Basic Electron Microscopy Arthur Rowe The Knowledge Base

Basic Electron Microscopy Arthur Rowe The Knowledge Base at a Simple Level

Introduction These 3 presentations cover the fundamental theory of electron microscopy In presentation #3 we cover: – requirements for imaging macromolecules _ – – the negative staining method the metal-shadowing method _ – – aids such as gold-labelled antibodies Including high-resolution modifications vitritied ice technology examples of each type of method

requirements for imaging macromolecules • sufficient CONTRAST must be attainable, but > bio-molecules are made up of low A. N. atoms > & are of small dimensions (4+ nm) > hence contrast must usually be added • sufficient STABILITY in the beam is needed > to enable an image to be recorded > low dose ‘random’ imaging mandatory for any high resolution work

ways of imaging macromolecules • ADDING CONTRAST (with heavy metals) > negative contrast + computer analysis + immunogold labels > metal shadowing + computer enhancement • USING INTRINSIC CONTRAST > particles in thin film of vitrified ice + computer acquisition & processing
ways of imaging macromolecules • using immunogold labels to localise epitopes > widely used in cell biology > beginning to be of importance for macromolecules Au sphere Mab epitope macromolecule

negative staining particles Electron dense negative stain

negative staining • requires minimal interaction between particle & ‘stain’ • to avoid binding, heavy metal ion should be of same charge +/as the particle • positive staining usually destructive of bio-particles • biological material usually -ve charge at neutral p. H • widely used negative contrast media include: anionic cationic phosphotungstate uranyl actetate/formate molybdate (ammonium) (@ p. H ~ 4)

metal shadowing - 1 -directional

metal shadowing - 1 -directional • Contrast usually inverted to give dark shadows > resolution 2 - 3 nm - single 2 -fold a-helix detectable - historic use for surface detail - now replaced by SEM > detail on ‘shadow’ side of the particle can be lost > apparent ‘shape’ can be distorted > problems with orientation of elongated specimens - detail can be lost when direction of shadowing same as that of feature > very limited modern use for macromolecular work

metal shadowing - rotary

metal shadowing - rotary • Contrast usually inverted to give dark shadows > resolution 2 - 3 nm - single DNA strand detectable - historic use for ‘molecular biology’ (e. g. heteroduplex mapping) > good preservation of shape, but enlargement of apparent dimensions > in very recent modification (MCD - microcrystallite decoration), resolution ~1. 1 nm

particle in vitrified ice: low contrast particles examined at v. low temperature, frozen in a thin layer of vitrified (structureless) ice - i. e. no contrast added

particle in vitrified ice: low contrast average of large numbers (thousands +) of very low contrast particles enables a structure to be determined

particle in vitrified ice: low contrast average of large numbers (thousands +) of very low contrast particles enables a structure to be determined: • resolution may be typically 1 nm or better • this is enough to define the “outline” (or ‘envelope’) of a large structure • detailed high resolution data give us models for domains (or sub-domains) which can be ‘fitted into’ the envelope • ultimate resolution of the method ~0. 2 nm, rivalling XRC/NMR
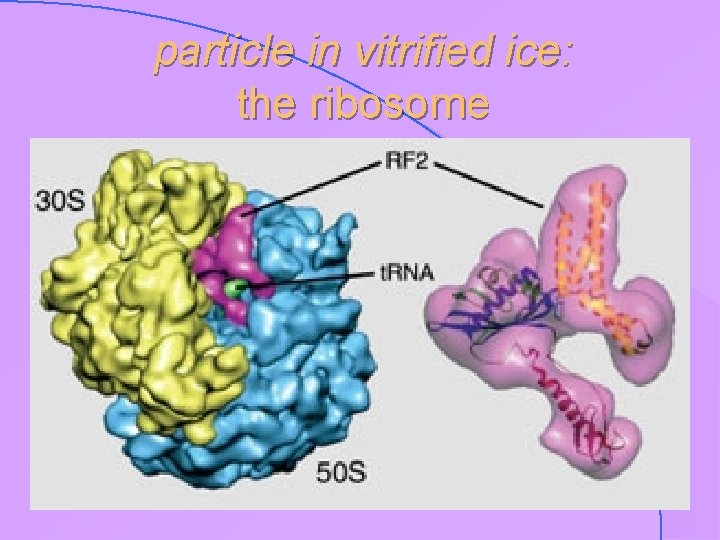
particle in vitrified ice: the ribosome
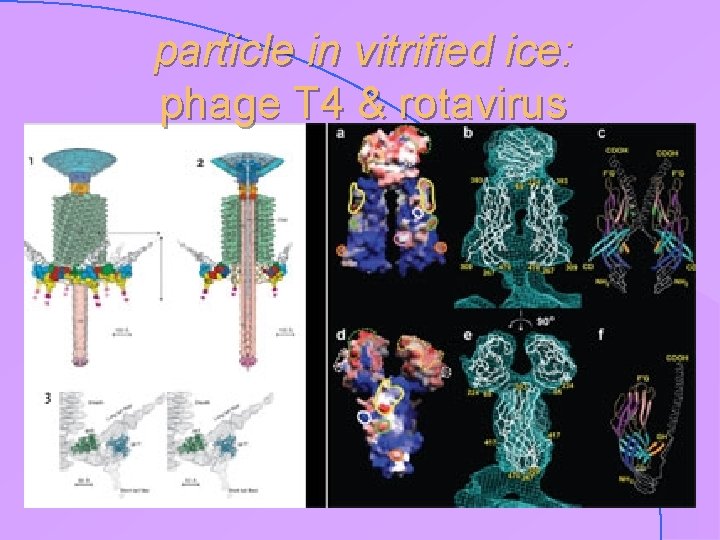
particle in vitrified ice: phage T 4 & rotavirus

case study : Gro. EL-Gro. ES • important chaperonins • hollow structure • appear to require ATP (hydrolysis ? ) for activity

particle in vitrified ice: low contrast the chaperonin protein Gro. EL visualised in vitrified ice (Helen Saibil & co-workers)
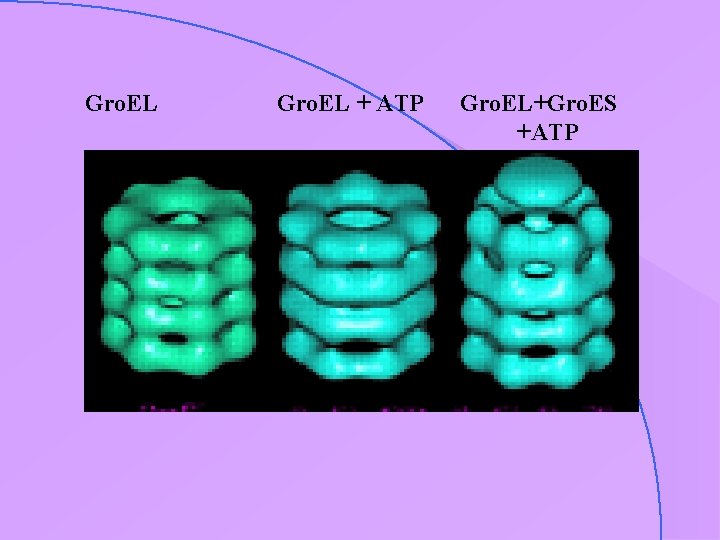
Gro. EL + ATP Gro. EL+Gro. ES +ATP

DLS as a probe for conformational change in Gro. EL/ES
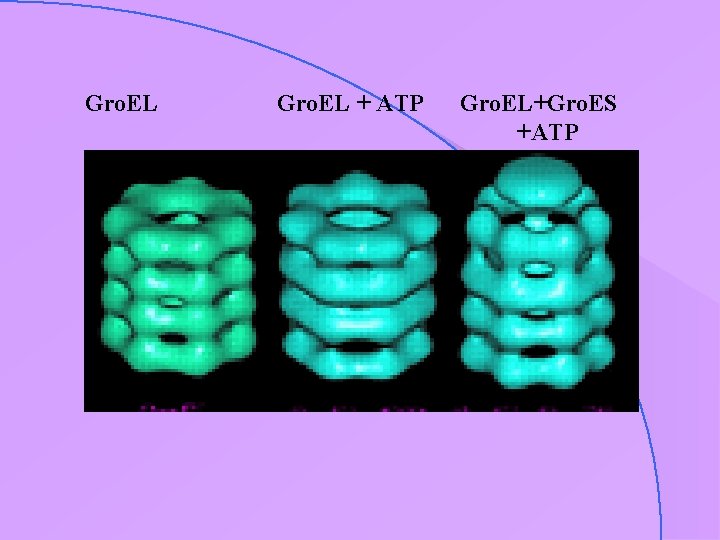
Gro. EL + ATP Gro. EL+Gro. ES +ATP


case study : pneumolysin • 53 k. D protein, toxin secreted from Pneumococcus pneumoniae • among other effects, damages membrane by forming pores • major causative agent of clinical symptoms in pneumonia
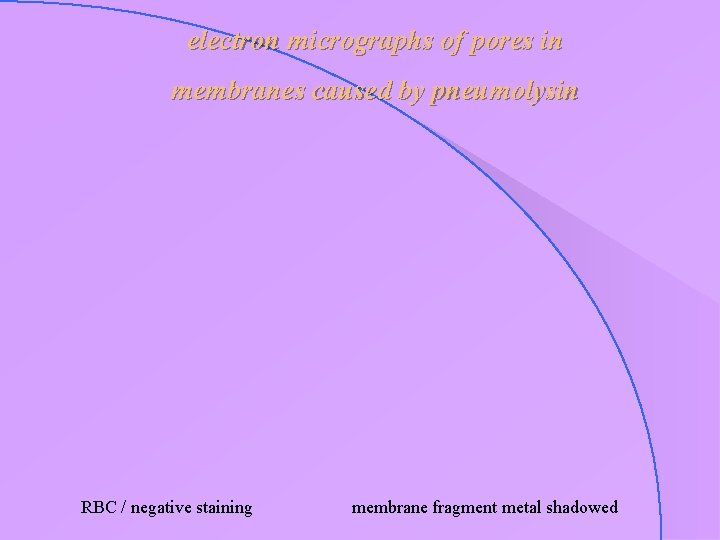
electron micrographs of pores in membranes caused by pneumolysin RBC / negative staining membrane fragment metal shadowed
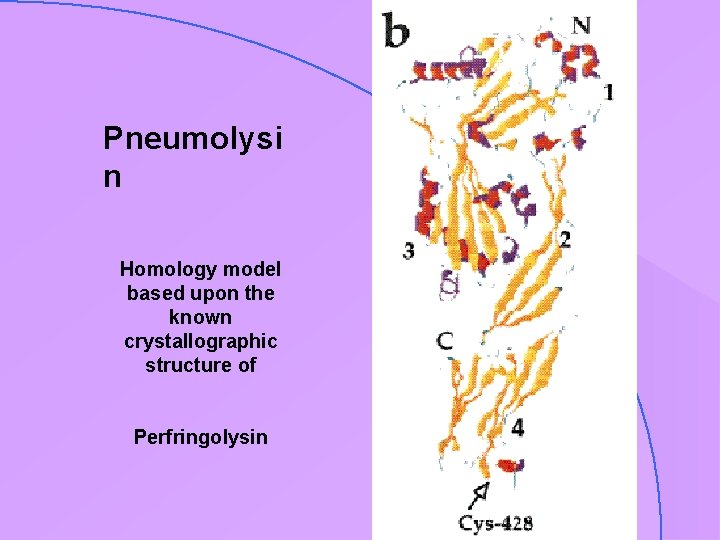
Pneumolysi n Homology model based upon the known crystallographic structure of Perfringolysin
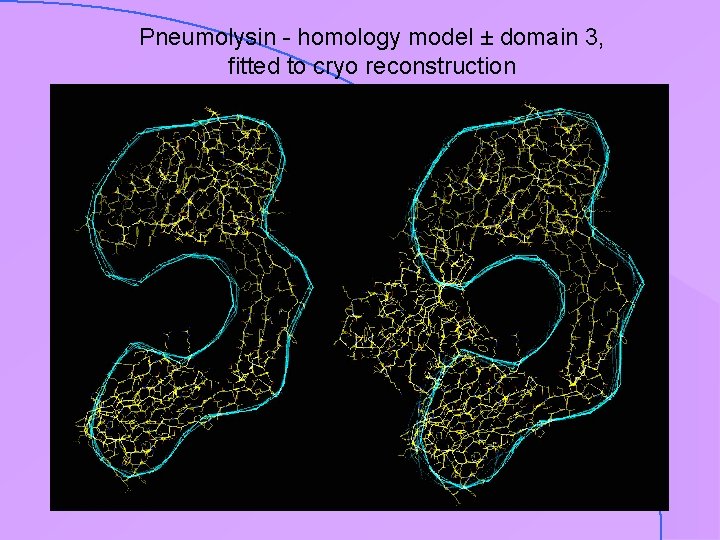
Pneumolysin - homology model ± domain 3, fitted to cryo reconstruction

Pneumolysin - EM by microcrystallite decoration (MCD) reveals orientation of domains
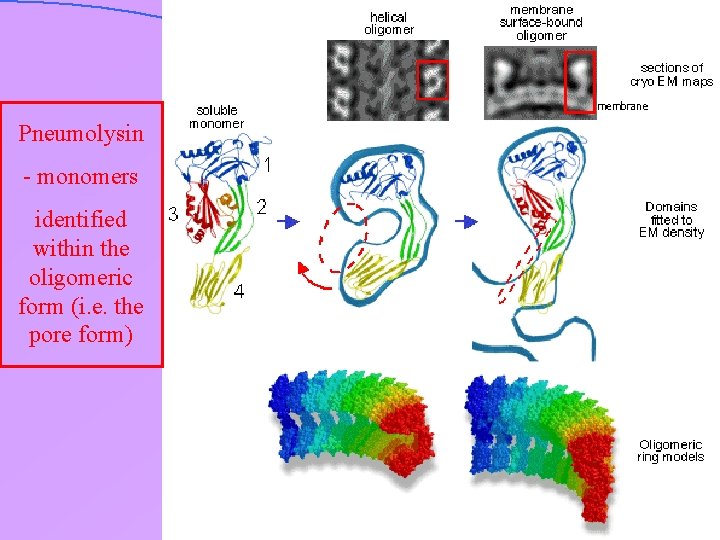
Pneumolysin - monomers identified within the oligomeric form (i. e. the pore form)

case study : myosin S 1 • motor domain of the skeletal muscle protein myosin • 2 S 1’s / myosin, mass c. 120 k. D • ‘cross-bridge’ between myosin and actin filaments, thought to be source of force generation

myosin is a 2 -stranded coiled-coil protein, with 2 globular (S 1) ‘heads’ S 1 unit

Each S 1 unit has a compact region, & a ‘lever arm’ connected via a ‘hinge’ to the main extended ‘tail’

Myosin S 1 imaged by Microcrystallite Decoration (no nucleotide present)

Effect of nucleotide (ADP) on the conformation of myosin S 1 as seen by MCD electron microscopy -ADP +ADP

case study : epitope localisation in an engineered vaccine • a new vaccine for Hepatitis B contains 3 antigens, S, S 1 & S 2, with epitopes on each • but does every particle of ‘hepagene’ contain all 3 of these epitopes ? • Mabs against S, S 1 & S 2 have been made & conjugated with gold: S 15 nm S 1 10 nm S 2 5 nm
immunolabelling of one epitope (S 1) in hepagene using 10 nm-Au labelled Mab

triple labelling of 3 epitopes on hepagene

Basic Electron Microscopy Arthur Rowe End
- Slides: 37