Bacterial chemotaxis Dr habil Khidai Lszl 2017 Diverse
Bacterial chemotaxis Dr. habil. Kőhidai László 2017.
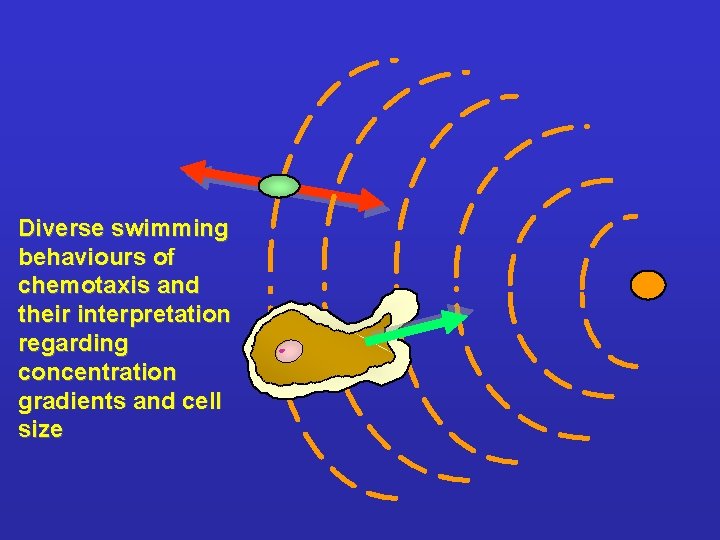
Diverse swimming behaviours of chemotaxis and their interpretation regarding concentration gradients and cell size
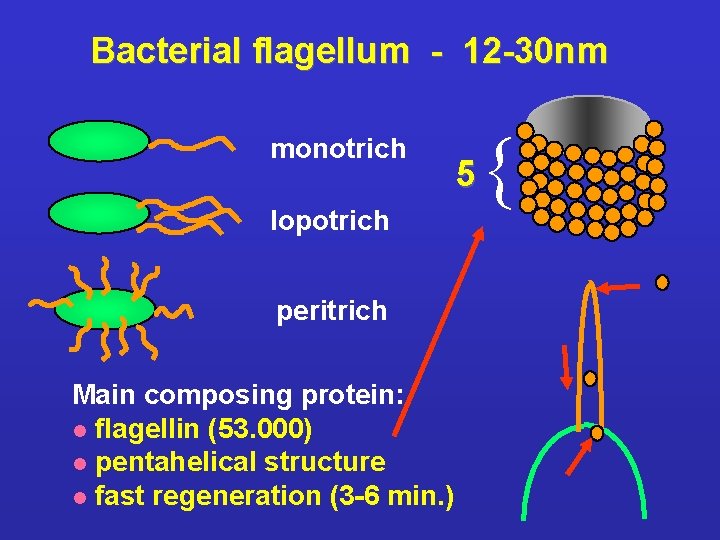
Bacterial flagellum - 12 -30 nm monotrich lopotrich peritrich Main composing protein: l flagellin (53. 000) l pentahelical structure l fast regeneration (3 -6 min. ) 5 {
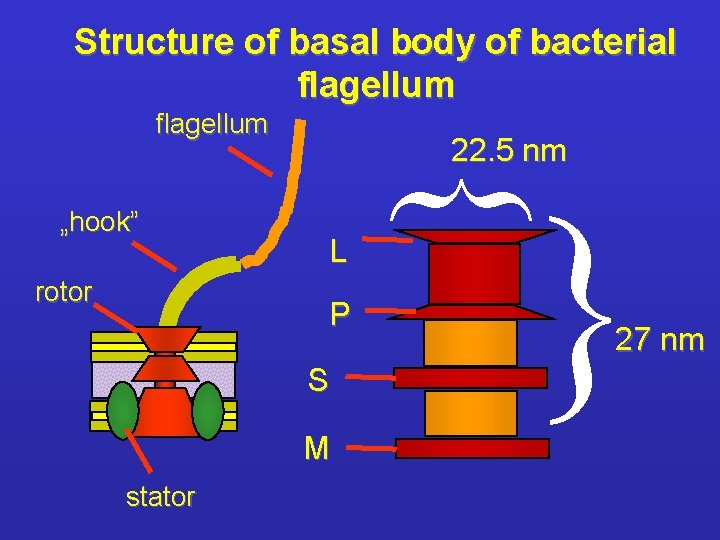
Structure of basal body of bacterial flagellum P S M stator { rotor L { „hook” 22. 5 nm 27 nm
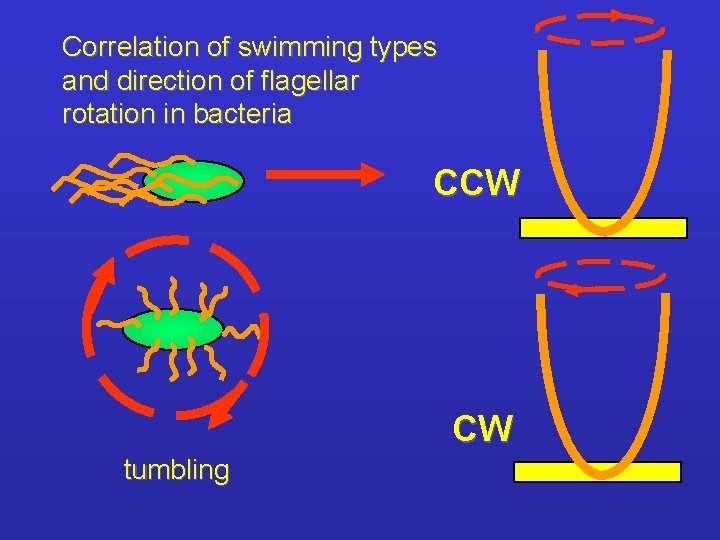
Correlation of swimming types and direction of flagellar rotation in bacteria CCW CW tumbling
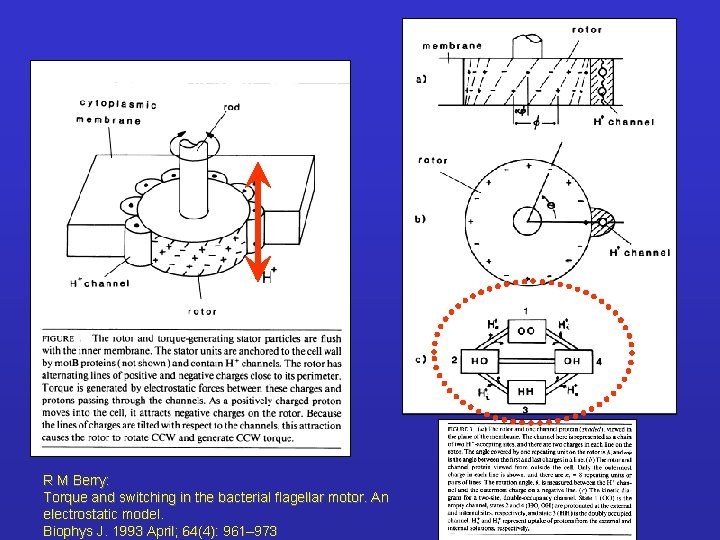
R M Berry: Torque and switching in the bacterial flagellar motor. An electrostatic model. Biophys J. 1993 April; 64(4): 961– 973
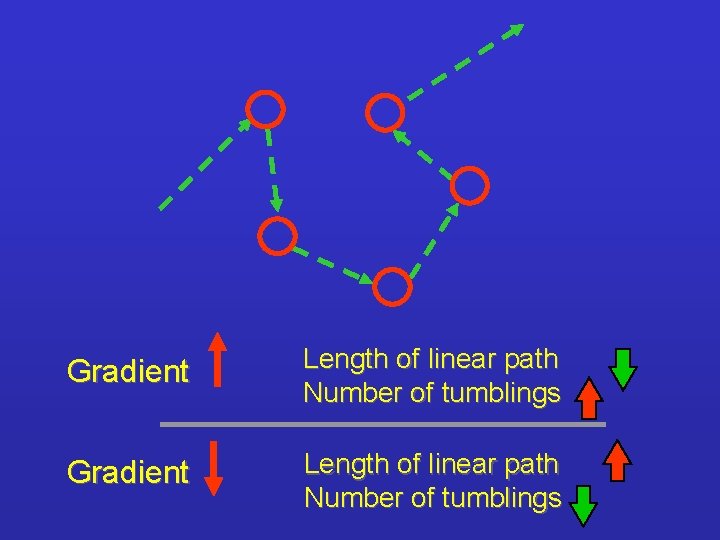
Gradient Length of linear path Number of tumblings
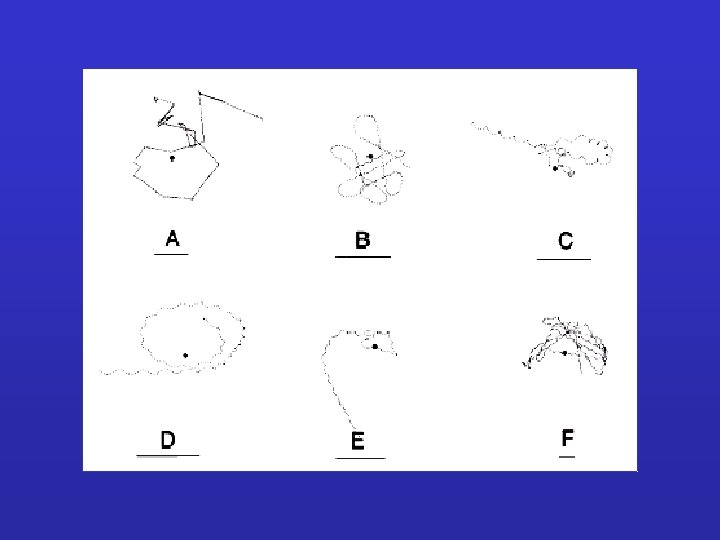
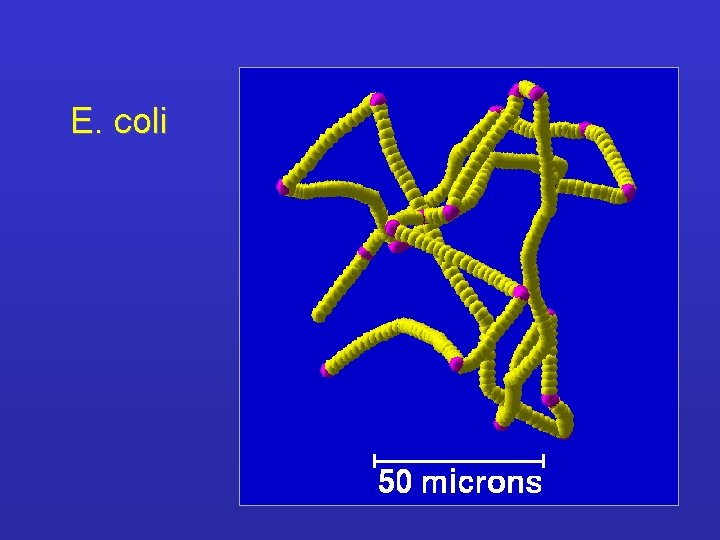
E. coli
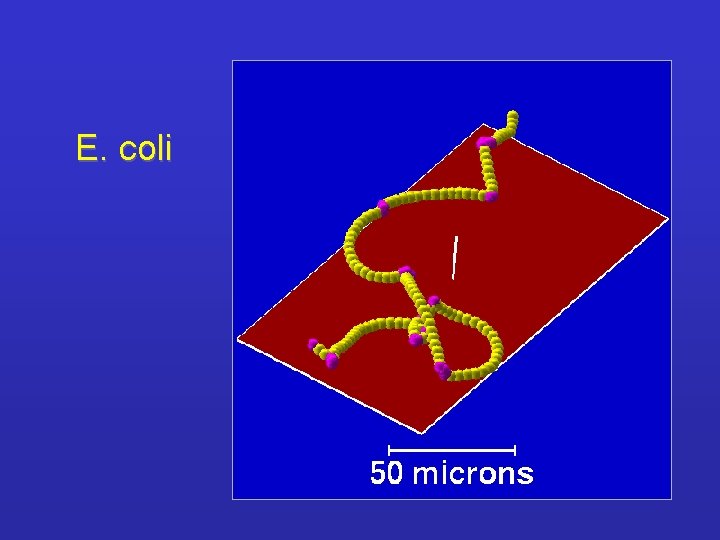
E. coli

Bacterial chemotaxis and adaptation Swimming of cells is influenced NOT ONLY by the changes of concentration of the ligand. ! Adaptation mechanisms refer to the presence of a ‘primitive’ memory of cells
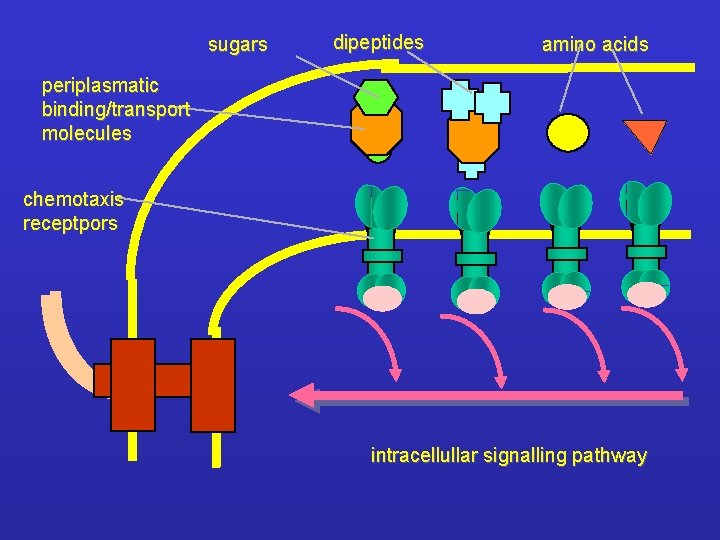
sugars dipeptides amino acids periplasmatic binding/transport molecules chemotaxis receptpors intracellullar signalling pathway
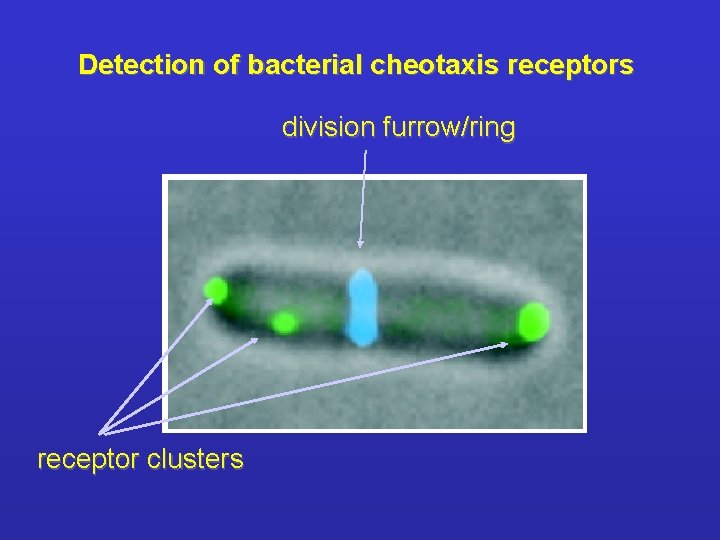
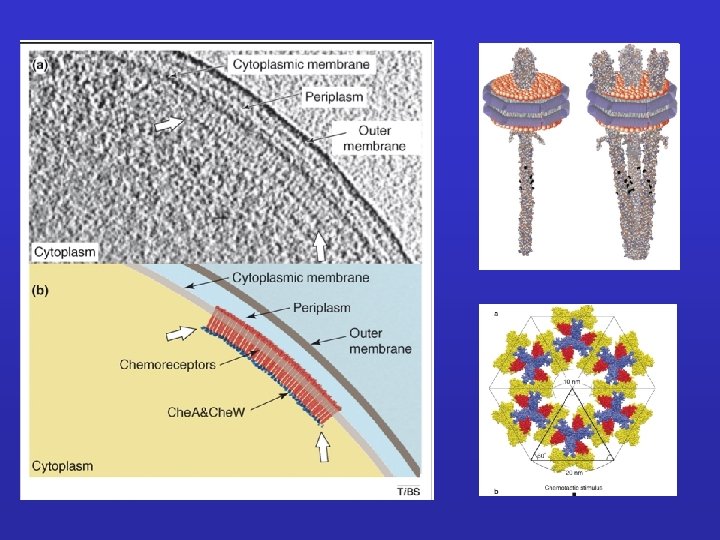
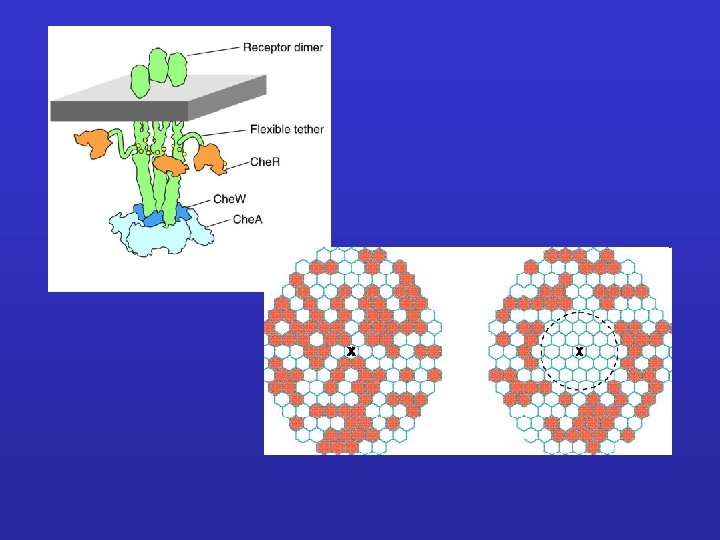
Detection of bacterial cheotaxis receptors division furrow/ring receptor clusters
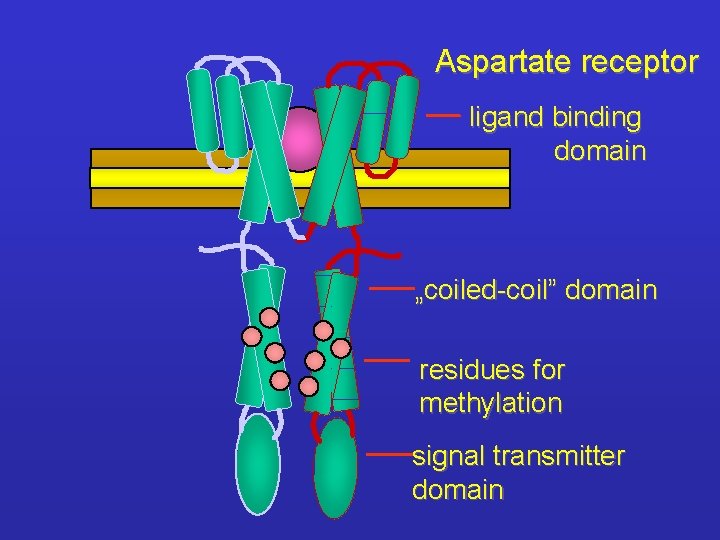
Aspartate receptor ligand binding domain „coiled-coil” domain residues for methylation signal transmitter domain
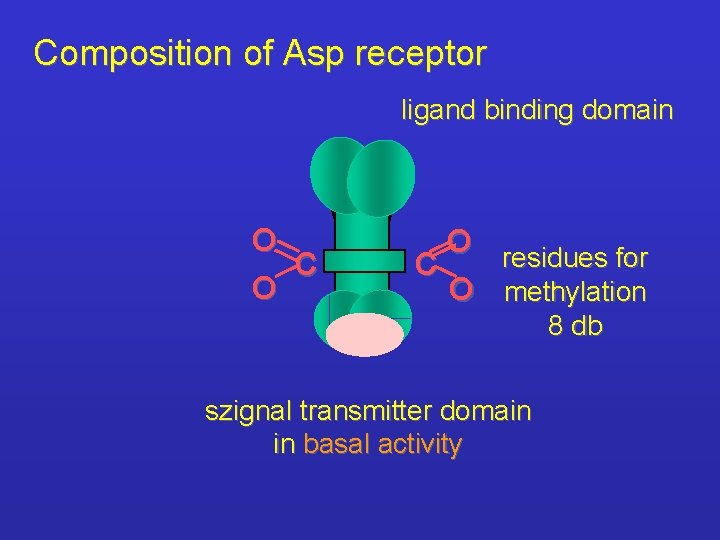
Composition of Asp receptor ligand binding domain O C O O residues for C O methylation 8 db szignal transmitter domain in basal activity
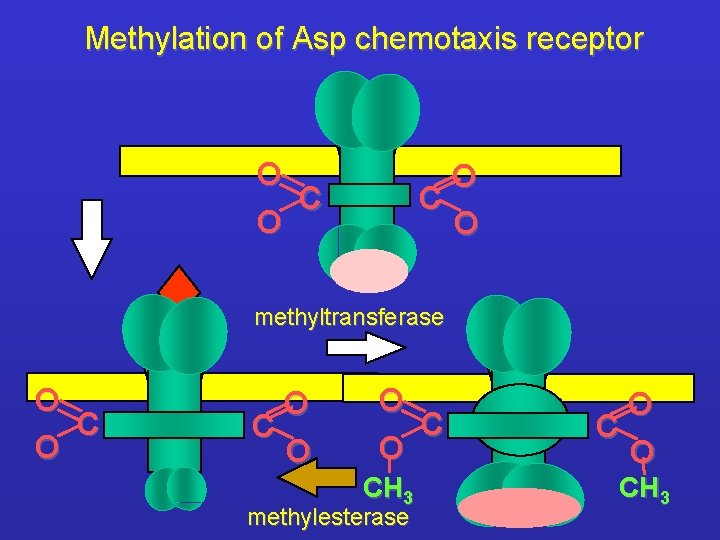
Methylation of Asp chemotaxis receptor O C O methyltransferase O C O O CH 3 methylesterase O CH 3
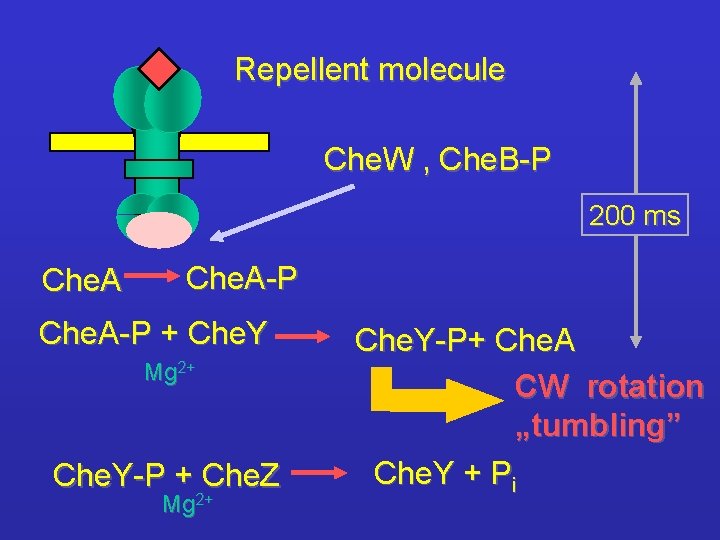
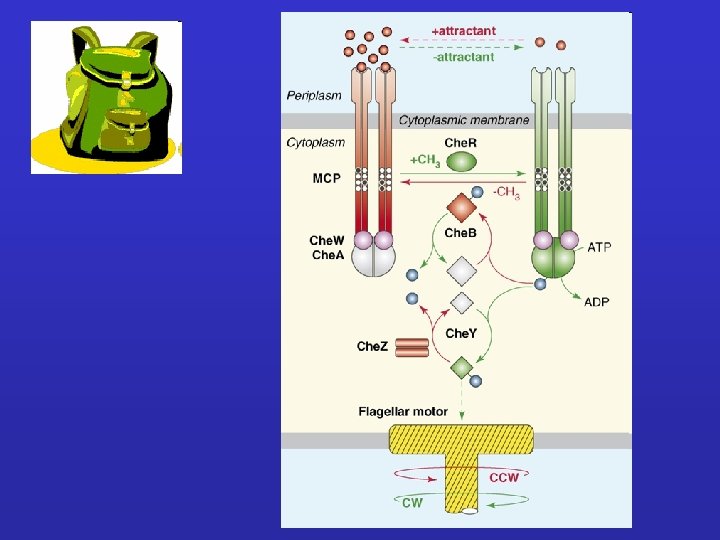
Repellent molecule Che. W , Che. B-P 200 ms Che. A-P + Che. Y Mg 2+ Che. Y-P + Che. Z Mg 2+ Che. Y-P+ Che. A CW rotation „tumbling” Che. Y + Pi
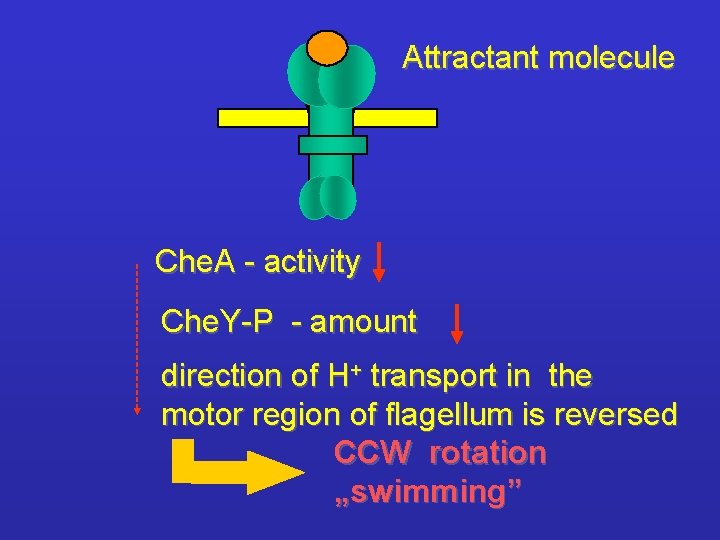
Attractant molecule Che. A - activity Che. Y-P - amount direction of H+ transport in the motor region of flagellum is reversed CCW rotation „swimming”
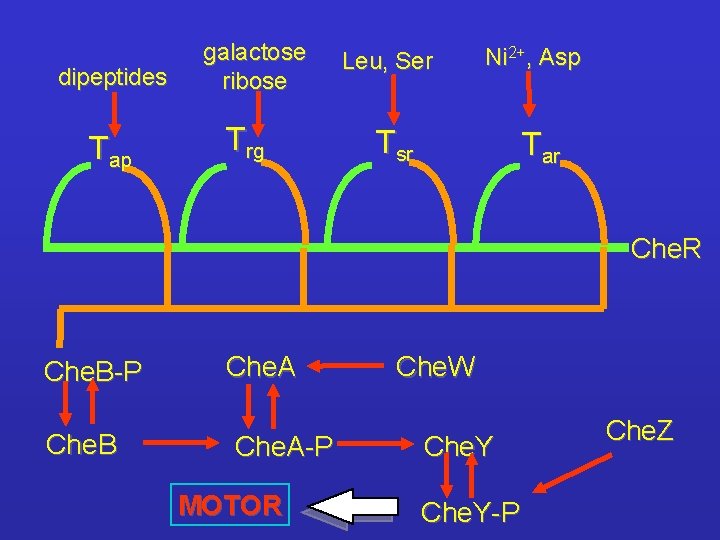
dipeptides Tap galactose ribose Trg Leu, Ser Ni 2+, Asp Tsr Tar Che. R Che. B-P Che. B Che. A-P MOTOR Che. W Che. Y-P Che. Z
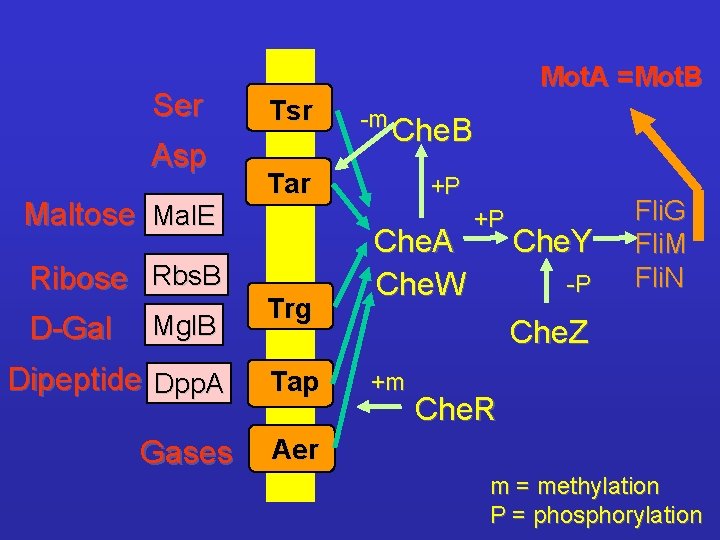
Ser Asp Maltose Mal. E Ribose Rbs. B D-Gal Mgl. B Dipeptide Dpp. A Gases Mot. A =Mot. B Tsr -m Che. B Tar Trg Tap +P Che. A Che. W +P Che. Y -P Fli. G Fli. M Fli. N Che. Z +m Che. R Aer m = methylation P = phosphorylation
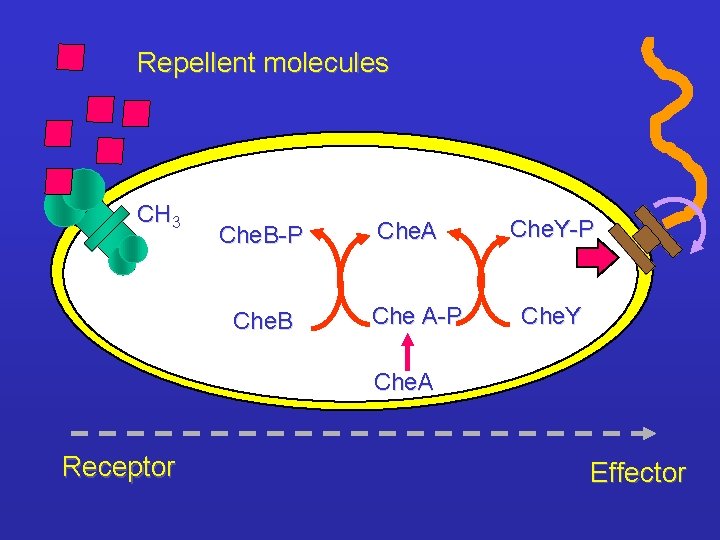
Repellent molecules CH 3 Che. B-P Che. B Che. A Che A-P Che. Y Che. A Receptor Effector
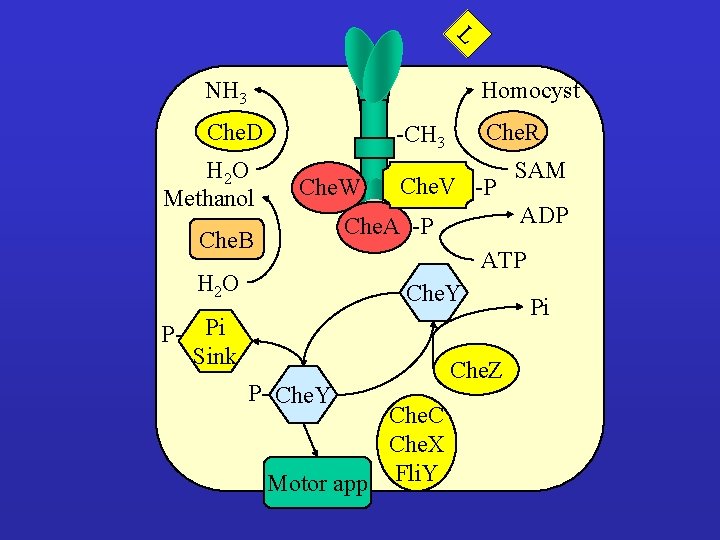
L NH 3 Homocyst Che. D H 2 O Methanol Che. R -CH 3 Che. W Che. V -P ADP Che. A -P Che. B SAM ATP H 2 O Che. Y P- Pi Sink P- Che. Y Che. C Che. X Motor app Fli. Y Che. Z Pi
Structure of Che. Y
Structure of Ch. A - Ch. Y complex
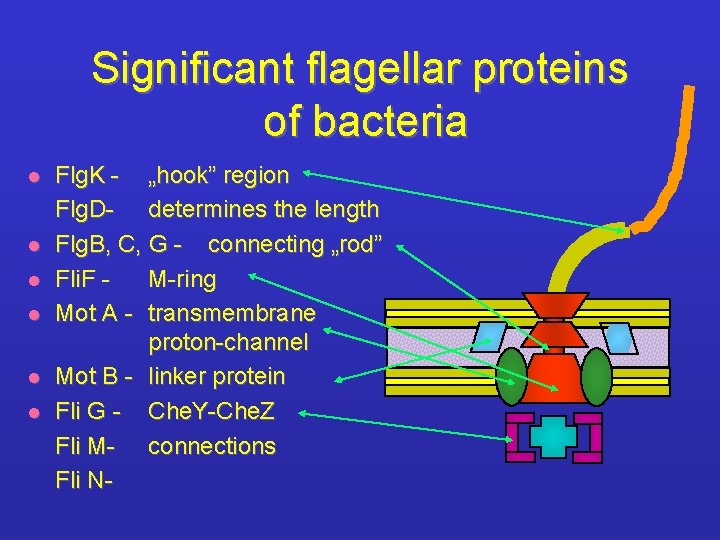
Significant flagellar proteins of bacteria l l l Flg. K - „hook” region Flg. D- determines the length Flg. B, C, G - connecting „rod” Fli. F M-ring Mot A - transmembrane proton-channel Mot B - linker protein Fli G - Che. Y-Che. Z Fli M- connections Fli N-

Flagellar proteins l Determined by more than 30 genes organized into several operons l Their synthesis / expression is regulated by Sigma 28 factor l „Hook associated protein” (HAP) : - nucleation point of flagellins - increases the mechanical stability l Main classes: Fli, Flg, Flh

Characterization of bacterial chemotaxis proteins l Che. A - histidine autokinase P 1 - 22 amino acids, non inhibited region P 2 - 25 amino acis, interacts with Che. Y Ø Che. AL (long) - His 48 autophosphorylation which is a component of the Che. Y and Che. B activation Ø Che. AL – its function is p. H-dependent. Optimal p. H 8. 1 - 8. 9 - Tar és Trg receptors signalling is turned on when cytopl. p. H decreses below p. H 7. 6 Ø Ch. AS (short) – possesses kinase activity, but the subunit does not autophosphorylating - the aminoterminal 97 aa. long sequence is missing

Characterization of bacterial chemotaxis proteins l Che. A hyper kinase – ponit mutation in Pro 337 which results a faster phosphorylation l Che. A - regulates phsphorylation of Che. V l Che. N - present in Bacillis substilisban and homologue to Che. A of E. coli

Characterization of bacterial chemotaxis proteins l Che. Y - Composed by 128 aa. , its phosphorylation results a conformational change in positions listed below: 17, 21, 23, 39, 60, 63, 64, 66, 67, 68, 69, 85, 86, 87, 88, 94, 107, 109, 112, 113, 114, 121 l Presence of Mg 2+ is essential for activation of Che. Y; Mg 2+ results the release of salt bond Lys 109 - Asp 57 which makes possible the phosphorylation
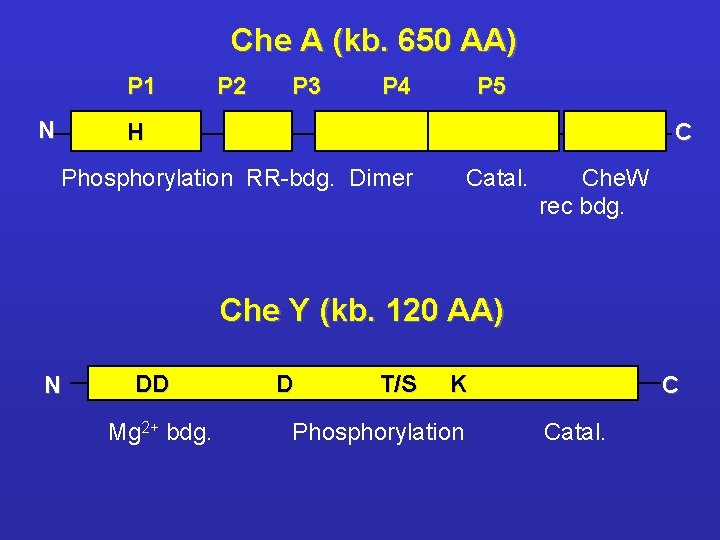
Che A (kb. 650 AA) P 1 N P 2 P 3 P 4 P 5 C H Phosphorylation RR-bdg. Dimer Catal. Che. W rec bdg. Che Y (kb. 120 AA) N DD Mg 2+ bdg. D T/S K Phosphorylation C Catal.
Characterization of Methyl-Accepting Chemotaxis proteines (MCP) l MCP 1 - Tsr, MCP 2 - Tar, MCP 3 - Trg, MCP 4 - Tap l. H 1 - 97 k. D p. I 5. 1; H 2 - 86 k. D p. I 5. 1; H 3 - 76 k. D p. I 5. 3 l. Dcr. A l - composed by 668 aa. , oxygen sensor composed by hem and 2 hydrophobic sequences induced by changes in redox-potential (Desulfovibrio vulgaris) Tlpc - 30% homology with E. coli MCP; its defect resulst the loss of pathological chemotaxis

Characterization of Methyl-Accepting Chemotaxis proteines (MCP) Methylation is a food molecule dependent process (e. g. E. coli) l Starvation results the methylation of a membrane associated 43 k. D protein; - in the presence of food the methylation is stopped l The link between the methylation system and activation of chemotaxis points to the essential common phylogenetical background of chemotaxis receptor and the signalling process.
Characterization of Methyl-Accepting Chemotaxis proteines (MCP) l MCP-k demethylation -CH 3 Attractant CARRIER-CH 3 MCP-CH 3 rapid CARRIER-CH 3 Methanol + CARRIER slow The non methylated intermedier results „tumbling”, then the ADAPTATION takes place.
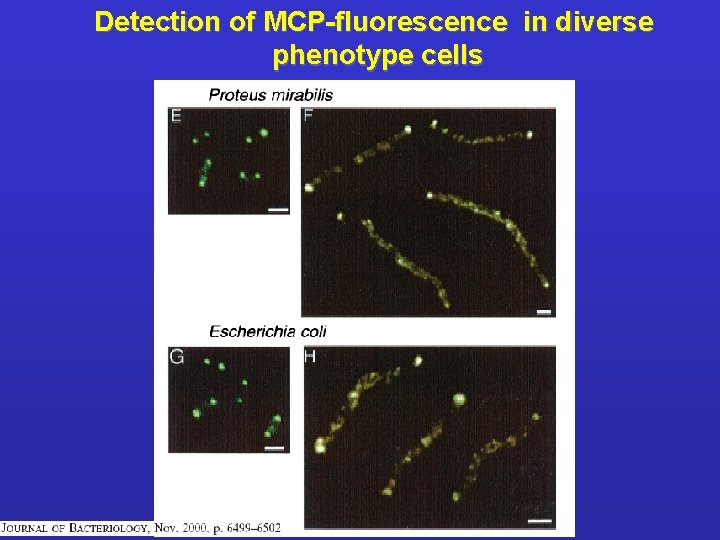
Detection of MCP-fluorescence in diverse phenotype cells
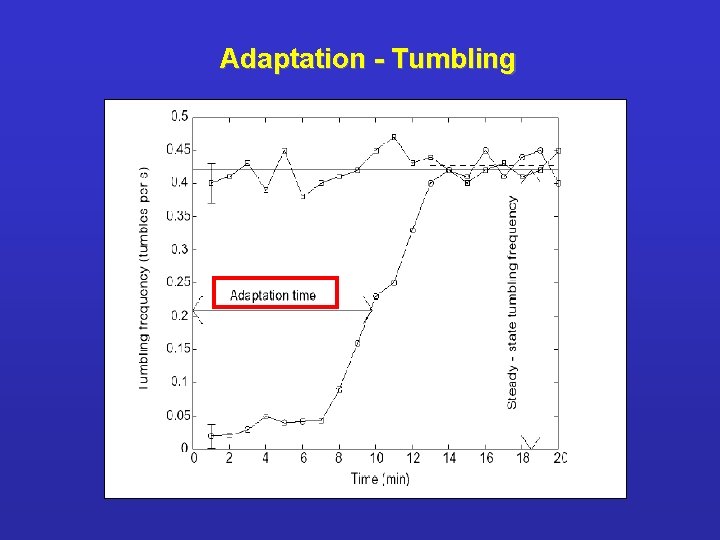
Adaptation - Tumbling
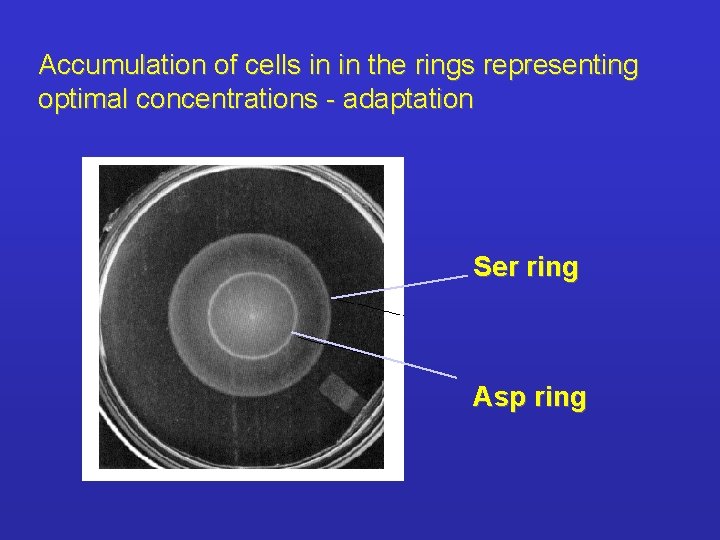
Accumulation of cells in in the rings representing optimal concentrations - adaptation Ser ring Asp ring

Methylation – Effect of carbohydrate type ligands
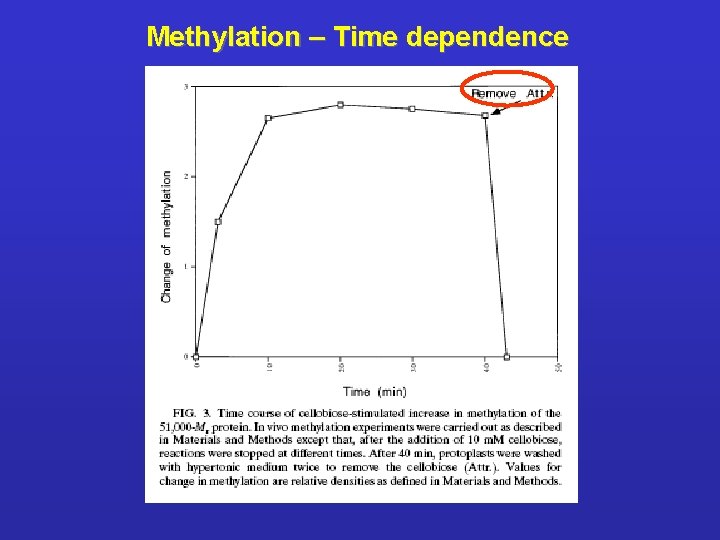
Methylation – Time dependence

Chemotaxis - Evolution Methyl-transferases Che. R Homology: E. coli methyl-transferase methylates MCP of Bac. subst. Difference: Bac. subst. Che. RB E. coli Che. RE Adaptation to repellents Adaptation to attractants

Chemotaxis - Evolution Methyl-esterases Che. B Homology: Bac. subst. MCP + ATTRACTANT E. coli Che. B DEMETHYLATION Bac. subst. Che. B + ATTRACTANT E. coli MCP DEMETHYLATION MCP determines the kinetics of reactions
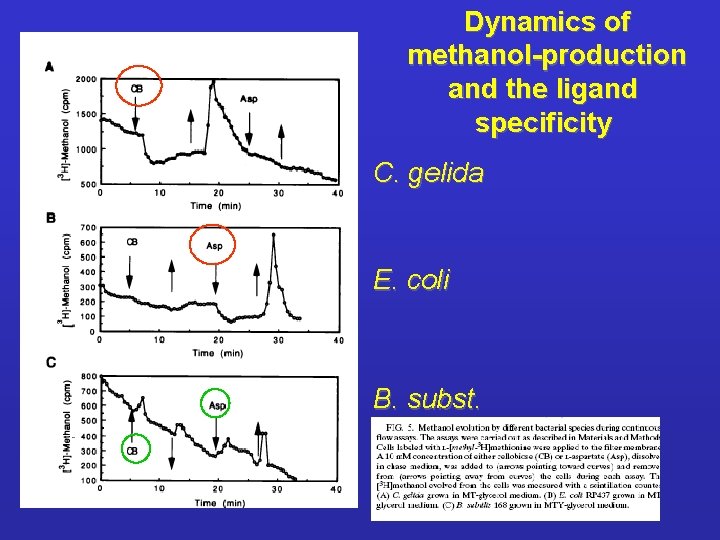
Dynamics of methanol-production and the ligand specificity C. gelida E. coli B. subst.
Chemotaxis - Evolution l. Bac. subst. Che. Y E. coli Che. A l. Bac. subst. Che. Y-P E. coli Che. Z l. Bac. subst. l. E. coli Che. Y-P Che. Y positive chemotaxis l. Bac. subst. - Che. Y-P - Chey-P and E. coli Che. W 28. 6% homology l. Bac. subst. Che. B and E. coli Che. Y 36% homology l. Bac. subst. and E. coli - M ring and rod

Effect of Ca 2+ on the bacterial chemotaxis l 38 k. D, Ca 2+-binding protein is detectable l Ca 2+ channel blockers (e. g. verapamil, La. Cl 3) disturbs chemotaxis
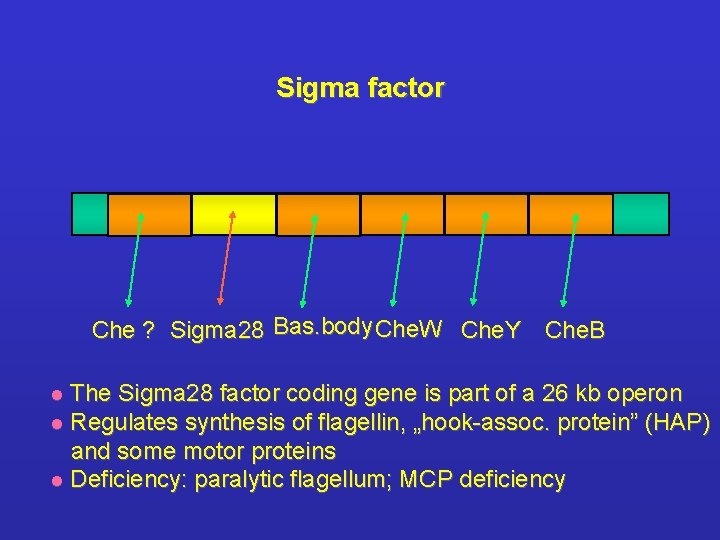
Sigma factor Che ? Sigma 28 Bas. body Che. W Che. Y Che. B The Sigma 28 factor coding gene is part of a 26 kb operon l Regulates synthesis of flagellin, „hook-assoc. protein” (HAP) and some motor proteins l Deficiency: paralytic flagellum; MCP deficiency l
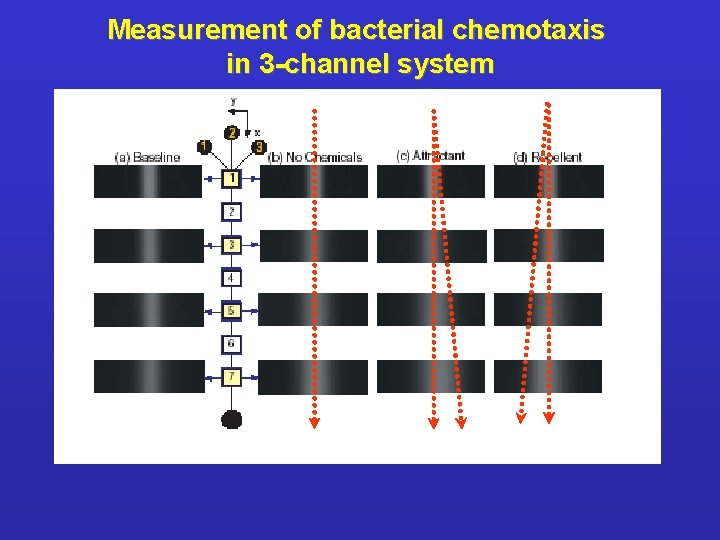
Measurement of bacterial chemotaxis in 3 -channel system
- Slides: 48