Basic concepts on population genetics OUTLINE Allele frequency

Basic concepts on population genetics OUTLINE Ø Allele frequency and genotypic frequency Ø Hardy Weinberg equilibrium Ø Linkage and Linkage disequilibrium (LD)

Allele and genotypic frequency under HW Characteristics of a population Genotypic frequencies Allele frequencies ‘n’ diploid plants are equivalent to ‘ 2 n’ alleles In F 1 and F 2 generations p and (1 -p)=0. 5 at all loci that differ between the two parental inbreeds. What is the Hardy-Weinberg Equilibrium?

HARDY-WEINBERG EQUILIBRIUM FOR ONE LOCUS OR SEVERAL LOCI CONSIDERED SEPARATELY 3 KEY FEATURE Ø Allele frequencies [p and q=(1 -p)] remain constant from generation to generation Ø Square of the array of allele=array of genotypic frequency Ø If allele frequency change, one generation of random mating would lead to a new set of equilibrium in genotypic frequency for all loci considered separately (one locus). How about for genotypes with respect to loci considered jointly?

Assumptions of HW model Ø Ø Ø DIPLOID SEXUAL REPRODUCTION TWO ALLELES MATING IS AT RANDOM LARGE POPULATION SIZE NO MUTATION, NO MIGRATION NATURAL SELECTION DOES NOT AFFECT THE ALLELES UNDER CONSIDERATIONS

Geneticists and breeders use procedures that cause deviation from HW equilibrium Ø Linkage and lack or random mating Ø Small population sizes Ø Assortative mating Ø Selection Ø Inbreeding INBREEDING and SMALL POPULATION SIZES affects all loci in the population SELECTION affect only certain loci
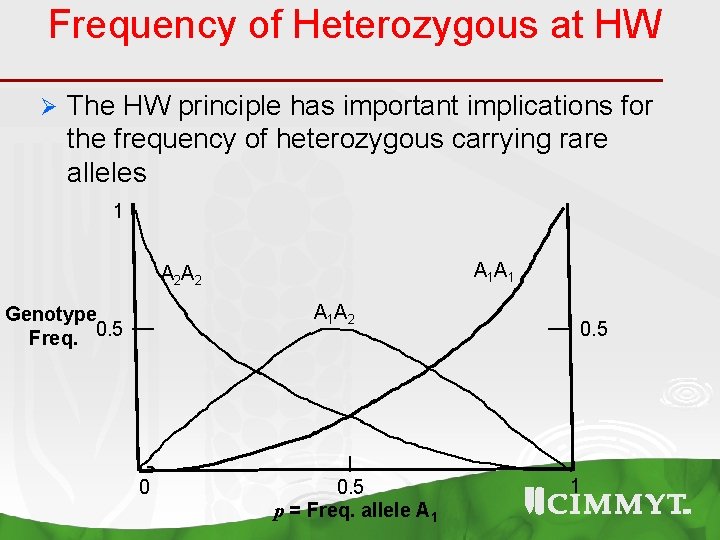
Frequency of Heterozygous at HW Ø The HW principle has important implications for the frequency of heterozygous carrying rare alleles 1 A 1 A 2 Genotype Freq. 0. 5 0 0. 5 p = Freq. allele A 1 0. 5 1
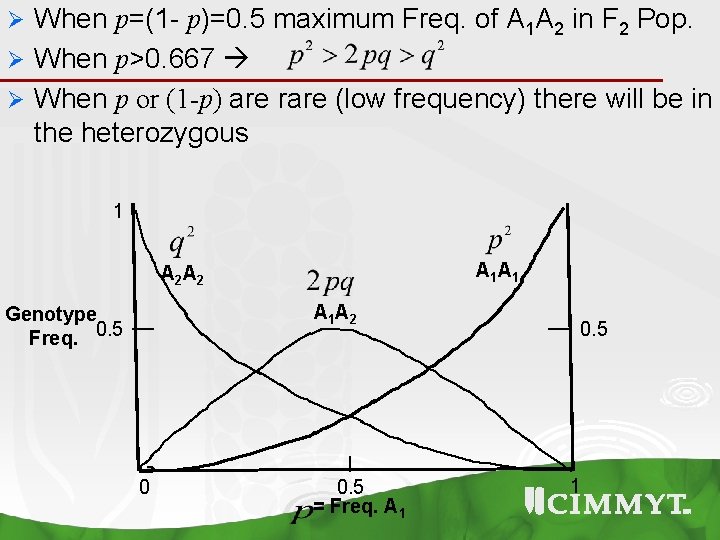
When p=(1 - p)=0. 5 maximum Freq. of A 1 A 2 in F 2 Pop. Ø When p>0. 667 Ø When p or (1 -p) are rare (low frequency) there will be in the heterozygous Ø 1 A 1 A 2 Genotype Freq. 0. 5 0 0. 5 = Freq. A 1 0. 5 1

HW for more than one locus Ø Two populations with two alleles at two loci. Equal number of individuals from both populations are mixed and mate at random A 1 A 1 B 1 B 1 x A 2 A 2 B 2 B 2 A 1 A 2 B 1 B 2 From 9 possible genotypes, in the first generation, there will be three types of genotypes A 1 A 1 B 1 B 1, A 1 A 2 B 1 B 2 , A 2 A 2 B 2 B 2 whereas all the other 6 possible genotype classes are ‘missings’

9 possible genotypes A 1 A 1 B 1 B 1 A 1 A 1 B 1 B 2 A 1 A 1 B 2 B 2 A 1 A 2 B 1 B 1 A 1 A 2 B 1 B 2 A 1 A 2 B 2 B 2 A 2 A 2 B 1 B 1 A 2 A 2 B 1 B 2 A 2 A 2 B 2 B 2

More than 1 generation of random mating Ø The missing genotypes would appear in subsequent generations. Ø If the two loci are linked then HW equilibrium will take longer because the appearance of the missing genotypes depends on the RECOMBINATION between the two loci. Ø Disequilibrium with respect to two or more loci is called LINKAGE DISEQUILIBRIUM this is regardless of whether the loci are linked or not
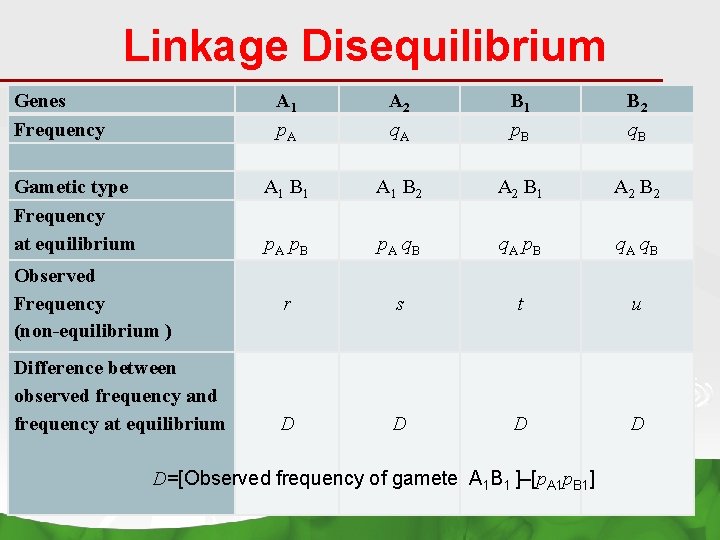
Linkage Disequilibrium Genes Frequency Gametic type Frequency at equilibrium A 1 p. A A 1 B 1 p. A p. B A 2 q. A A 1 B 2 p. A q. B B 1 p. B A 2 B 1 q. A p. B B 2 q. B A 2 B 2 q. A q. B Observed Frequency (non-equilibrium ) r s t u Difference between observed frequency and frequency at equilibrium D D D=[Observed frequency of gamete A 1 B 1 ]–[p. A 1 p. B 1]

When alleles at locus A are in random association with alleles in locus B? Ø LINKAGE A 1 B 2 ---A 2 B 1 ---- EQUILIBRIUM p. A 1 p. B 1 p. A 1 p. B 2 p. A 2 p. B 1 p. A 2 p. B 2 When alleles at locus A are NOT in random association with alleles in locus B? Ø LINKAGE DISEQUILIBRIUM D=[Observed frequency of gamete A 1 B 1 ]–[p. A 1 p. B 1] D=p. A 1 B 1 –[p. A 1 p. B 1]

Linkage Random association of alleles of different loci linkage equilibrium D=0 Ø Non Random association of alleles of different loci linkage disequilibrium D≠ 0 Ø For unlinked loci D decreases by ½ in each generation meiosis Ø For linked loci D decreases by c (recombination frequency) in each generation meiosis Ø Causes of linkage loci in the same chromosome Admixture of populations with different allele frequencies Natural selection differentially favoring some genotypes

Decay of LD in a population (in LD) mating at random Ø In this case the amount of disequilibrium is progressively reduced with successive generation of random mating. Ø The rate at which LD reduces depends on the frequency of gametic types in two successive generations. Ø Assume two loci are linked in the same chromosome. Ø The Disequilibrium D in the progeny can be obtained from the frequency of the four gametic types

Decay of LD in a population (in LD) mating at random Ø Consider only A 1 B 1 gametic type. This can be produces in two ways As non-recombinant (parental) A 1 B 1/Ax. Bx gametes In this case the frequency is r(1 -c) where r=freq. of A 1 B 1 in the parental gametes and c=recombination frequency 1. As recombinant A 1 Bx/Ax. B 1. The frequency of A 1 Bx chromosome is p. A and that of chromosome Ax. B 1 is p. B. In this case the frequency with which A 1 B 1 arises is p. A p. Bc 1. Therefore the frequency of A 1 B 1 in the progeny gametes is r’=r(1 -c)+ p. A p. Bc

Decay of LD in a population (in LD) mating at random The frequency of A 1 B 1 in the progeny gametes is r’=r(1 -c)+ p. A p. Bc and the disequilibrium in the progeny generation is D 1=r’-p. A p. B =r(1 -c)+ p. A p. Bc - p. A p. B = r(1 -c)- p. A p. B(1 -c) =(r-p. A p. B) (1 -c)=D 0(1 -c) For one more generation of random mating D 2=D 1(1 -c)=D 0(1 -c)2 For t more generation of random mating Dt=D 0(1 -c)t The loci do not have to be linked to be in LD. With unlinked loci c=1/2 and the amount of LD is ½ in each generation of random mating

Decay of LD in a population (in LD) mating at random Ø Ø Ø Ø Two concepts LINKAGE and LD. A population may be in LE but the loci remain linked by virtue of their being close together in the same chromosome Meiosis offer opportunities for crossover Selfing and random mating disrupt linkage Each generation of random mating reduces LD in a population Linkage affects the gametic output only if individuals are heterozygous at two or more loci. Close linkage does not automatically mean the loci are in LD Closely linked loci may be in LE and unlinked loci may in LD if, for example, two populations with different gene freq. at the loci are mixed Knowledge of LD can tell something about the breeding history of the population

The Haldane (1919) function Ø The Haldane function establishes that the recombination (c) frequency between a QTL and its closest marker can be expressed as where x is the map distance in centi. Morgans between the QTL and its closest marker

The Haldane (1919) function Ø From the Haldane function we get Ø This equation relates the recombination frequency (c) with the map distance (x) in centi. Morgan indicates that if c=1/2 the value of x is undefined Ø For this reason a QTL not linked to a marker must have a c close to 1/2 Ø This
- Slides: 19