Presentasi 5 E

Pseudo-steady assumption Note the quadratic behaviour Aulani " Biokimia Enzim Lanjut" Presentasi 5

Independent binding sites S S 2 k+ E C 1 k. S k+ 2 k- C 2 S P P E E Just twice the single binding rate, as expected Aulani " Biokimia Enzim Lanjut" Presentasi 5
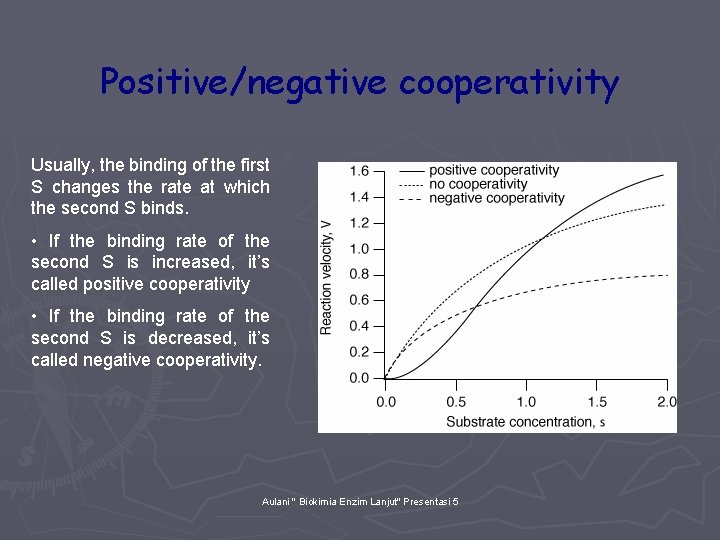
Positive/negative cooperativity Usually, the binding of the first S changes the rate at which the second S binds. • If the binding rate of the second S is increased, it’s called positive cooperativity • If the binding rate of the second S is decreased, it’s called negative cooperativity. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Hill equation In the limit as the binding of the second S becomes infinitely fast, we get a nice reduction. VERY special assumptions, note. This equation is used all the time to describe a cooperative reaction. Mostly use of this equation is just a heuristic kludge. Aulani " Biokimia Enzim Lanjut" Presentasi 5 Hill equation, with Hill coefficient of 2.

Another fast equilibrium model of cooperativity S E S k 1 C 1 k-1 S S k 2 P Let C=C 1+C 2 S+E E k 1 k-1 C f(s) k 3 P+E Aulani " Biokimia Enzim Lanjut" Presentasi 5 k-3 C 2 P k 4 E

Monod-Wyman-Changeux model A more mechanistic realisation of cooperativity. Aulani " Biokimia Enzim Lanjut" Presentasi 5

Equilibrium approximation Don’t even think about a pseudo-steady approach. Waste of valuable time. occupancy fraction and so on for all the other states which gives Note the sigmoidal character of this curve Aulani " Biokimia Enzim Lanjut" Presentasi 5

Reversible enzymes Of course, all enzymes HAVE to be reversible, so it’s naughty to put no back reaction from P to C. Should use S+E k 1 k-1 C k 2 k-2 I leave it as an exercise to calculate that Aulani " Biokimia Enzim Lanjut" Presentasi 5 P+E
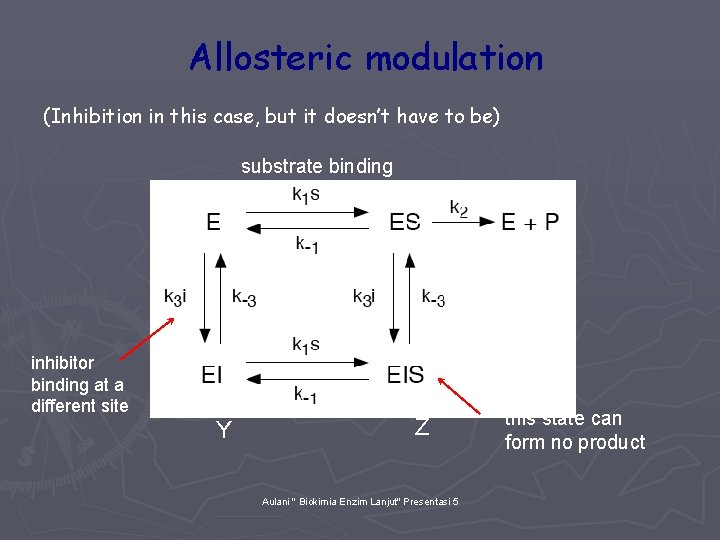
Allosteric modulation (Inhibition in this case, but it doesn’t have to be) substrate binding X inhibitor binding at a different site Y Z Aulani " Biokimia Enzim Lanjut" Presentasi 5 this state can form no product

Equilibrium approximation X Y Inhibition decreases the Vmax in this model Z Could change these rate constants, also. Aulani " Biokimia Enzim Lanjut" Presentasi 5
- Slides: 21