Bacterial genome plasticity and integrons Didier Mazel Unit

Bacterial genome plasticity and integrons Didier Mazel Unité Plasticité du Génome Bactérien http: //www. pasteur. fr/recherche/unites/pgb/

- Introduction what we know How bacteria do evolve: » Mutations » Acquisitions (horizontal gene transfer) - Amplitude of the phenomenon - HGT Limits - Mechanisms of mobility - Mechanisms of gene capture – the exemple of the antibiotic resistance development and the integrons – superintegrons

Introduction We now know the intimity of bacterial genomes quite well in september 2005 • 257 bacterial genome had been sequenced (+24 Archeal genomes) • 521 were on going (+25 Archaea) http: //www. ncbi. nlm. nih. gov/genomes/lproks. cgi

• The sequenced bacterial genome* sizes vary from 0. 580 Mb (Mycoplasma genitalium) to ≈ 10. 0 Mb (cyanobacteria (Lyngbya) and actinobacteria (Streptomuyces-Rhodococcus) ) for comparison : S. cerevisiae 12 Mb • first rule: direct correlation between the adptive capabilities (“versatility”) and the genome size. e. g. intracellular parasite bacteria (constant and rich environnement) have small genomes. *, genome broad meaning : chromosome(s) + episomes


Plasticity ? large variation in size and content of bacterial genomes , - between different genera and species - but also among strains of the same species Large variation in the bacterial genome organisation, even between closely related species.

Variation in the bacterial genome organisation: 2 nd rule: Variations using 2 mechanisms: Transposition and homologous Recombinaison => repeated sequences play a major role, notably the mobile DNA elements.
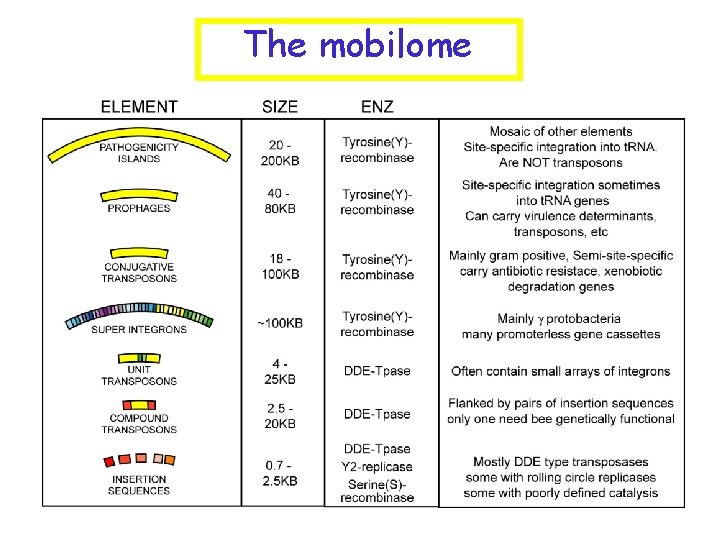
The mobilome
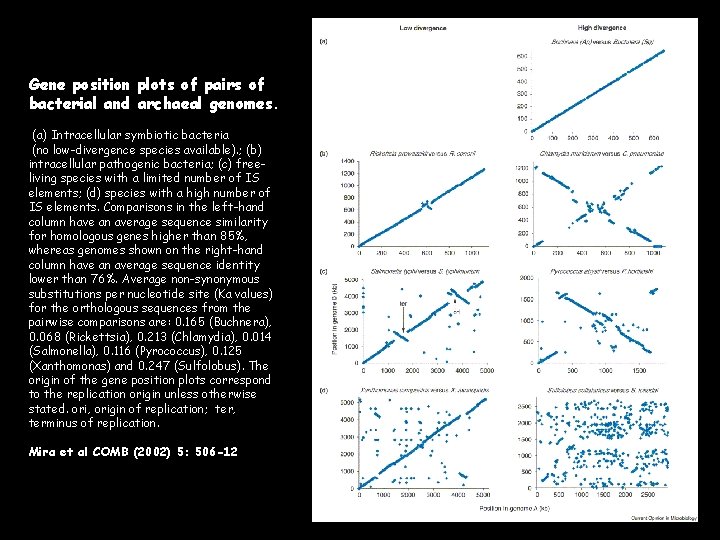
Gene position plots of pairs of bacterial and archaeal genomes. (a) Intracellular symbiotic bacteria (no low-divergence species available). ; (b) intracellular pathogenic bacteria; (c) freeliving species with a limited number of IS elements; (d) species with a high number of IS elements. Comparisons in the left-hand column have an average sequence similarity for homologous genes higher than 85%, whereas genomes shown on the right-hand column have an average sequence identity lower than 76%. Average non-synonymous substitutions per nucleotide site (Ka values) for the orthologous sequences from the pairwise comparisons are: 0. 165 (Buchnera), 0. 068 (Rickettsia), 0. 213 (Chlamydia), 0. 014 (Salmonella), 0. 116 (Pyrococcus), 0. 125 (Xanthomonas) and 0. 247 (Sulfolobus). The origin of the gene position plots correspond to the replication origin unless otherwise stated. ori, origin of replication; ter, terminus of replication. Mira et al COMB (2002) 5: 506 -12
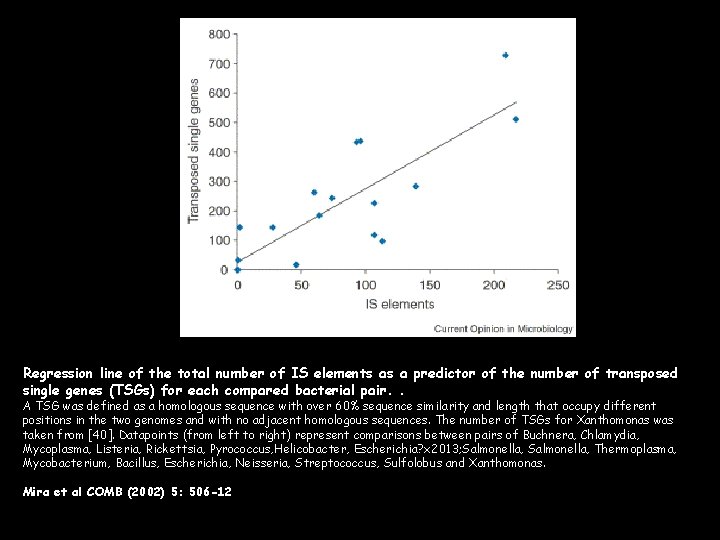
Regression line of the total number of IS elements as a predictor of the number of transposed single genes (TSGs) for each compared bacterial pair. . A TSG was defined as a homologous sequence with over 60% sequence similarity and length that occupy different positions in the two genomes and with no adjacent homologous sequences. The number of TSGs for Xanthomonas was taken from [40]. Datapoints (from left to right) represent comparisons between pairs of Buchnera, Chlamydia, Mycoplasma, Listeria, Rickettsia, Pyrococcus, Helicobacter, Escherichia? x 2013; Salmonella, Thermoplasma, Mycobacterium, Bacillus, Escherichia, Neisseria, Streptococcus, Sulfolobus and Xanthomonas. Mira et al COMB (2002) 5: 506 -12

Bacterial evolution routes • Mutations … • Duplication and divergent evolution (paralogues) (Rares ! in général paralogues viennent d’orthologues différents) • Horizontal gene transfer (up to 15% of the genome) The three kind of phenomenon co-exist and act in synergy. Therefore it is difficult to measure their respective contribution to complex phenotypes development => a well known exemple : the antibiotic resistance

Horizontal gene transfer signatures: • Comparative Analysis = > differences • Mobile elements characteristic boundaries: insertions at highly conserved loci (t. RNAs, …) presence of repeated sequences in direct or inverted orientation, transposase or recombinase genes • base composition or codon usage deviations

Limits of the exchanges : none ? ? ? • based on sequence similarity: Eucaryotes bactéries • Exprimentaly : idem ! archae • Natural systems: Ti DNA Agrobacterium -> plants migration of the organites genes in the nucleus The limit is more a question of sharing the same ecological niche

Mobility mechanims • Transformation: Limitée à un certain nombre d’espèces intégration par recombinaison homologue • Transduction: Phages très abondants, spectre d’hôtes restreint // PAI de type prophage • Conjugaison: Spectre d’hôte variable, transfert de réplicon ou machinerie d’intégration par recombinaison spécifique de site (affranchi limite)

an example : the antibiotics resistance • recent phenomenon: 60 years • Extremely documented • 80 % of resistances due to exogenous gene acquisitions
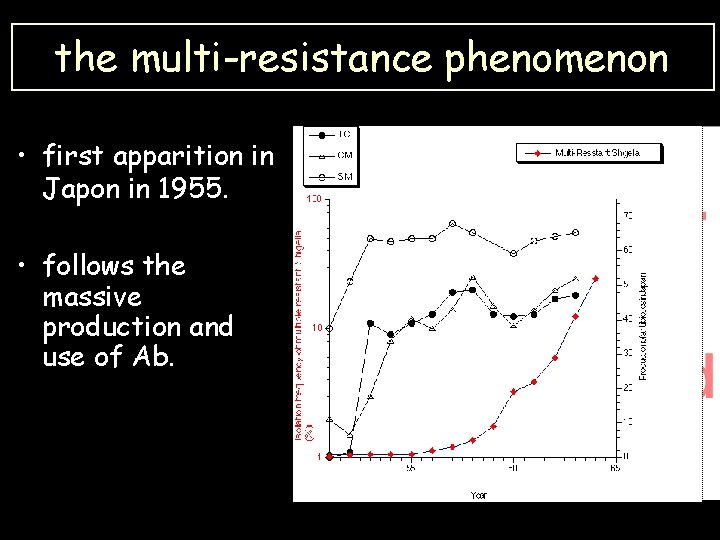
the multi-resistance phenomenon • first apparition in Japon in 1955. • follows the massive production and use of Ab.

R Ab genes origin • in most cases : ? ? • bacterial producers have had to develop protection mechanisms => potential source
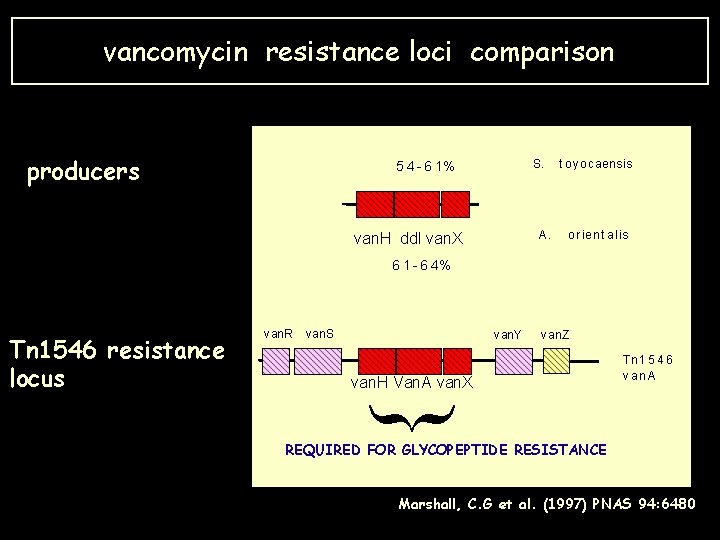
vancomycin resistance loci comparison producers 5 4 - 6 1% van. H ddl van. X S. t oy ocaensis A. o r ien t alis 6 1 - 6 4% van. Y van. Z van. H Van. A van. X Tn 1 5 4 6 v an A { Tn 1546 resistance locus van. R van. S REQUIRED FOR GLYCOPEPTIDE RESISTANCE Marshall, C. G et al. (1997) PNAS 94: 6480
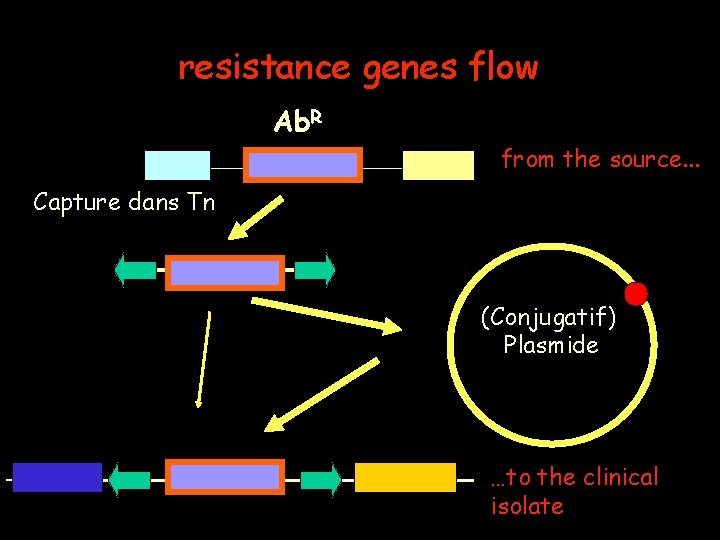
resistance genes flow Ab. R from the source. . . Capture dans Tn (Conjugatif) Plasmide …to the clinical isolate

A crucial question: how do Transposons acquièrent-ils ces gènes ? the answer has been obtained for two examples: • Composite. Transposons • Transposons carrying an integron
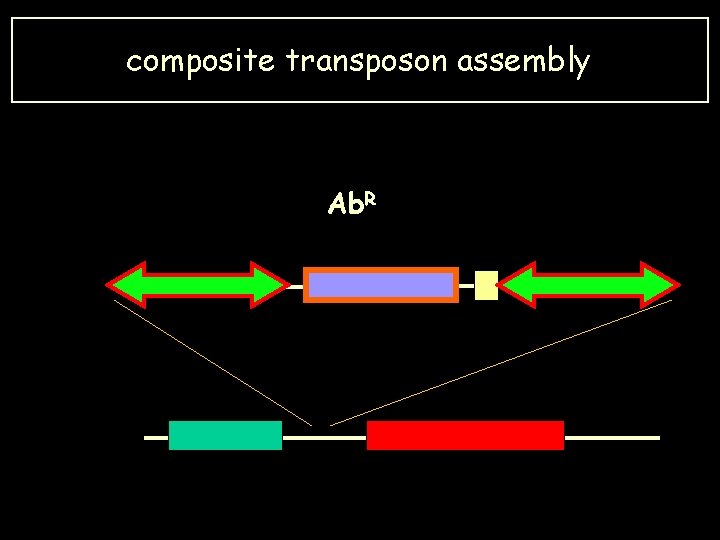
composite transposon assembly Ab. R

The integrons • They constitute what can be defined as a natural genetic engineering system: • Incorporate ORFs • Express them
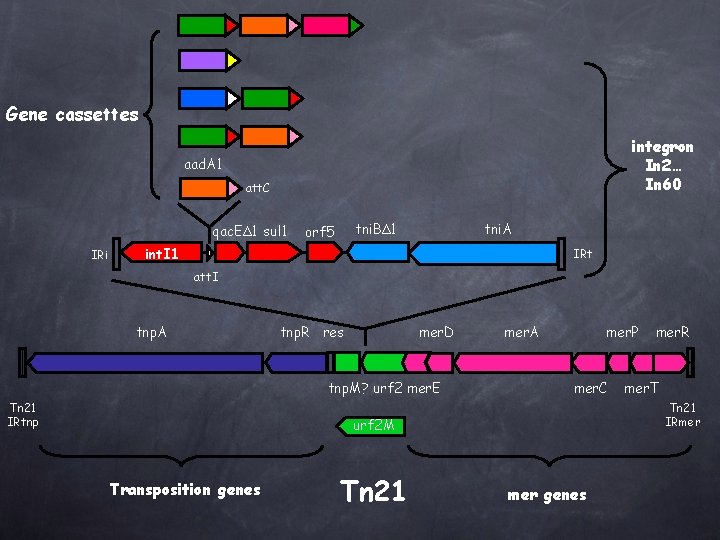
Gene cassettes integron In 2… In 60 aad. A 1 att. C qac. EΔ 1 sul 1 IRi tni. BΔ 1 orf 5 tni. A int. I 1 IRt att. I tnp. A tnp. R res mer. D tnp. M? urf 2 mer. E Tn 21 IRtnp mer. A mer. P mer. C Tn 21 mer. T Tn 21 IRmer urf 2 M Transposition genes mer. R mer genes

More than 85 different gene cassettes encoding antibiotic resistance have been found in integrons. These cassettes allow to resist to all classes of antibiotics used against human Gram-negative pathogens (β-lactams, aminoglycosides, chloramphenicol, trimethoprim, streptothricin, rifampin, erythromycin, …antiseptics).
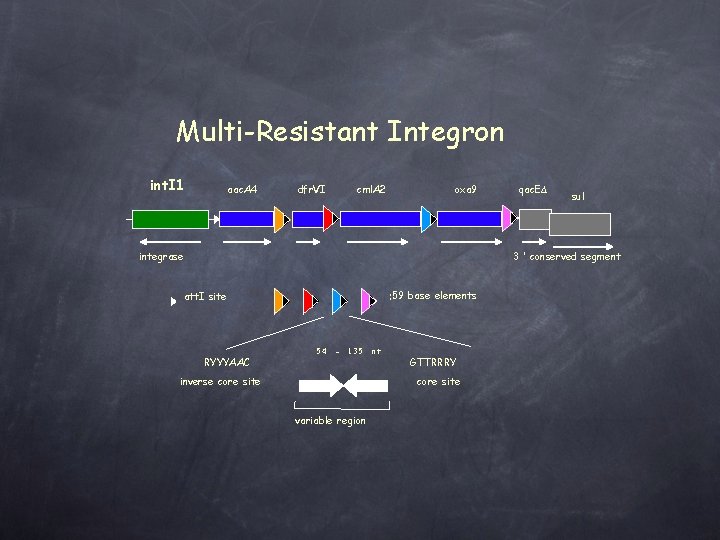
Multi-Resistant Integron int. I 1 aac. A 4 dfr. VI cml. A 2 oxa 9 integrase qac. EΔ sul 3 ’ conserved segment : 59 base elements att. I site RYYYAAC 54 - 135 inverse core site nt GTTRRRY core site variable region
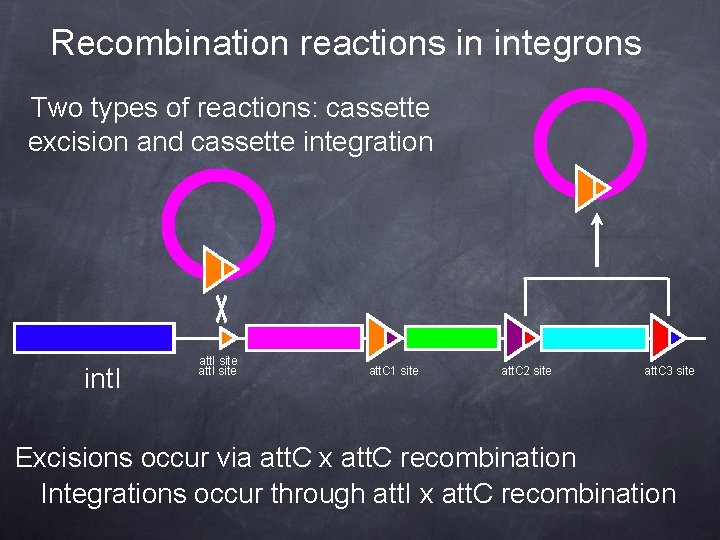
Recombination reactions in integrons Two types of reactions: cassette excision and cassette integration int. I att. I site att. C 1 site att. C 2 site att. C 3 site Excisions occur via att. C x att. C recombination Integrations occur through att. I x att. C recombination
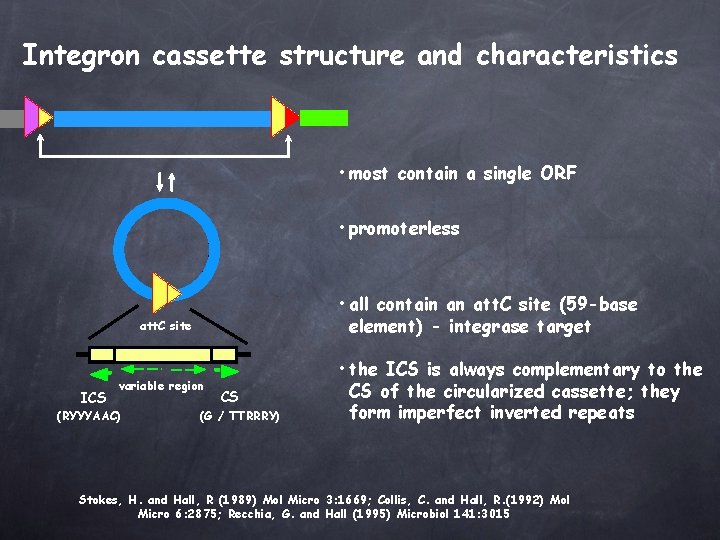
Integron cassette structure and characteristics • most contain a single ORF • promoterless • all contain an att. C site (59 -base element) - integrase target att. C site ICS variable region (RYYYAAC) CS (G / TTRRRY) • the ICS is always complementary to the CS of the circularized cassette; they form imperfect inverted repeats Stokes, H. and Hall, R (1989) Mol Micro 3: 1669; Collis, C. and Hall, R. (1992) Mol Micro 6: 2875; Recchia, G. and Hall (1995) Microbiol 141: 3015

The Mobile Integrons and the antibiotic Resistance : Five “classes” of Integrons (MIs) • Share between 45 -58% amino acid identity • All are associated with mobile DNA elements • Carry at most 8 resistance cassettes class 1 class 2* class 3 class 4 class 5 Tn 21 family (most ubiquitous) Tn 7 family self-transmissible plasmid SXT element (constin)

What is the origin of the multiresistance integrons and their cassettes ?
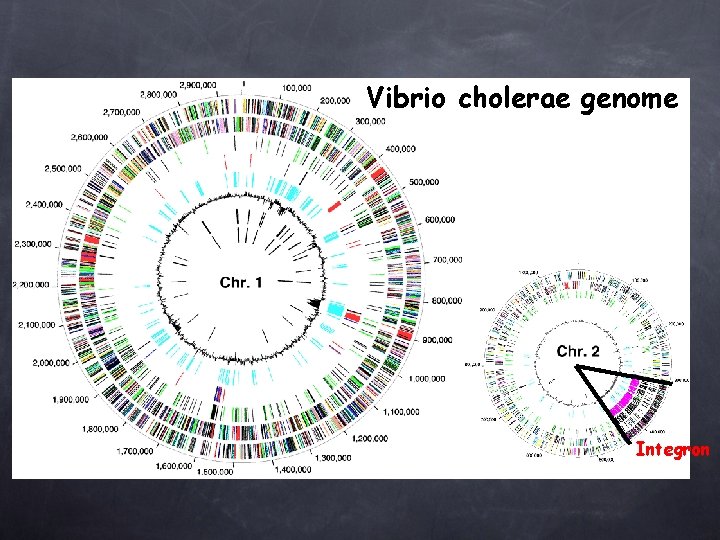
Vibrio cholerae genome Integron
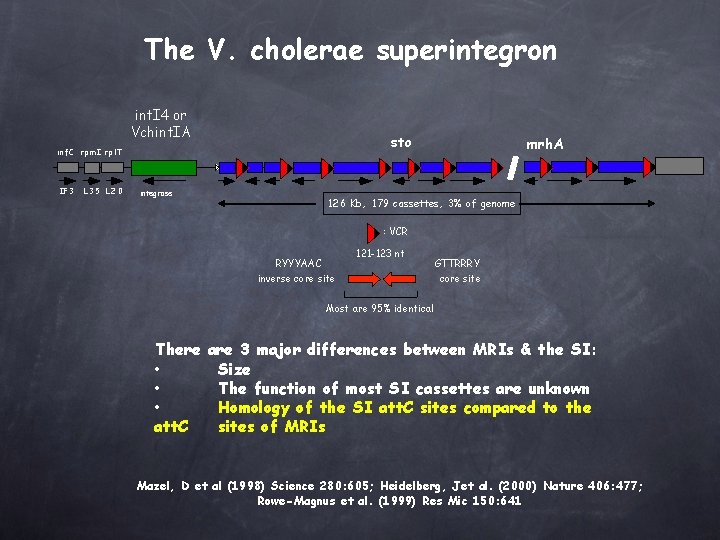
The V. cholerae superintegron int. I 4 or Vchint. IA sto i n f. C rp m I rpl. T IF 3 L 35 L 20 integrase mrh. A 126 Kb, 179 cassettes, 3% of genome : VCR RYYYAAC inverse core site 121 -123 nt GTTRRRY core site Most are 95% identical There are 3 major differences between MRIs & the SI: • Size • The function of most SI cassettes are unknown • Homology of the SI att. C sites compared to the att. C sites of MRIs Mazel, D et al (1998) Science 280: 605; Heidelberg, Jet al. (2000) Nature 406: 477; Rowe-Magnus et al. (1999) Res Mic 150: 641
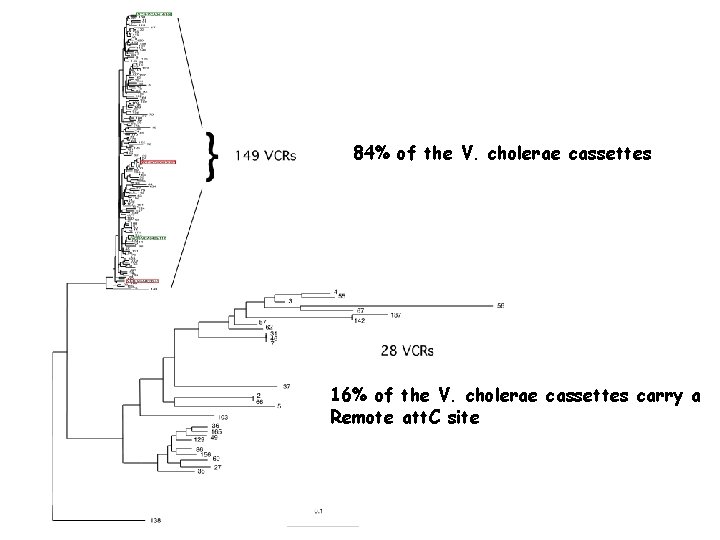
84% of the V. cholerae cassettes 16% of the V. cholerae cassettes carry a Remote att. C site
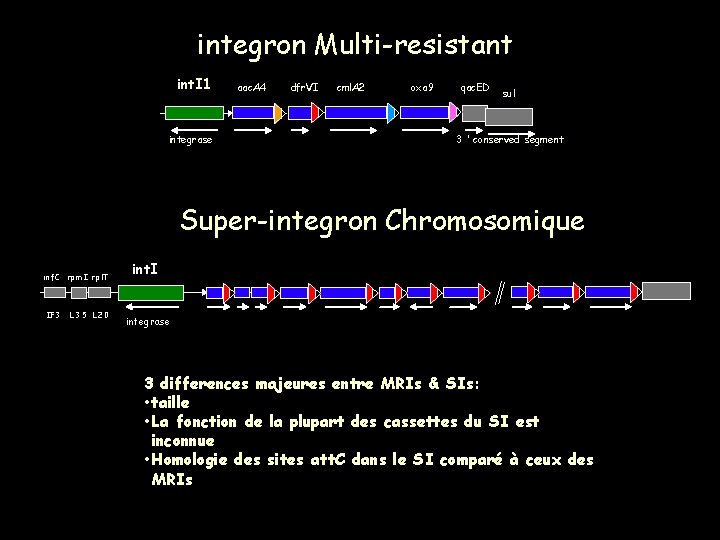
integron Multi-resistant int. I 1 integrase aac. A 4 dfr. VI cml. A 2 oxa 9 qac. ED sul 3 ’ conserved segment Super-integron Chromosomique in f. C rp m I rp l. T IF 3 L 35 L 20 int. I integrase 3 differences majeures entre MRIs & SIs: • taille • La fonction de la plupart des cassettes du SI est inconnue • Homologie des sites att. C dans le SI comparé à ceux des MRIs

Are SIs the source of MRIs?
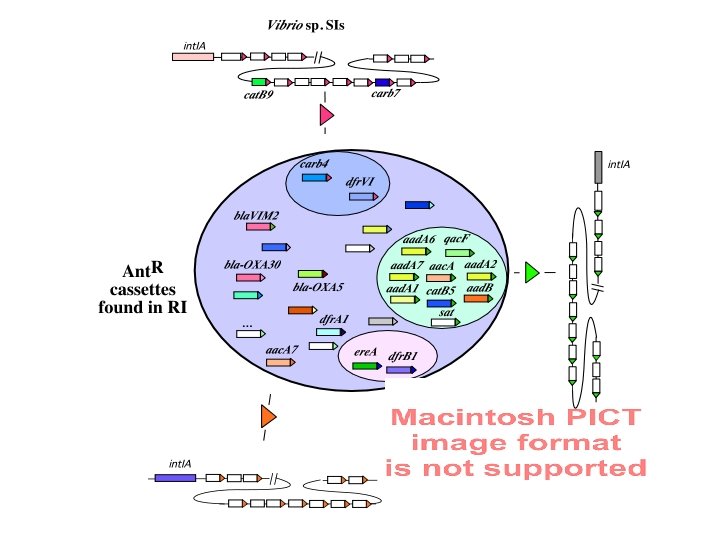

SIs are widespread among the proteobacteria => We characterized SIs / Is structures in >30 proteobacterial species (mainly γ ) Vibrio Listonella Alteromonas Xanthomonas Pseudomonas Shewanella Treponema alcaligenes * mendocina * pseudoalcaligenes oneidensis MR-1 putrefaciens denticola (spirochete) Microbulbifer Nitrosomonas europaea (β) Thiobacillus ferrooxidans (β) Geobacter sulfurreducens (δ ) Photobacterium Moritella + 2 new types of MRIs + 19 int. Is and hundreds of cassettes from soil extracted DNA
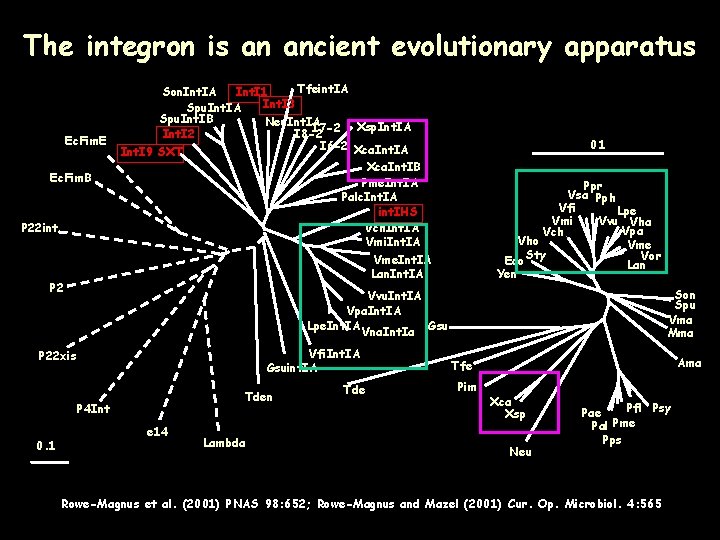
The integron is an ancient evolutionary apparatus Ec. Fim. E Ec. Fim. B P 22 int P 2 Tfeint. IA Son. Int. IA Int. I 1 Int. I 3 Spu. Int. IA Spu. Int. IB Neu. Int. IA I 7 -2 Xsp. Int. IA Int. I 2 I 8 -2 I 6 -2 Xca. Int. IA Int. I 9 SXT Xca. Int. IB Pme. Int. IA Palc. Int. IA int. IHS Vch. Int. IA Vmi. Int. IA Vme. Int. IA Lan. Int. IA Ppr Vsa Pph Vfi Lpe Vvu Vha Vmi Vpa Vch Vho Vme Sty Vor Eco Lan Yen Son Spu Vma Mma Vvu. Int. IA Vpa. Int. IA Lpe. Int. IA Vna. Int. Ia Gsu Vfi. Int. IA Gsuint. IA P 22 xis Tden P 4 Int 0. 1 e 14 Lambda Tde Ama Tfe Pim Xca Xsp Neu Pfl Psy Pae Pme Pal Pps Rowe-Magnus et al. (2001) PNAS 98: 652; Rowe-Magnus and Mazel (2001) Cur. Op. Microbiol. 4: 565
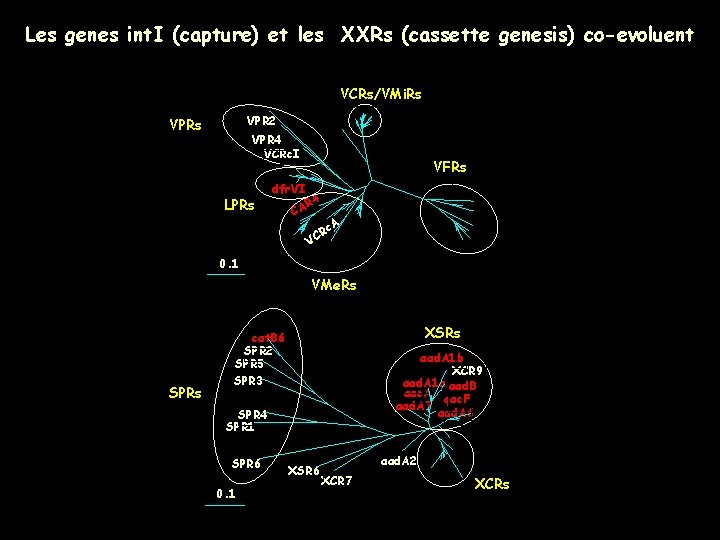
. Les genes int. I (capture) et les XXRs (cassette genesis) co-evoluent VCRs/VMi. Rs VPR 2 VPRs VPR 4 VCRc. I LPRs dfr. VI VFRs R 4 CA R VC c. A 0. 1 VMe. Rs SPRs XSRs cat. B 6 SPR 2 SPR 5 SPR 3 aad. A 1 b XCR 9 aad. A 1 a aad. B aac. A qac. F aad. A 7 aad. A 6 SPR 4 SPR 1 SPR 6 0. 1 XSR 6 aad. A 2 XCR 7 XCRs

Altogether: • • • SI functional platforms (int. I + att. I) are not mobile and co-evolve with host genomes Integrases and att. C sites co-evolve The Vibrio and the Xanthomonas ancestors carried a SI (evolutionary history of > 300 Myrs).

The Gene Cassette Reservoir To what extent do the different SIs share cassettes? What kind of functions are found in the cassettes?
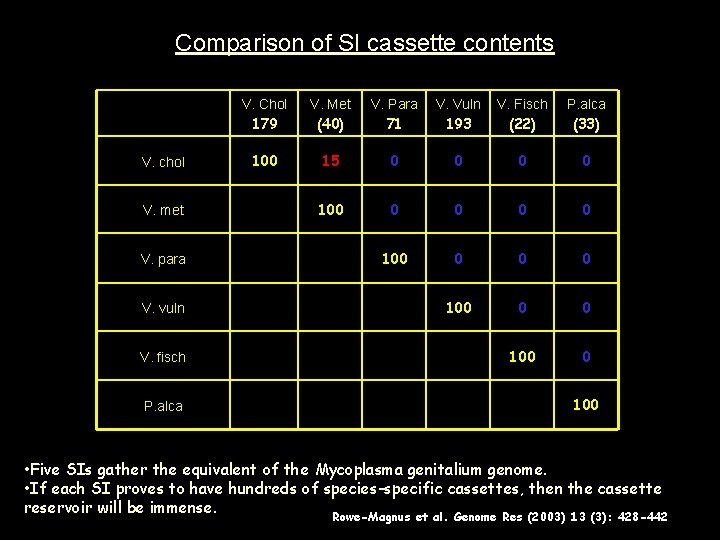
Comparison of SI cassette contents V. chol V. met V. para V. vuln V. fisch P. alca V. Chol 179 V. Met (40) V. Para 71 V. Vuln 193 V. Fisch (22) P. alca (33) 100 15 0 0 0 0 100 0 100 • Five SIs gather the equivalent of the Mycoplasma genitalium genome. • If each SI proves to have hundreds of species-specific cassettes, then the cassette reservoir will be immense. Rowe-Magnus et al. Genome Res (2003) 13 (3): 428 -442
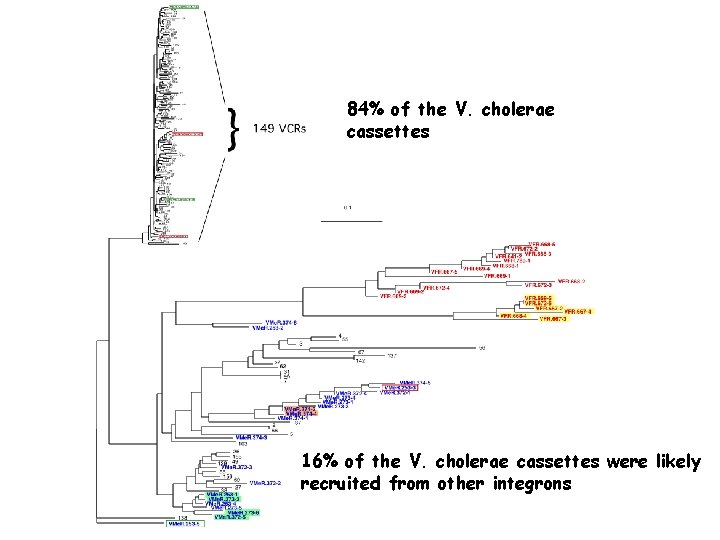
84% of the V. cholerae cassettes 16% of the V. cholerae cassettes were likely recruited from other integrons
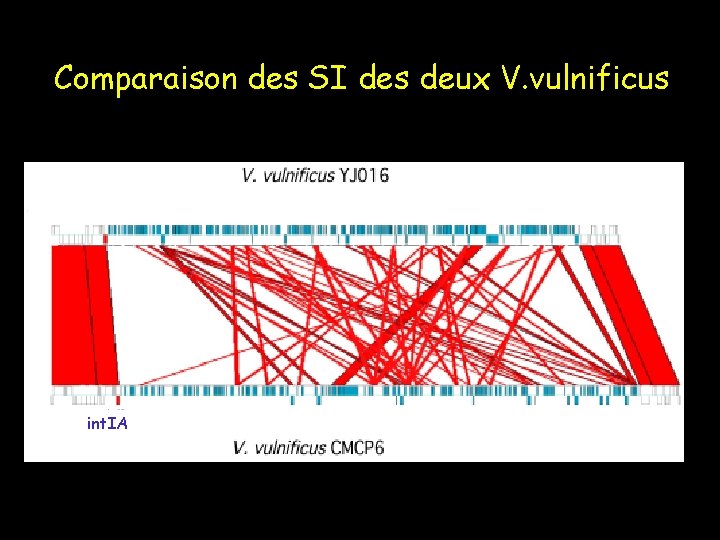
Comparaison des SI des deux V. vulnificus int. IA

Partial inventory of the functions encoded in SI cassettes Various demonstrated adaptive functions : Pathogenicity, Metabolic activities, DNA repair, But also homologues to several Ab. R genes (aminoglycoside, chloramphenicol, fosfomycin, phosphinothricine, streptothricine, microcin immunity, . . . ) … and many plasmid addiction modules
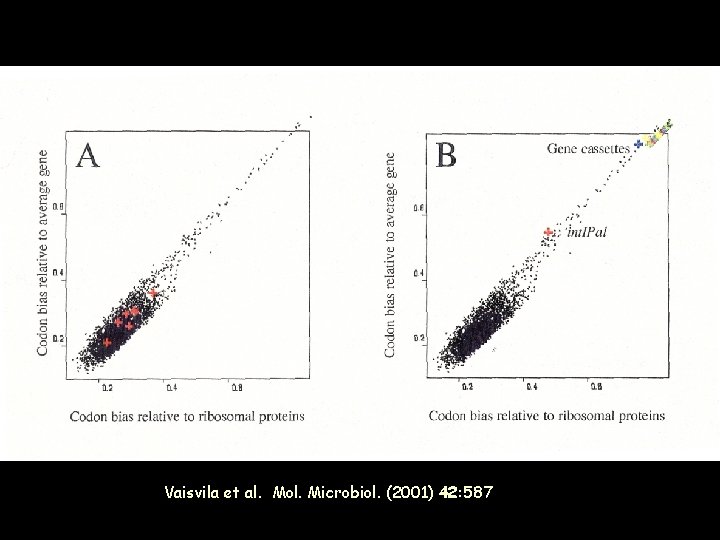
Vaisvila et al. Mol. Microbiol. (2001) 42: 587

Paradoxically, despite the fact that most of the hundreds of cassettes have NO PROMOTER, the SI cassette content is extremely stable ! How does the selection apply to maintain such large arrays of mobile elements (the cassettes), which, in addition, are silent in most cases ? What can explain this ?

Mobile elements, repeated sequences and bacterial genome stability • Repeated sequences play a primordial role in the overall genome plasticity, especially through recombination. • It is postulated, and observed, that “useless” or somehow deleterious genes are lost from bacterial genomes (reduction). • It is then considered that without a positive selective pressure, a gene will be lost quite rapidly, even faster if this gene is “mobile”.
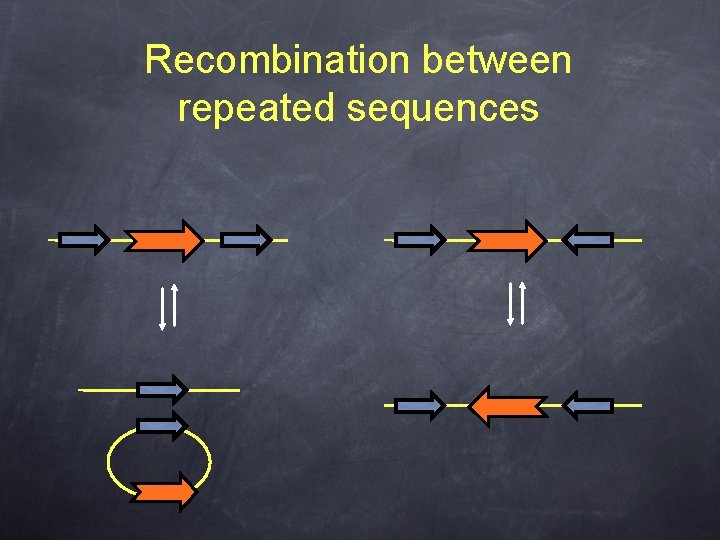
Recombination between repeated sequences
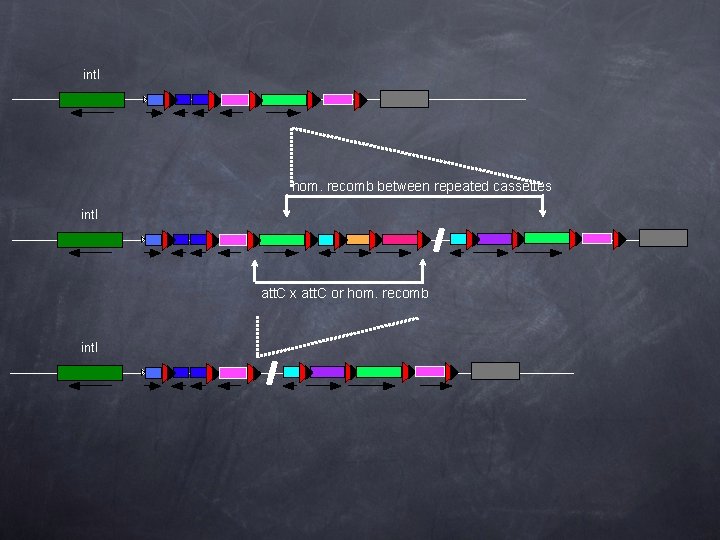
int. I hom. recomb between repeated cassettes int. I att. C x att. C or hom. recomb int. I

How does the selection apply to maintain such large arrays of mobile elements (the cassettes), which, in addition, are silent in most cases ? What can explain this ? 1 - Does this apparent stability reflect the absence of integrase expression?

Yes ! The int. I genes, even the one of class 1 RI, are completely silent in laboratory conditions ! The integrase expression is tightly controlled (However, the catalytic properties of the SI integrases are identical to those of the MRI integrases)

2. The addiction cassettes: an explanation for the SI cassette arrays stability ? Paradox: • each of the characterized SI carries a few repeated cassettes (up to 24 X in cholerae) • Most cassettes are silent How are such structures maintained? How are deletions counter selected ?
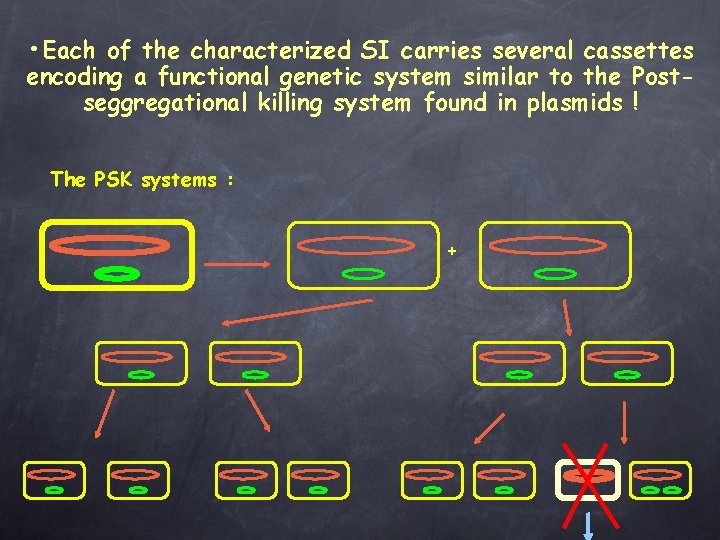
• Each of the characterized SI carries several cassettes encoding a functional genetic system similar to the Postseggregational killing system found in plasmids ! The PSK systems : +
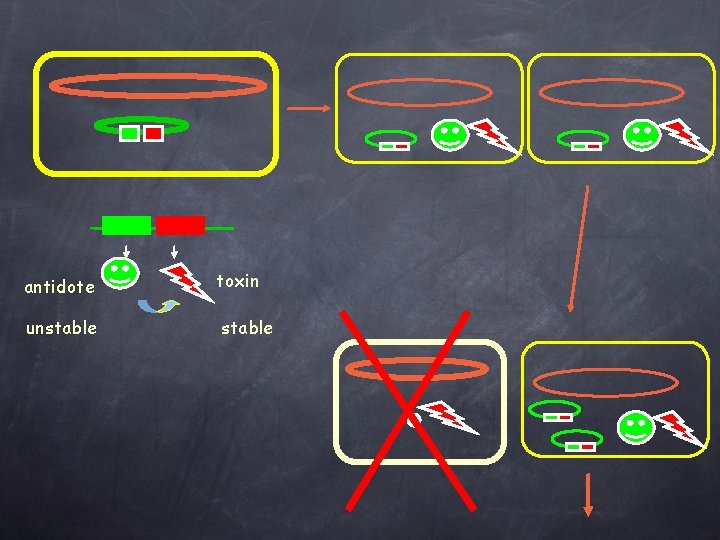
antidote toxin unstable

Each of the characterized SI carries at least one cassette encoding a functional genetic system similar to the Post-seggregational killing system found in plasmids : • ccd. AB in V. fischeri // F plasmid • Phd-doc in cholerae and metschnikovii // P 1 phage • hig. AB in cholerae • par. DE in cholerae • rel. BE in parahaemolyticus, vulnificus, cholerae • Xba. I RE and methylase in X. campestris badrii • RE in Pseudomonas, V. metschnikovii, , V. vulnificus functional 5 different types found in the V. cholerae SI, giving a total of at least 14 psk cassettes! => Recombination between repeated cassettes is counter selected and the concomitant loss of large part of the tandem cassette array is avoided.

How adaptative are SIs ? . . . but : – A great majority of cassettes bear ORF of unknown function – The only known promoter (inside Int. I gene) cannot account for expression of the whole set of cassette – Integration preferently occur at att. I (integrase) site, while excision occur between two att. C (cassette) site
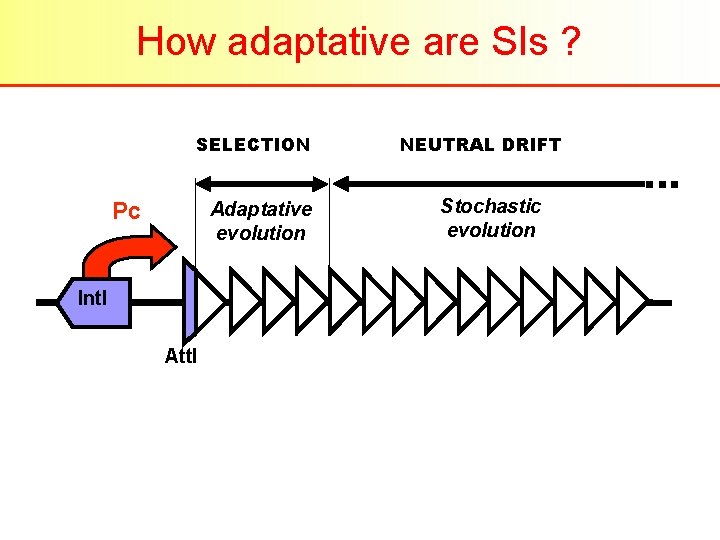
How adaptative are SIs ? SELECTION Pc Adaptative evolution Int. I Att. I NEUTRAL DRIFT Stochastic evolution
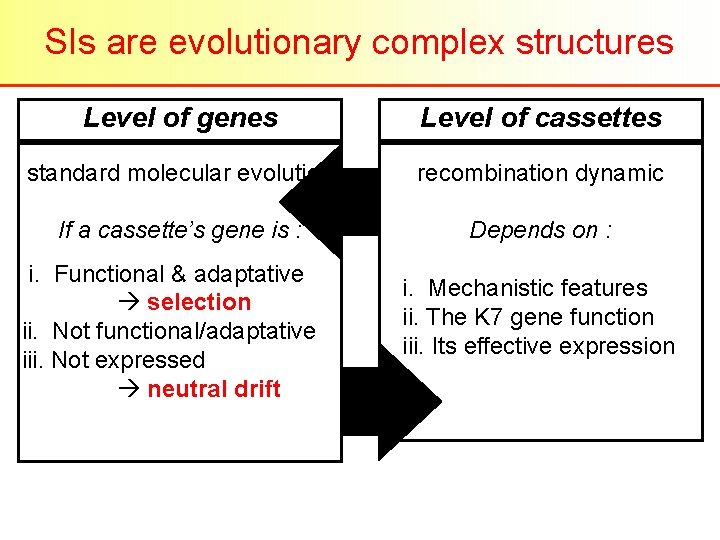
SIs are evolutionary complex structures Level of genes Level of cassettes standard molecular evolution recombination dynamic If a cassette’s gene is : Depends on : i. Functional & adaptative selection ii. Not functional/adaptative iii. Not expressed neutral drift i. Mechanistic features ii. The K 7 gene function iii. Its effective expression

Ka / Ks : A comparative method for detecting selection For pairs of sufficiently diverse sequences : • Ka / Ks = 1 : Neutrality • Ka / Ks >> 1 : More non-syn. mutations Diversifying selection • Ka / Ks << 1 : more syn. mutations Purifying selection

Ka / Ks : Application to Vibrios SIs’ cassettes • Cassettes extraction : “IRMA” > ~ 1500 cassettes • Pairing : local blast on the cassettes bank > ~ 200 groups • Alignment and test for selection

Ka / Ks analysis : Negative selection • 25 % of the groups show strong features of purifying selection functionality • Physical link between adaptative cassettes ? promoter identification. . . • Lots of TA system • Proteins involved in stress response
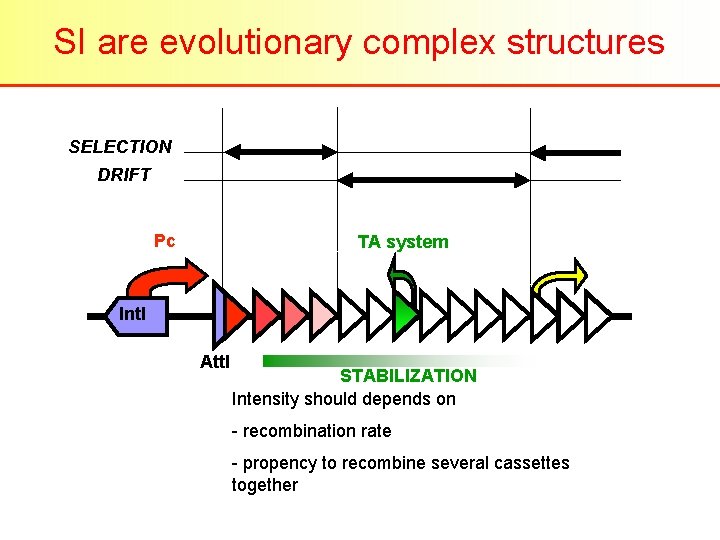
SI are evolutionary complex structures SELECTION DRIFT Pc TA system Int. I Att. I STABILIZATION Intensity should depends on - recombination rate - propency to recombine several cassettes together

The genesis of a MRI
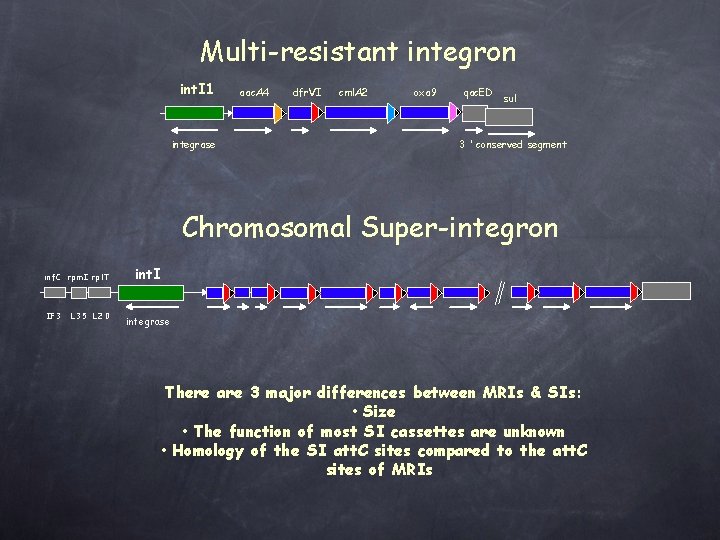
Multi-resistant integron int. I 1 integrase aac. A 4 dfr. VI cml. A 2 oxa 9 qac. ED sul 3 ’ conserved segment Chromosomal Super-integron i n f. C rp m I rpl. T IF 3 L 35 L 20 int. I integrase There are 3 major differences between MRIs & SIs: • Size • The function of most SI cassettes are unknown • Homology of the SI att. C sites compared to the att. C sites of MRIs
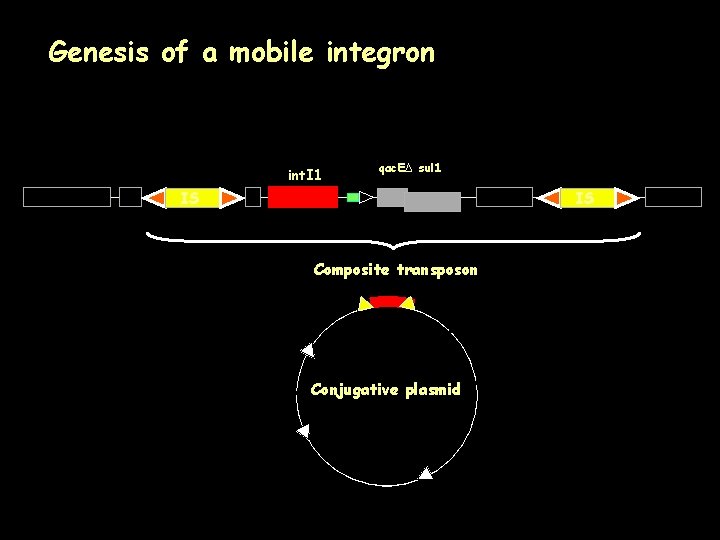
Genesis of a mobile integron int. I 1 qac. EΔ sul 1 IS IS Composite transposon Conjugative plasmid IS
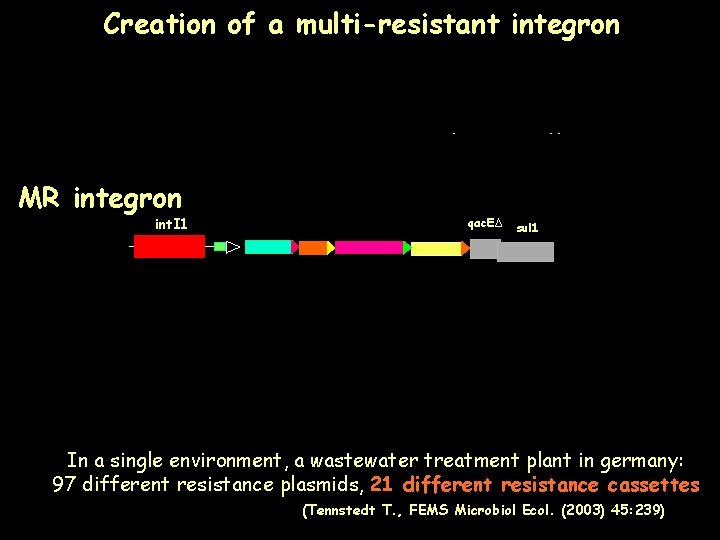
Creation of a multi-resistant integron SI Vibrio sp SIShewanellasp MR integron int. I 1 qac. EΔ sul 1 qac. EΔ qac. ED sul 1 qac. EΔ sul 1 SIXanthomonassp In a single environment, a wastewater treatment plant in germany: 97 different resistance plasmids, different species resistance cassettes SI 21 unknown (Tennstedt T. , FEMS Microbiol Ecol. (2003) 45: 239)

Bacterial resistance evolution by recruitment of super-integron gene cassettes

Potential antibiotic resistance cassettes in the V. cholerae N 16961 SI ORF Number on Chromosome IIb Proposed Function VCA 0300 Chloramphenicol acetyltransferase* VCA 0328, VCA 0341, VCA 0463 glutathione-transferase Fosfomycin VCA 0387 Phosphinothricin acetyltransferase VCA 0473 Streptogramin acetyltransferase b, the 214 ORFs in the 179 cassettes in the SI of V. cholerae strain N 16961 are numbered from VCA 0292 to VCA 0506 (Heidelberg et al. , 2000). *, VCA 0300 is 60% identical to CATB 6 and 40% identical to Sat. A
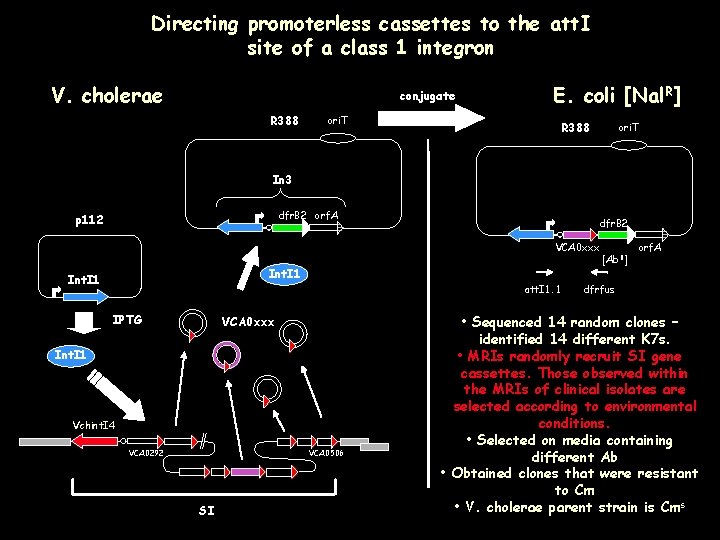
Directing promoterless cassettes to the att. I site of a class 1 integron V. cholerae conjugate R 388 ori. T E. coli [Nal. R] R 388 ori. T In 3 dfr. B 2 orf. A p 112 dfr. B 2 VCA 0 xxx Int. I 1 att. I 1. 1 IPTG VCA 0 xxx Int. I 1 Vchint. I 4 VCA 0292 VCA 0506 SI [Ab. R] orf. A dfrfus • Sequenced 14 random clones – identified 14 different K 7 s. • MRIs randomly recruit SI gene cassettes. Those observed within the MRIs of clinical isolates are selected according to environmental conditions. • Selected on media containing different Ab • Obtained clones that were resistant to Cm • V. cholerae parent strain is Cms

This is not limited to this unique example. Recently: • Two CARB genes have been identified in the SI of environmental Vibrio isolates. These genes show signatures that point them as the ancestors of all the cassette-encoded carbenicillinases found in multi-resistance integrons. (Petroni, AAC (2004) 48: 4042 -46) • We have identified two novel dfr cassettes in the SI of two environmental V. splendidus.

Conclusions • The widely spread multi-resistance integrons and their cassettes derive from the sedentary super-integrons and their cassette pools. • The stability of the silent SI cassette arrays, hundreds of cassettes long, is due to both a tight control of the integrase expression and the presence of multiple addiction cassettes. • If we made progress in understanding the recombination reactions, we still do not know the cassette genesis process…

Part 2 Cassette insertion in integrons : a novel recombination process involving a folded single stranded substrate.
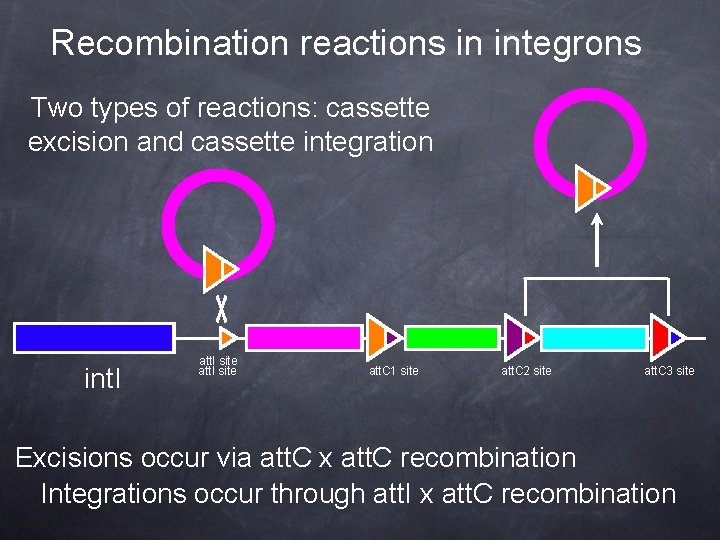
Recombination reactions in integrons Two types of reactions: cassette excision and cassette integration int. I att. I site att. C 1 site att. C 2 site att. C 3 site Excisions occur via att. C x att. C recombination Integrations occur through att. I x att. C recombination

The integron paradox The att. C and att. I sites structure differ from the canonical Tyrosine Recombinase Recombination Sites Cre, Flp, Xer. CD and Intλ : the recombination occurs between two very similar sites Simple Site RBE 6 Lox. P (Cre) Frt (Flp) 8 Arg. R Int 8 Cer (Xer. CD) 7 att. P (Intλ ) Xis Pep. A IHF Fis

Potential RBEs in the att. I sites ? att. I 1 cat-------------------------------------- ||| att. I 4 catatagttctcactgaatatttaactggttatttgtacagtatttgttgtttttatgtcaagaggctatacag att. I 1 -----ggcttgttatga-ctg-tttttttgtacagtctatgcctcgggcatccaagcagcaagcgcgtta-cgccg--t ||| | |||||| | | |||| | att. I 4 acatcagcaatctataagctgagatttttgaatggtgtgatgctcaacatactgatttagaaggttgttatggtagtat CS att. I 1 gggtcga-t-gtttgat-gttatggagcagcaacgat. GTTACGCagcagggcagtcgccct-aaaacaaa. GTTAGAT | | | | ||| | | ||| || || | | ||||| | att. I 4 gcacccagtggtctattagatagcggtagcctacctgttggaaaggtaagaagctgtctagaaagc. GTTAGTT

att. C sites characteristics • little primary sequence conservation among the att. C sites, except their boundaries, ICS and CS • the ICS is always complementary to the CS of the circularized cassette. • Sizes vary from 57 to 142 nt att. C site ICS variable region (RYYYAAC) • imperfect palindromic structures CS (G / TTRRRY) Stokes, H. and Hall, R (1989) Mol Micro 3: 1669; Collis, C. and Hall, R. (1992) Mol Micro 6: 2875; Recchia, G. and Hall (1995) Microbiol 141: 3015
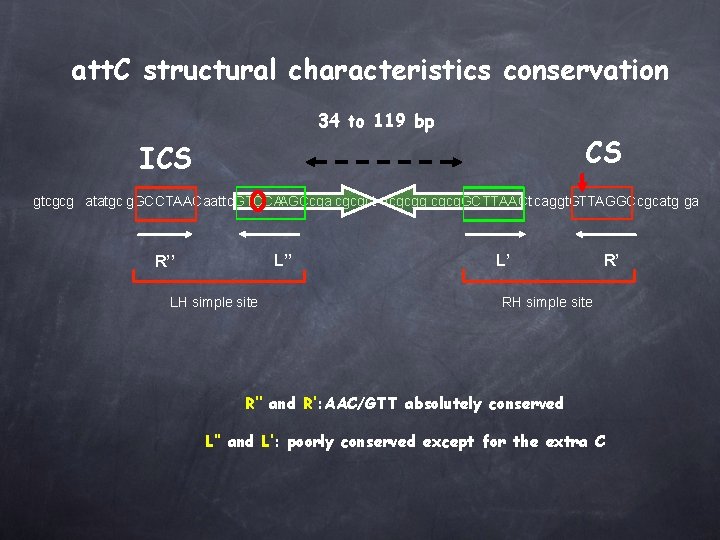
att. C structural characteristics conservation 34 to 119 bp CS ICS gtcgcg atatgc g. GCCTAACaattc. GTCCAAGCcga cgcgct tcgcgg cgcg. GCTTAACt caggt. GTTAGGCcgcatg ga L’’ R’’ LH simple site L’ R’ RH simple site R’’ and R’: AAC/GTT absolutely conserved L’’ and L’: poorly conserved except for the extra C

In 1999, the group of Fernando de la Cruz made the following observations in DNA / Integrase binding experiments + Int. I 1 Gel mobility shift assays with ds. DNA fragments containing aad. A 1 att. C (left) or the att. I site (right). Lanes 1, control ds. DNA; lanes 2, ds. DNA plus E. coli C 41 control extract; lanes 3, ds. DNA plus pure Int. I 1 -COOH; lanes 4, ds. DNA plus purified Int. I 1. F, free ds. DNA; B, DNAprotein complex. =>Int. I does not bind ds att. C Francia MV, Zabala JC, de la Cruz F, Garcia Lobo JM. J. Bacteriol. (1999) 181: 6844 -9.
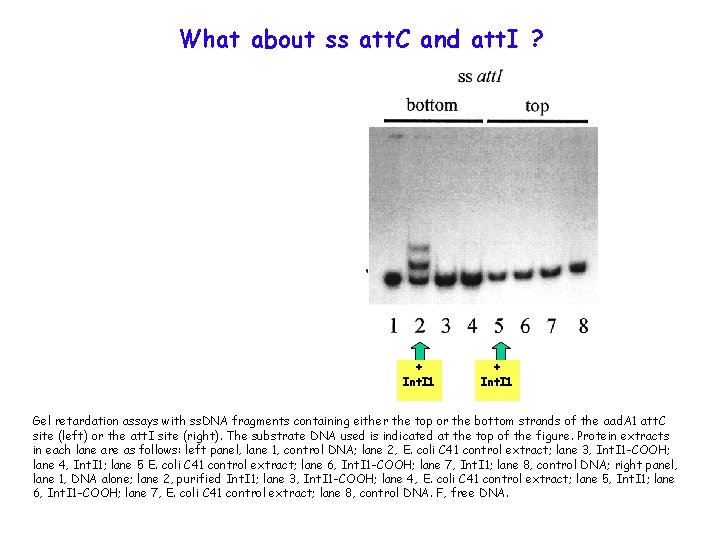
What about ss att. C and att. I ? + Int. I 1 Gel retardation assays with ss. DNA fragments containing either the top or the bottom strands of the aad. A 1 att. C site (left) or the att. I site (right). The substrate DNA used is indicated at the top of the figure. Protein extracts in each lane are as follows: left panel, lane 1, control DNA; lane 2, E. coli C 41 control extract; lane 3, Int. I 1 -COOH; lane 4, Int. I 1; lane 5 E. coli C 41 control extract; lane 6, Int. I 1 -COOH; lane 7, Int. I 1; lane 8, control DNA; right panel, lane 1, DNA alone; lane 2, purified Int. I 1; lane 3, Int. I 1 -COOH; lane 4, E. coli C 41 control extract; lane 5, Int. I 1; lane 6, Int. I 1 -COOH; lane 7, E. coli C 41 control extract; lane 8, control DNA. F, free DNA.
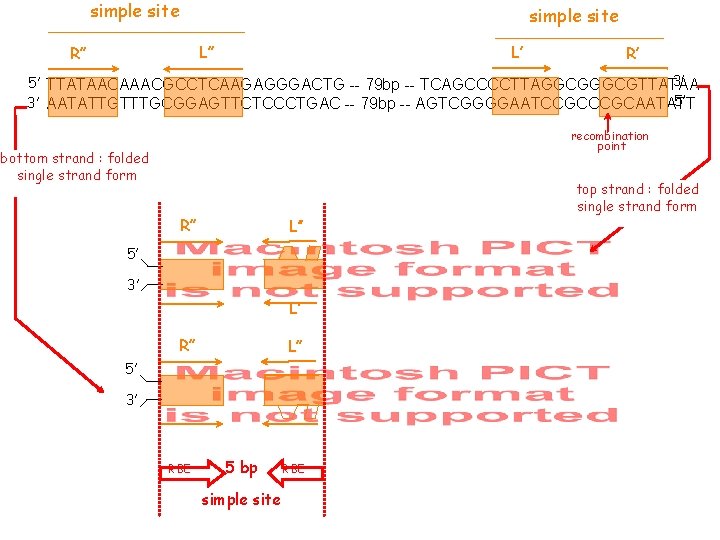
simple site L’ L’’ R’ 3’ 5’ TTATAACAAACGCCTCAAGAGGGACTG -- 79 bp -- TCAGCCCCTTAGGCGTTATAA 5’ 3’ AATATTGTTTGCGGAGTTCTCCCTGAC -- 79 bp -- AGTCGGGGAATCCGCAATATT recombination point bottom strand : folded single strand form R’’ L’’ R’ L’ 5’ 3’ RBE 5 bp simple site RBE top strand : folded single strand form
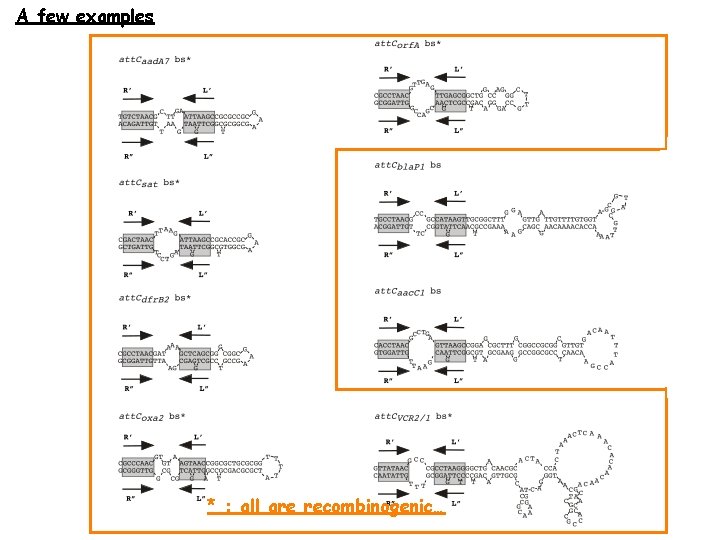
A few examples * : all are recombinogenic…
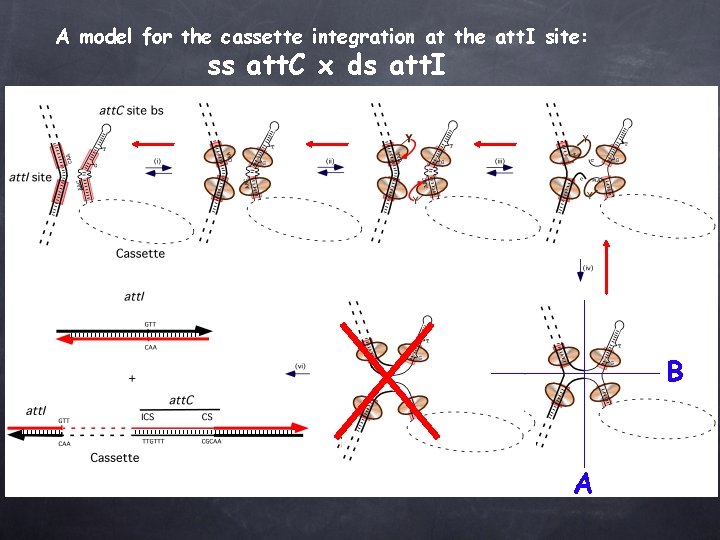
A model for the cassette integration at the att. I site: ss att. C x ds att. I B A

We thought that conjugation allowed to test the realism of a such a recombination model involving a folded single strand DNA: Indeed conjugation always goes through a single strand transfer - The transferred strand depends on the transfer origin orientation => It is always the same Plasmid promiscuity : meeting the challenge of DNA immigration control. B. Wilkins. Env Microbiology, 2002, 4: 495
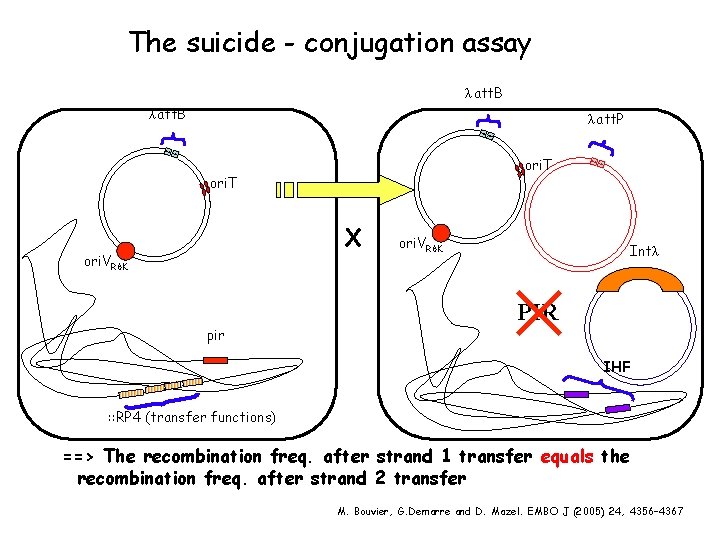
The suicide - conjugation assay λ att. B λ att. P ori. T X ori. VR 6 K pir ori. VR 6 K Intλ PIR IHF : : RP 4 (transfer functions) ==> The recombination freq. after strand 1 transfer equals the recombination freq. after strand 2 transfer M. Bouvier, G. Demarre and D. Mazel. EMBO J (2005) 24, 4356– 4367
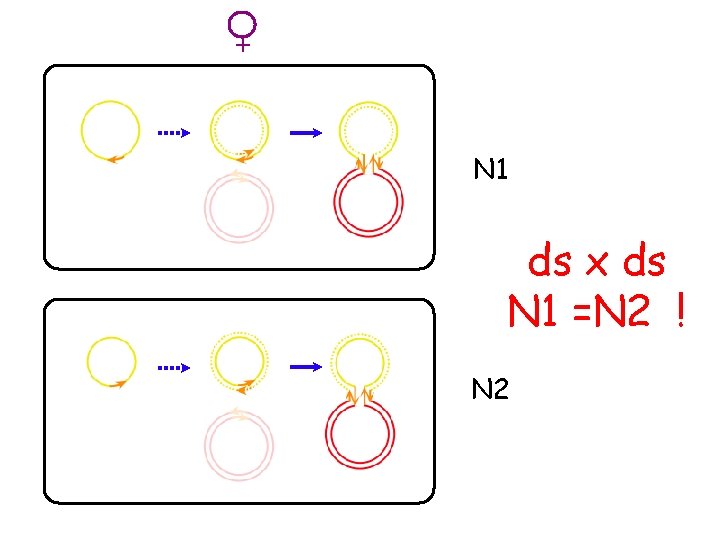
+ N 1 ds x ds N 1 =N 2 ! N 2
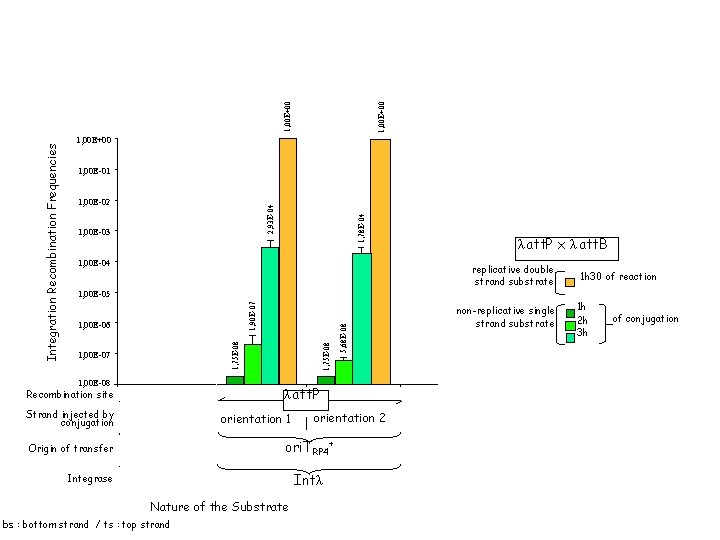
1, 00 E+00 1, 00 E-01 1, 78 E-04 2, 93 E-04 1, 02 E-05 1, 00 E-03 1, 00 E-04 replicative double strand substrate non-replicative single strand substrate 1, 99 E-03 1, 57 E-03 1, 00 E-02 9, 68 E-06 16 h of reaction 16 h of conjugation λ att. P x λ att. B replicative double strand substrate 1 h 30 of reaction 1, 00 E-06 1, 00 E-08 Strand injected by conjugation Origin of transfer Integrase λ att. P att. I 1 Recombination site ts bs 1, 75 E-08 1, 00 E-07 orientation 1 orientation 2 ori. TRP 4+ Int. I 1 Intλ Nature of the Substrate bs : bottom strand / ts : top strand 5, 68 E-08 1, 90 E-07 1, 00 E-05 1, 75 E-08 Integration Recombination Frequencies 1, 00 E+00 att. I 1 x att. Caad. A 7 non-replicative single strand substrate 1 h 2 h 3 h of conjugation
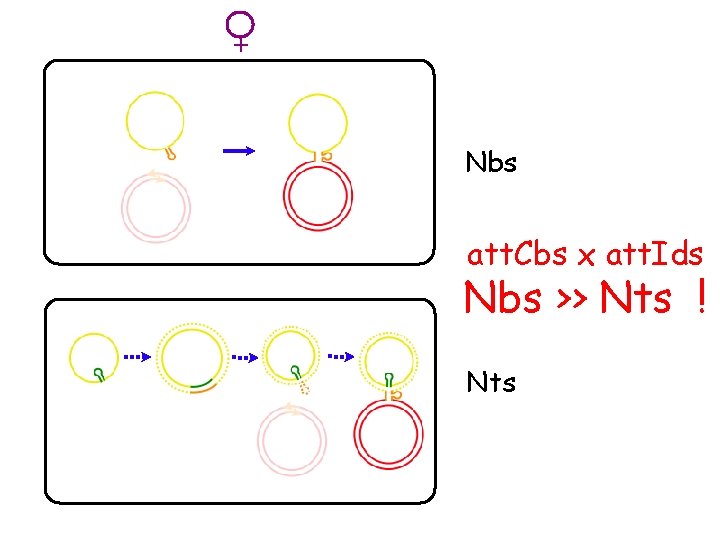
+ Nbs att. Cbs x att. Ids Nbs >> Nts ! Nts
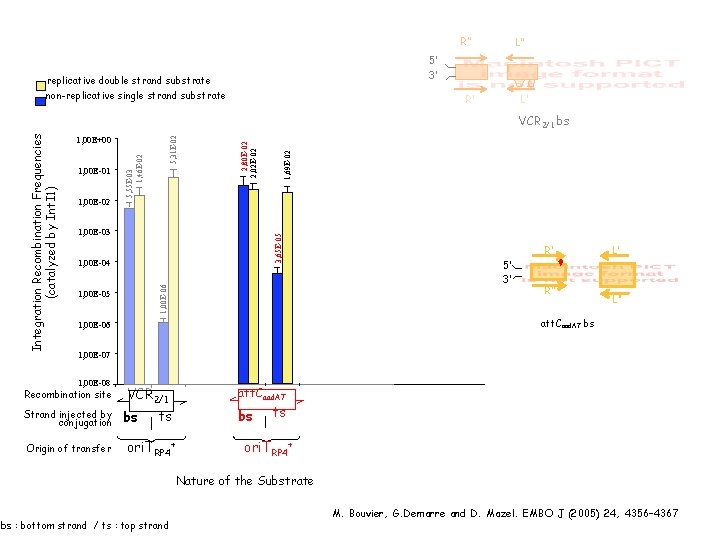
R’’ L’’ 5’ 3’ replicative double strand substrate non-replicative single strand substrate L’ R’ 1, 00 E-02 3, 65 E-05 1, 00 E-03 1, 69 E-02 1, 00 E-01 2, 80 E-02 2, 02 E-02 5, 31 E-02 1, 00 E+00 5, 55 E-03 1, 46 E-02 1, 00 E-04 1, 00 E-06 Integration Recombination Frequencies (catalyzed by Int. I 1) VCR 2/1 bs 1, 00 E-05 R’ 5’ 3’ R’’ L’ L’’ att. Caad. A 7 bs att. Caad. A 7 WT 1, 00 E-06 1, 00 E-07 1, 00 E-08 Recombination site Strand injected by conjugation Origin of transfer VCR 2/1 bs att. Caad. A 7 bs ts ori. TRP 4+ Nature of the Substrate bs : bottom strand / ts : top strand M. Bouvier, G. Demarre and D. Mazel. EMBO J (2005) 24, 4356– 4367
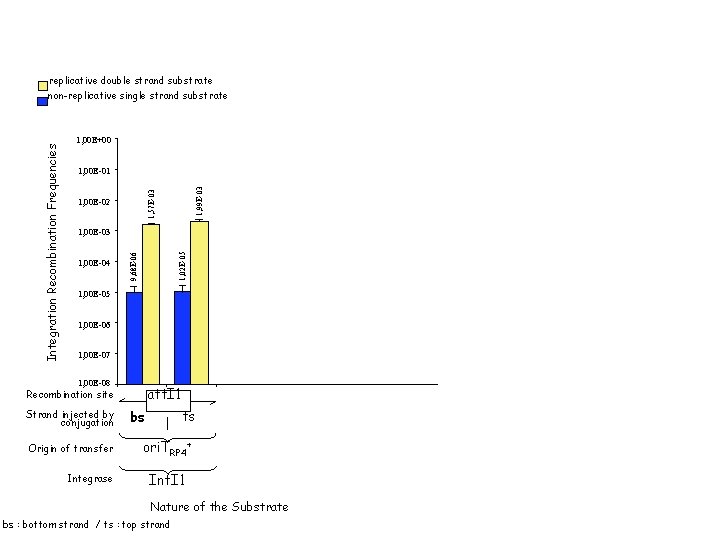
replicative double strand substrate 1, 00 E+00 1, 57 E-03 1, 99 E-03 1, 00 E-01 1, 00 E-02 1, 00 E-04 1, 02 E-05 1, 00 E-03 9, 68 E-06 Integration Recombination Frequencies non-replicative single strand substrate 1, 00 E-05 1, 00 E-06 1, 00 E-07 1, 00 E-08 att. I 1 Recombination site Strand injected by conjugation Origin of transfer Integrase ts bs ori. TRP 4+ Int. I 1 Nature of the Substrate bs : bottom strand / ts : top strand
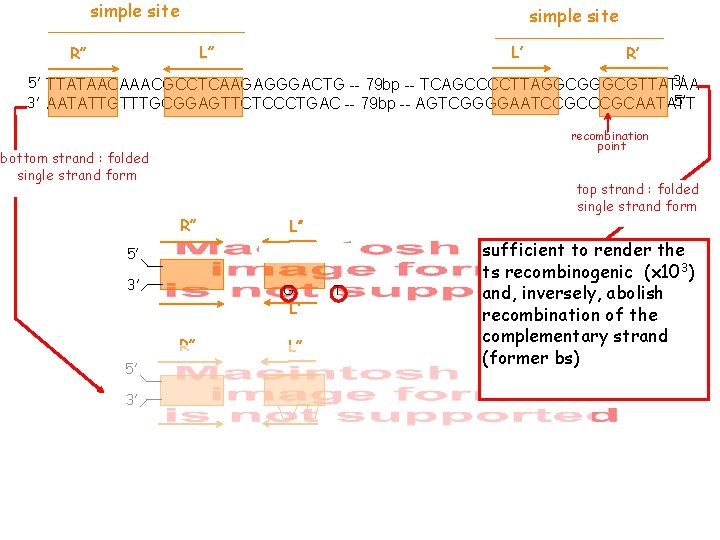
simple site L’ L’’ R’ 3’ 5’ TTATAACAAACGCCTCAAGAGGGACTG -- 79 bp -- TCAGCCCCTTAGGCGTTATAA 5’ 3’ AATATTGTTTGCGGAGTTCTCCCTGAC -- 79 bp -- AGTCGGGGAATCCGCAATATT recombination point bottom strand : folded single strand form R’’ top strand : folded single strand form L’’ 5’ 3’ G R’ L’ R’’ L’’ R’ L’ 5’ 3’ T sufficient to render the ts recombinogenic (x 103) and, inversely, abolish recombination of the complementary strand (former bs)
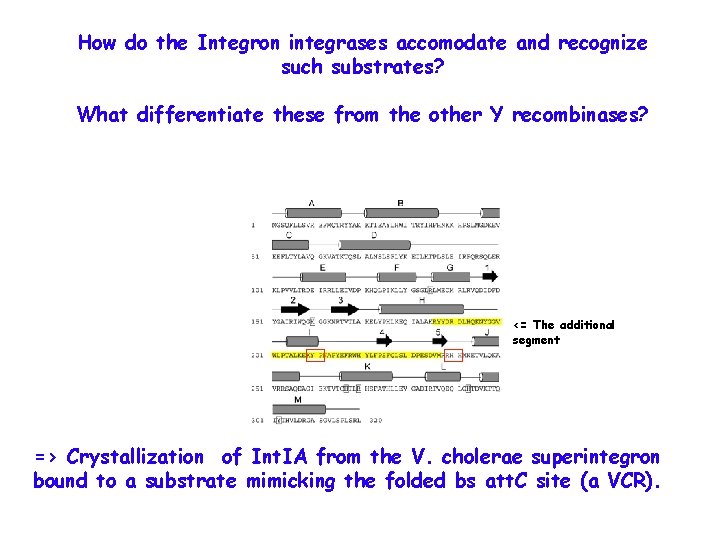
How do the Integron integrases accomodate and recognize such substrates? What differentiate these from the other Y recombinases? <= The additional segment => Crystallization of Int. IA from the V. cholerae superintegron bound to a substrate mimicking the folded bs att. C site (a VCR).

Vch. Int. IA – VCRbs Hairpin tetrameric synapse structure 2 fold symmetry, not 4 X : -A and C active, bound to G 20 in trans -B and D bound to T 12 in cis => disorganization of their catalytic domains. D. Mac. Donald, G. Demarre, M. Bouvier, D. Mazel and D. N. Gopaul. Nature (2006) 440, 1157 -62)
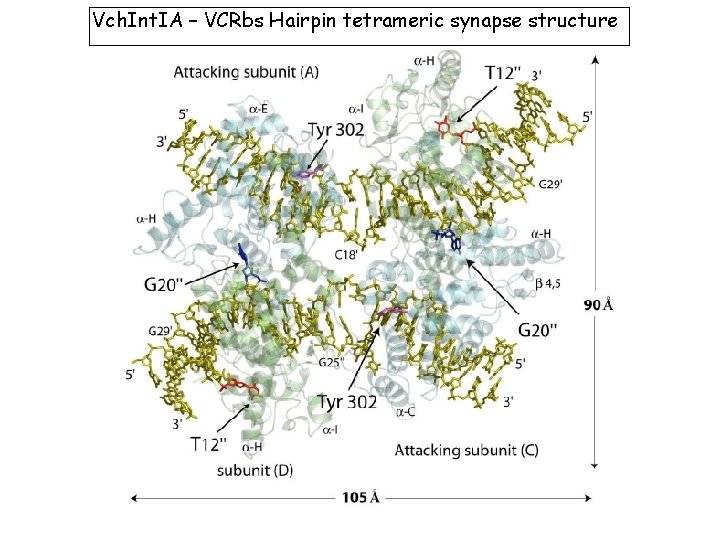
Vch. Int. IA – VCRbs Hairpin tetrameric synapse structure

94
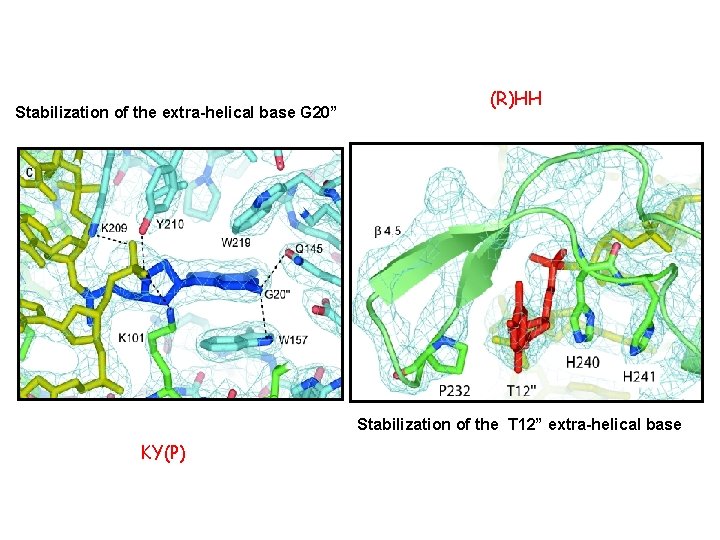
Stabilization of the extra-helical base G 20” (R)HH Stabilization of the T 12” extra-helical base KY(P)
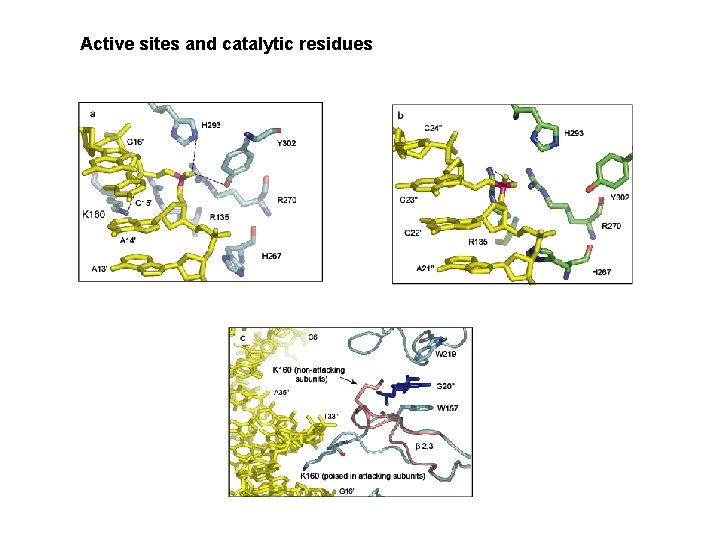
Active sites and catalytic residues
- Slides: 96