Bacterial Chemotaxis Bacteria swim toward attractants and away

Bacterial Chemotaxis • Bacteria swim toward attractants and away from repellents. • Their motion is a biased random walk due to control of tumbling frequency. • They measure gradients in time, not space. They adapt over time. • Chemotaxis probably relies on clustering and cooperation among receptors. Jason Kahn, UMCP

Fundamental mechanics of chemotaxis • Textbooks and Review Sources: • Macnab, R. M. (1987). Motility and chemotaxis. In Escherichia coli and Salmonella typhimurium: Cellular and molecular biology. 1 st ed. (Neidhardt, F. C. , ed. ), pp. 732 -759. Am. Soc. Microbiol. , Washington, D. C. • Stock, J. B. & Surette, M. G. (1996). Chemotaxis. In Escherichia coli and Salmonella typhimurium: Cellular and molecular biology. 2 nd ed. (Neidhardt, F. C. , ed. ), pp. 1103 -1129. Am. Soc. Microbiol. , Washington, D. C. • White, D. (2000). The physiology and biochemistry of prokaryotes. 2 nd ed. , Oxford University Press, Oxford. • Parkinson, J. S. (2004). Signal amplification in bacterial chemotaxis through receptor teamwork. ASM News 70, 575 -582. Jason Kahn, UMCP

Swimming driven by a flagellar motor • The flagellum is an organelle consisting of (1) a basal body that is a rotary motor, (2) a connecting hook, and (3) a long filament. • Counter-clockwise rotation leads to all of the filaments of the cell twining together about a common axis, and swimming in a straight line. • Clockwise rotation leads to separation of separate filaments and tumbling, leading to random reorientation of the cell. • Typically the cell swims for 1 sec at 25 um/sec and then tumbles for 0. 1 sec. A random walk. Chemoattractants increase the time spent swimming when the bacterium is going up the gradient. A biased random walk. Jason Kahn, UMCP
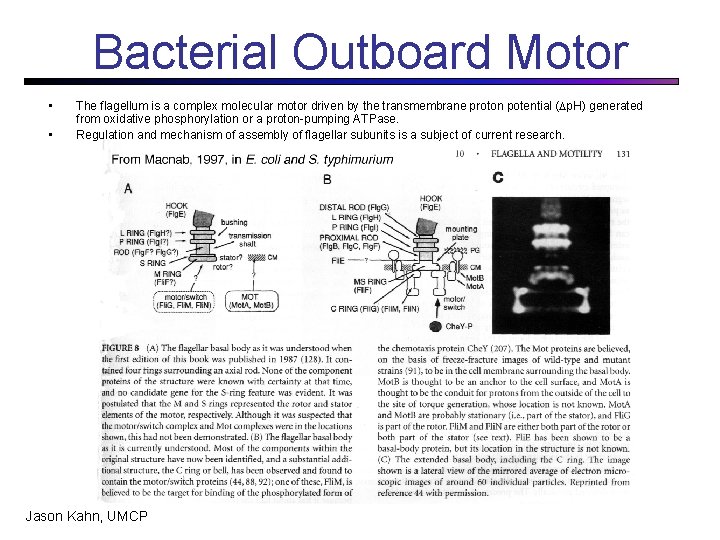
Bacterial Outboard Motor • • The flagellum is a complex molecular motor driven by the transmembrane proton potential ( p. H) generated from oxidative phosphorylation or a proton-pumping ATPase. Regulation and mechanism of assembly of flagellar subunits is a subject of current research. Jason Kahn, UMCP

• • • Tumbling and the Biased Random Walk From Macnab, 1987, E. coli and S. typhimurium Left hand side shows CCW flagellar rotation. The corkscrewing action of the left-handed helical filament drives the bacterium in a straight line. Right hand side shows. CW flagellar rotation, with the different filaments separating and driving in different directions, tumbling the cell. Increasing the time spent swimming between tumbles leads to net motion toward attractant. Decreasing the time spent swimming leads to motion away from a repellent. Jason Kahn, UMCP

Assays for chemotaxis • Swarm plates: on semisolid agar, observe motion away from the center of a self-generated nutrient gradient. • Microscopic observation of free-swimming From Berg, Physics Today, 1999 cells. • Tethering to an immobile surface allows observation of the rotation of the body of the cell. • Temporal stimulation (rapid changes in chemoeffector concentration). Jason Kahn, UMCP

Dynamics of Chemotaxis • Bacteria respond to temporal, not spatial signals. Given their size, the spatial gradient is undetectable unless they move! • Thus they have a short term “memory” of the chemoeffector concentration over the previous few seconds. • A change in concentration leads (after a brief latency phase) to excitation (a change in tumbling frequency) followed by adaptation: after a few seconds at a new effector concentration the cell resumes random tumbling. This allows the cell to continue following a gradient up or down. Jason Kahn, UMCP
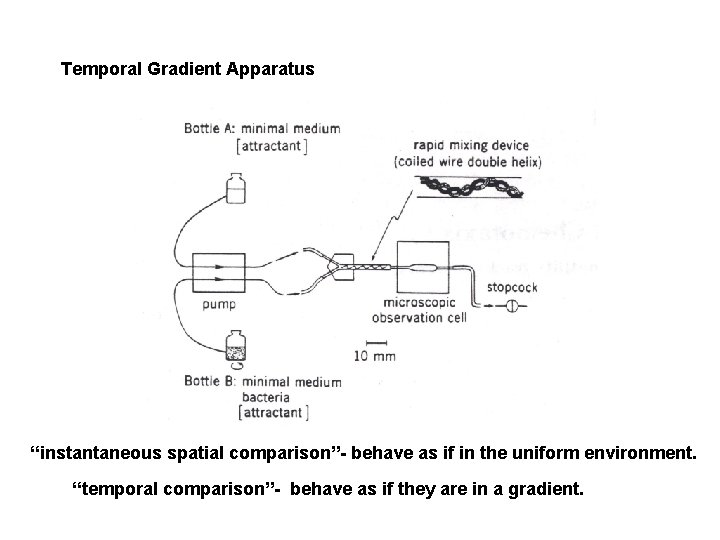
Temporal Gradient Apparatus “instantaneous spatial comparison”- behave as if in the uniform environment. “temporal comparison”- behave as if they are in a gradient.
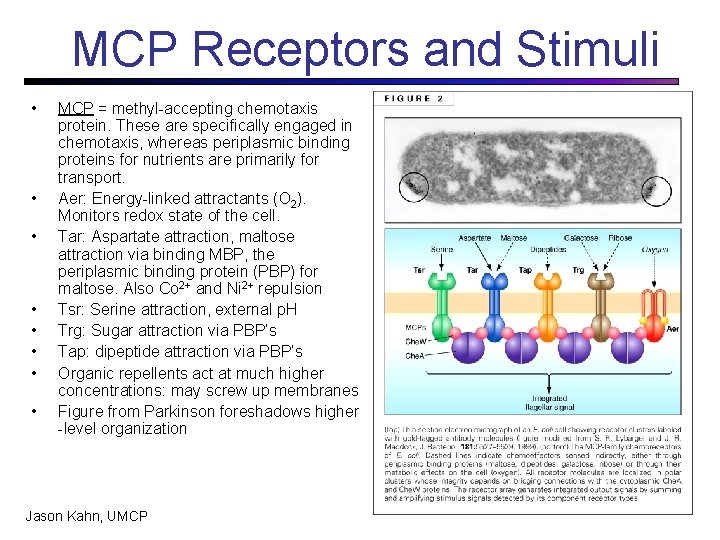
MCP Receptors and Stimuli • • MCP = methyl-accepting chemotaxis protein. These are specifically engaged in chemotaxis, whereas periplasmic binding proteins for nutrients are primarily for transport. Aer: Energy-linked attractants (O 2). Monitors redox state of the cell. Tar: Aspartate attraction, maltose attraction via binding MBP, the periplasmic binding protein (PBP) for maltose. Also Co 2+ and Ni 2+ repulsion Tsr: Serine attraction, external p. H Trg: Sugar attraction via PBP’s Tap: dipeptide attraction via PBP’s Organic repellents act at much higher concentrations: may screw up membranes Figure from Parkinson foreshadows higher -level organization Jason Kahn, UMCP

Che Proteins: Che. A • Che = Chemotaxis genes. (Mot = motility, and many other flagellum mutants are defective in chemotaxis as well. ) • Chemotaxis is a classic bacterial two-component signalling system. • Is general, these systems have a histidine kinase sensor protein and a response regulator that is controlled by phosphorylation via phosphotransfer from the kinase • The chemotaxis genes are Che. A, Che. W, Che. Y, Che. Z, Che. R, and Che. B. • Extensively studied! • Che. A histidine kinase is a dimer in solution. It autophosphorylates using phosphate from ATP: Che. A + ATP Che. A~P + ADP (on His 48) • Che. A auto-phosphorylation is enhanced by Che. W Jason Kahn, UMCP
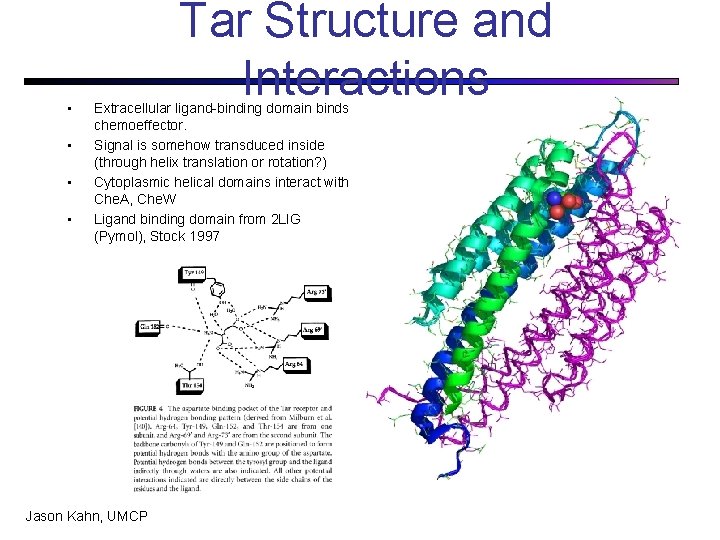
• • Tar Structure and Interactions Extracellular ligand-binding domain binds chemoeffector. Signal is somehow transduced inside (through helix translation or rotation? ) Cytoplasmic helical domains interact with Che. A, Che. W Ligand binding domain from 2 LIG (Pymol), Stock 1997 Jason Kahn, UMCP
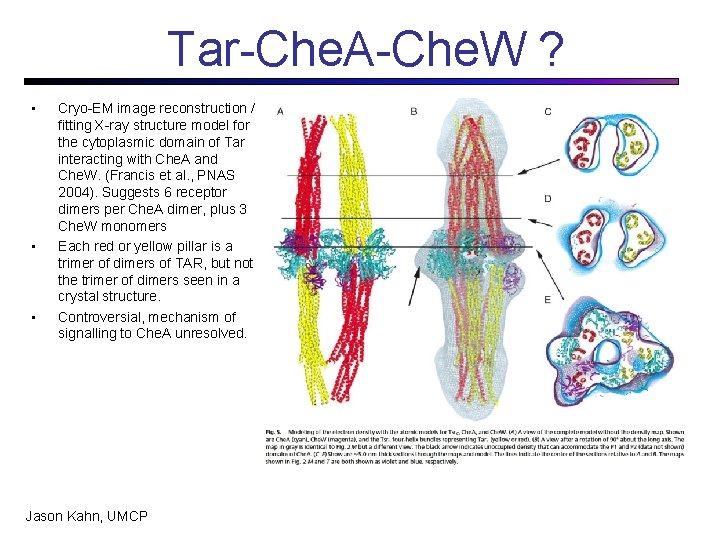
Tar-Che. A-Che. W ? • • • Cryo-EM image reconstruction / fitting X-ray structure model for the cytoplasmic domain of Tar interacting with Che. A and Che. W. (Francis et al. , PNAS 2004). Suggests 6 receptor dimers per Che. A dimer, plus 3 Che. W monomers Each red or yellow pillar is a trimer of dimers of TAR, but not the trimer of dimers seen in a crystal structure. Controversial, mechanism of signalling to Che. A unresolved. Jason Kahn, UMCP

Signal Transduction: Che. Y~P • Phosphorylated Che. A transfers its phosphate to Che. Y, the response regulator: Che. A~P + Che. Y Che. A + Che. Y~P (on Asp 57) • Che. Y~P interacts with Fli. M and other flagellar proteins to switch the motor to CW rotation: tumbling. • Thus inactivation of Che. A upon attractant binding to MCP’s leads to more swimming and less tumbling. Jason Kahn, UMCP

Stimulation: Control of Che. Y • Che. Y has an autophosphatase activity stimulated by Che. Z: Che. Z Che. Y~P + H 2 O Che. Y + Pi • So the steady-state level of Che. Y~P leads to the swimming / tumbling random walk, and the steady-state level changes in the presence of chemoeffectors. • Che. Z may be unregulated, may interact with Che. A. • This is a rapid response (< 1 sec) Jason Kahn, UMCP

Adaptation: Methylation • • • Adaptation means that bacteria stop responding to chemoeffectors when their concentrations do not change. They have a memory of the concentration over the previous few seconds. Hence adaptation is slower than excitation, allowing for a brief episode of biased random walking after a change in chemoeffector level. Adaptation is mediated by the Che. R methyltransferase and the Che. B methylesterase. Che. R methylates the MCP’s using (Tar, Tsr, et al. ). MCP-Glu + S-Adenosyl-Methionine (SAM) MCP-Glu-CH 3 + Adenosyl-homocysteine Methylated MCP’s are less responsive to attractant: Che. A is not inactivated, Che. Y continues to be phosphorylated, bacteria continue to tumble. There are 4 or 5 methylation sites per MCP, and if they occur in clusters this means there are many possible methylation states for a cluster of receptors: wide dynamic range. Che. R is slow, leading to the delay time in adaptation. Jason Kahn, UMCP

Adaptation II • Che. B methylesterase removes methyl groups from MCP’s: MCP-Glu-CH 3 + H 2 O MCP-Glu + CH 3 OH • Allows system to reset to be responsive to low concentrations of attractants again. • Che. B is activated by phosphorylation by Che. A. • Thus, Che. A causes immediate excitation via phosphorylation of Che. Y, but also initiates a slower negative feedback loop that reduces the sensitivity of the MCP’s that regulate it! • MCP’s have allosteric responses to attractants that make them more susceptible to methylation: damping. • Che. B auto-dephosphorylates slowly, perhaps unregulated. Jason Kahn, UMCP

Overall Summary of Circuitry Jason Kahn, UMCP
- Slides: 17