Bacterial Cell Structure Assist Prof Emrah Ruh NEU

Bacterial Cell Structure Assist. Prof. Emrah Ruh NEU Faculty of Medicine Department of Medical Microbiology
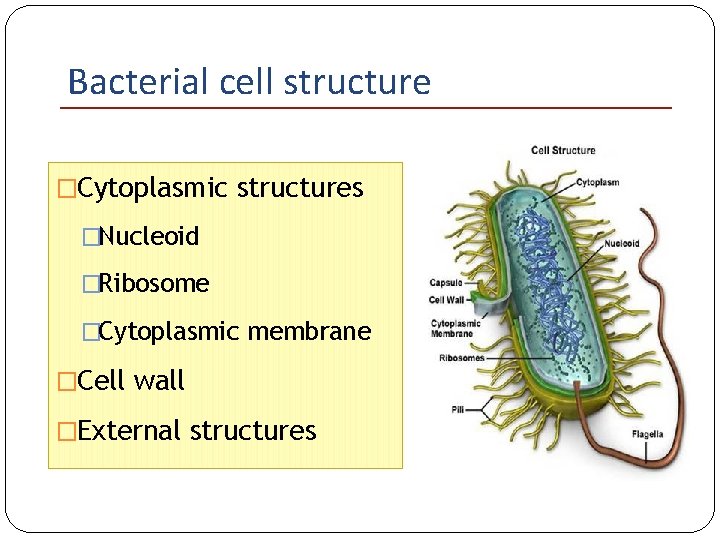
Bacterial cell structure �Cytoplasmic structures �Nucleoid �Ribosome �Cytoplasmic membrane �Cell wall �External structures
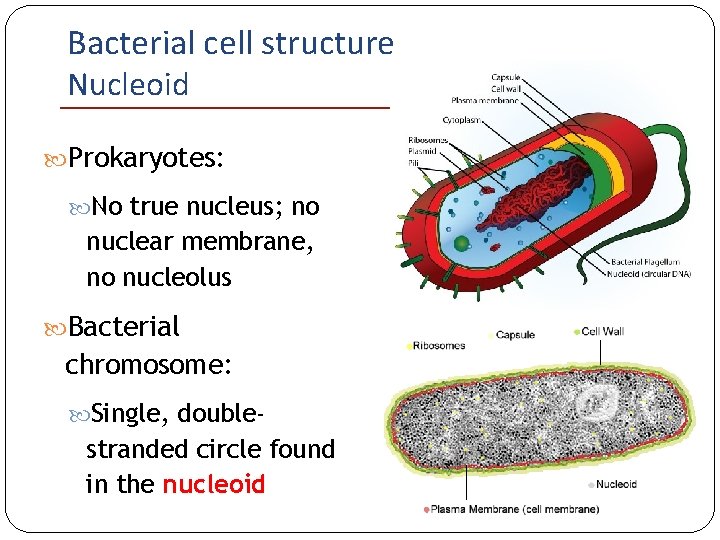
Bacterial cell structure Nucleoid Prokaryotes: No true nucleus; no nuclear membrane, no nucleolus Bacterial chromosome: Single, double- stranded circle found in the nucleoid
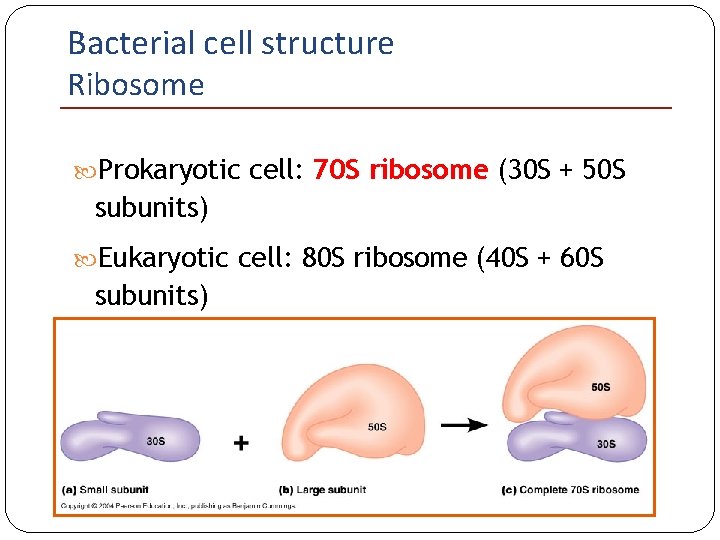
Bacterial cell structure Ribosome Prokaryotic cell: 70 S ribosome (30 S + 50 S subunits) Eukaryotic cell: 80 S ribosome (40 S + 60 S subunits)
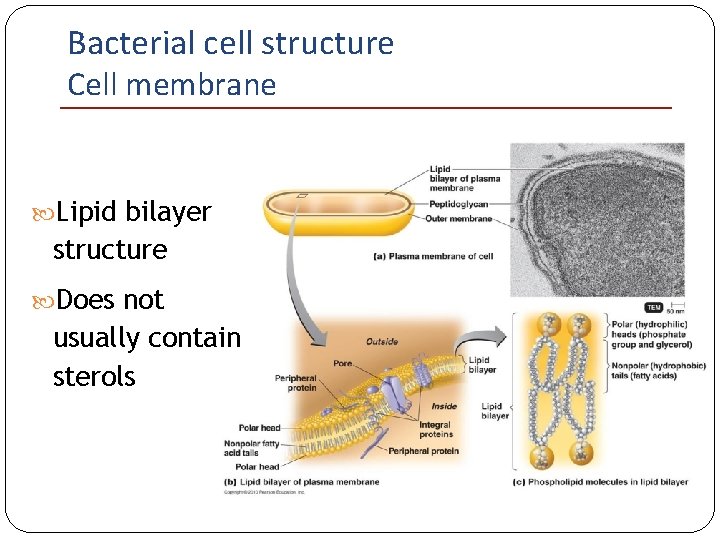
Bacterial cell structure Cell membrane Lipid bilayer structure Does not usually contain sterols

Bacterial cell structure Cell membrane – Functions �Selective permeability and transport of solutes �Electron transport and oxidative phosphorylation �Excretion of hydrolytic exoenzymes �Functioning in DNA and cell wall synthesis �Bearing the receptors of the chemotactic and other sensory transduction systems
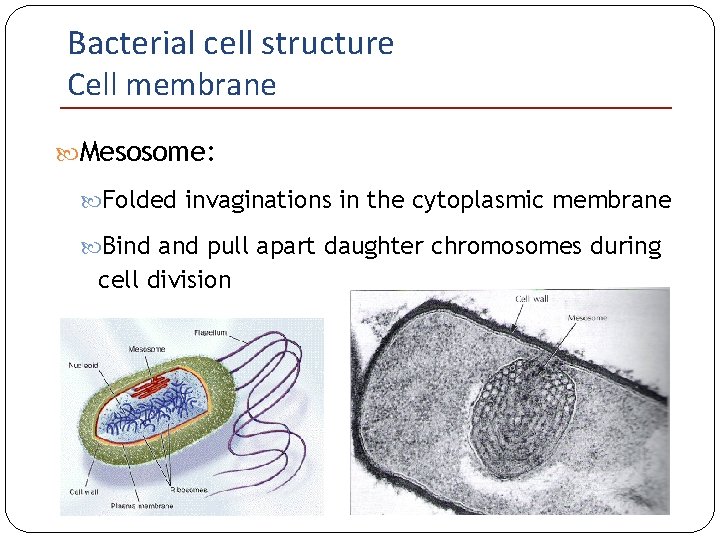
Bacterial cell structure Cell membrane Mesosome: Folded invaginations in the cytoplasmic membrane Bind and pull apart daughter chromosomes during cell division
Bacterial cell structure Cell membrane – Transport of substances 1. Passive transport • Simple diffusion, faciliated diffusion, channel proteins 2. Active transport • Ion-coupled transport, ATP-binding cassette (ABC) transport 3. Group translocation 4. Special transport process
Bacterial cell structure Cell membrane – Transport of substances 1. Passive transport: (Diffusion; no energy) A. Simple diffusion: Not selective; (eg, dissolved O 2, CO 2, and H 2 O) B. Faciliated diffusion: Selective C. Channel proteins: Rare in prokaryotes; selective channels passage of specific molecules (eg, glycerol)

Bacterial cell structure Cell membrane – Transport of substances 1. Passive transport: (Diffusion; no energy)

Bacterial cell structure Cell membrane – Transport of substances 2. Active transport: A. Ion-coupled transport: I. Uniport: single transport of a solute II. Symport: cotransport of a solute and H+ in same direction III. Antiport: transport of two similar solutes in opposite directions

Bacterial cell structure Cell membrane – Transport of substances 2. Active transport: A. Ion-coupled transport

Bacterial cell structure Cell membrane – Transport of substances 2. Active transport: B. ATP-binding cassette (ABC) transport: �Uses ATP �Binding proteins �Gram-negative �Gram-positive bacteria periplasmic space bacteria outer surface of the cell membrane

Bacterial cell structure Cell membrane – Transport of substances 2. Active transport: B. ATP-binding cassette (ABC) transport: �Bound substrate is transferred to a membrane- bound protein complex �Hydrolysis of ATP �Energy membrane pore open movement of substrate into the cell

Bacterial cell structure Cell membrane – Transport of substances 2. Active transport: B. ATP-binding cassette (ABC) transport

Bacterial cell structure Cell membrane – Transport of substances 3. Group translocation: Uptake of certain sugars (glucose, mannose…) Not active transport Phosphotransferase system: Membrane carrier protein phosphorylated (phosphoenolpyruvate) binds the free sugar transport into the cytoplasm: sugar phosphate

Bacterial cell structure Cell membrane – Transport of substances 3. Group translocation:

Bacterial cell structure Cell membrane – Transport of substances 4. Special transport process: Iron (Fe): essential nutrient for bacteria Siderophores transport Fe into the cell

Bacterial cell structure Cell membrane – Transport of substances 4. Special transport process:
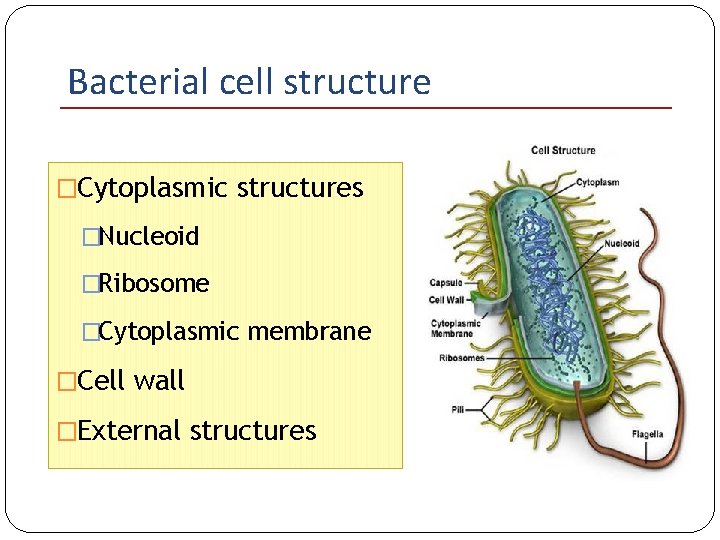
Bacterial cell structure �Cytoplasmic structures �Nucleoid �Ribosome �Cytoplasmic membrane �Cell wall �External structures

Bacterial cell structure Cell wall Distinguish Gram-positive from Gram- negative bacteria Most prokaryotes Peptidoglycan (murein) layer Rigidity, and the shape of the bacterial cell

Bacterial cell structure Cell wall

Bacterial cell structure Cell wall �Gram-positive �Gram-negative bacteria �Peptidoglycan �Teichoic acid �Lipoteichoic acid �Periplasmic space �Outer membrane �Proteins �Lipopolysaccharide

Bacterial cell structure Cell wall – Peptidoglycan �Gram-positive bacteria: �Thick peptidoglycan; ~40 sheets (50% of the cell wall) �Gram-negative bacteria: �Thin peptidoglycan; 1 or 2 sheets (5 -10% of the cell wall)

Bacterial cell structure Cell wall – Peptidoglycan �Peptidoglycan layer; �Provides the strength to the bacterial cell wall �Provides the osmotic stability to the bacterial cell �Synonyms: �Murein, mucopeptide

Bacterial cell structure Cell wall – Peptidoglycan Lysozyme (enzyme in tears, saliva and nasal secretions) degrades the peptidoglycan Hypotonic media Osmotic pressure differences water flows into the cell lysis

Bacterial cell structure Cell wall – Peptidoglycan Isotonic media: Gram-positive bacteria Lysozyme Protoplasts Gram-negative bacteria EDTA-lysozyme Spheroplasts If protoplasts/spheroplasts are able to grow and divide, they are called L-forms.
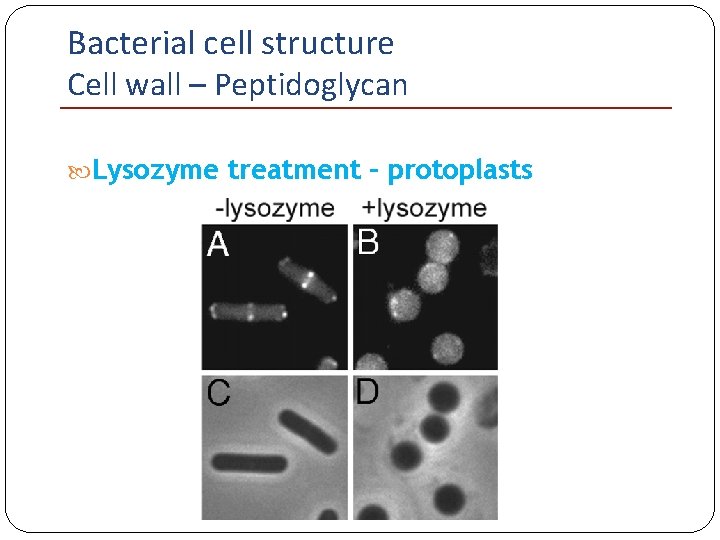
Bacterial cell structure Cell wall – Peptidoglycan Lysozyme treatment – protoplasts
Bacterial cell structure Cell wall – Peptidoglycan synthesis �Peptidoglycan: �Glycan portion: �N-acetylglucosamine (Glc. NAc, NAG) �N-acetylmuramic acid (Mur. NAc, NAM) �Peptide portion: �Tetrapeptide side chains �Peptide cross-bridges
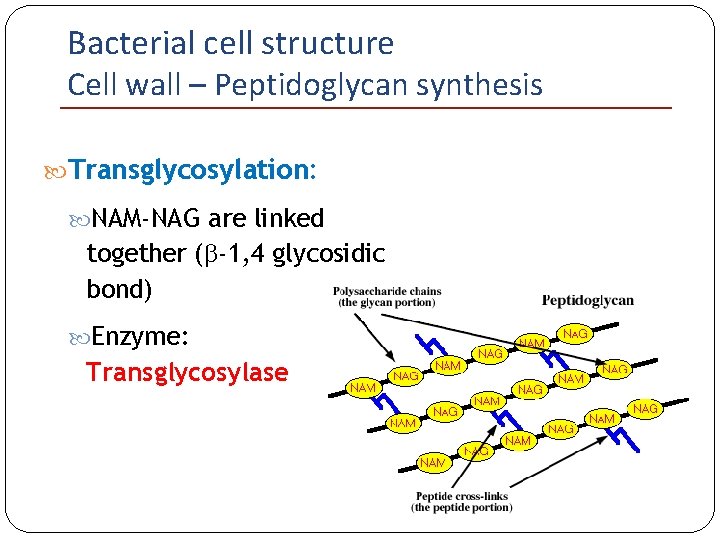
Bacterial cell structure Cell wall – Peptidoglycan synthesis Transglycosylation: NAM-NAG are linked together (b-1, 4 glycosidic bond) Enzyme: Transglycosylase

Bacterial cell structure Cell wall – Peptidoglycan synthesis Transpeptidation: Peptide chains attach to N-acetylmuramic acid Enzyme: Transpeptidase
Bacterial cell structure Cell wall – Peptidoglycan synthesis Transpeptidation: Pentapeptide: L-Alanine (L-Ala) D-Glutamate (D-Glu) L-Lysine (L-Lys) / Diaminopimelic acid (DAP) D-Alanine (D-Ala)
Bacterial cell structure Cell wall – Peptidoglycan synthesis Transpeptidation: Pentapeptide: The terminal D-Alanine (D-Ala) from pentapeptide is removed by transpeptidase and carboxypeptidase enzymes In mature peptidoglycan, peptide chains are tetrapeptides

Bacterial cell structure Cell wall – Peptidoglycan synthesis �Transglycosylase, transpeptidase and carboxypeptidase enzymes are called “Penicillin-binding proteins (PBPs)”
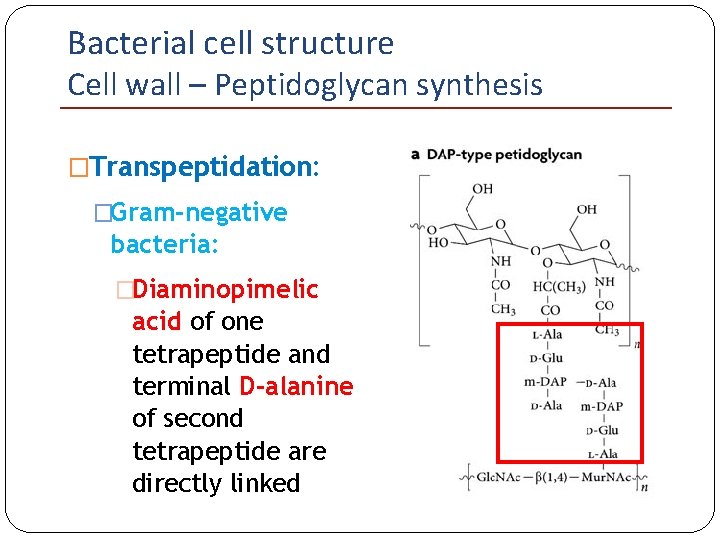
Bacterial cell structure Cell wall – Peptidoglycan synthesis �Transpeptidation: �Gram-negative bacteria: �Diaminopimelic acid of one tetrapeptide and terminal D-alanine of second tetrapeptide are directly linked
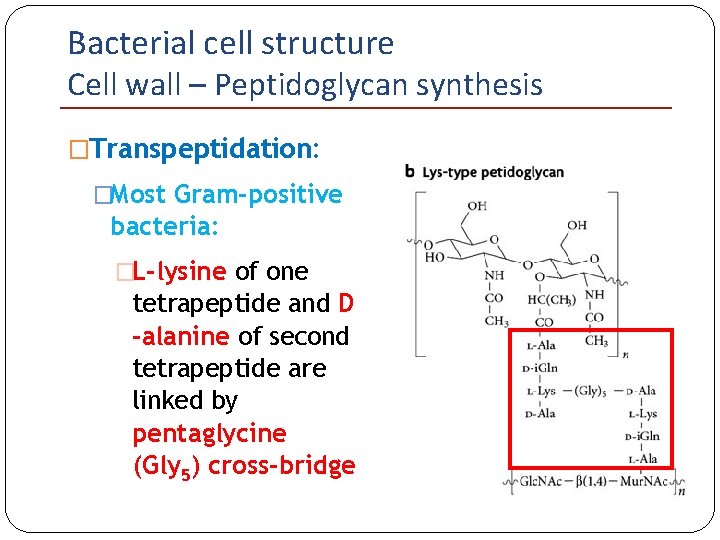
Bacterial cell structure Cell wall – Peptidoglycan synthesis �Transpeptidation: �Most Gram-positive bacteria: �L-lysine of one tetrapeptide and D -alanine of second tetrapeptide are linked by pentaglycine (Gly 5) cross-bridge
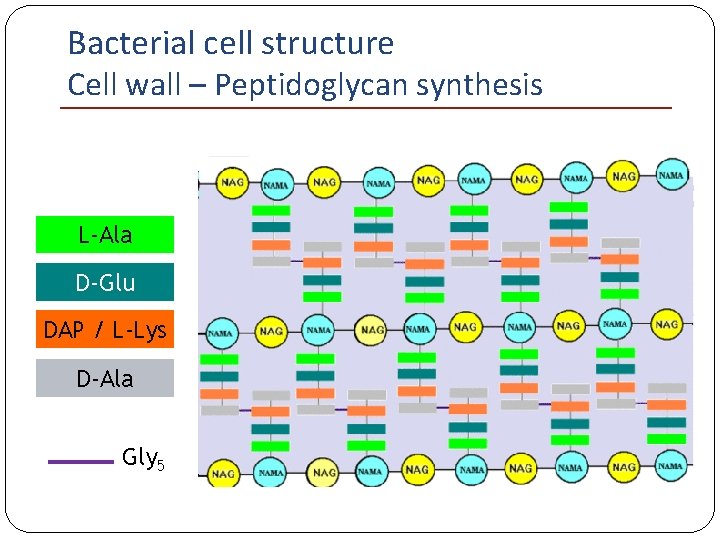
Bacterial cell structure Cell wall – Peptidoglycan synthesis L-Ala D-Glu DAP / L-Lys D-Ala Gly 5

Bacterial cell structure Cell wall – Teichoic acid / Lipoteichoic acid Gram-positive bacteria Common surface antigens (distinguish bacterial serotypes) Attachment (adherence) to the host cell Important factors in virulence Teichoic acids peptidoglycan Lipoteichoic acids cytoplasmic membrane

Bacterial cell structure Cell wall – Teichoic acid / Lipoteichoic acid

Bacterial cell structure Cell wall – Periplasmic space Gram-negative bacteria The space between the inner and outer membranes Contains the peptidoglycan layer Possess hydrolytic enzymes (proteases, lipases, . . . ) (breakdown of large molecules for metabolism)
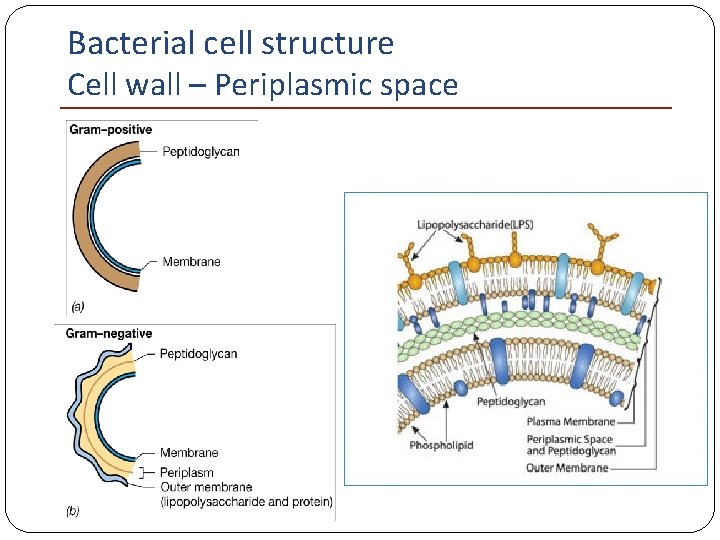
Bacterial cell structure Cell wall – Periplasmic space

Bacterial cell structure Cell wall – Outer membrane Gram-negative bacteria Bilayered structure: Inner leaflet similar composition with the cell membrane Outer leaflet contains lipopolysaccharide (LPS)

Bacterial cell structure Cell wall – Outer membrane

Bacterial cell structure Cell wall – Outer membrane Exclude hydrophobic molecules (unusual feature among biological membranes!) Protect the bacterium (eg, digestive system) Connected to both the peptidoglycan layer and the cytoplasmic membrane Lipoprotein: connects peptidoglycan with outer membrane

Bacterial cell structure Cell wall – Outer membrane Possess special channels (porins): passive diffusion of low-molecular-weight hydrophilic compounds (sugars, amino acids and ions) Large antibiotic molecules penetrate slowly: antibiotic resistance!
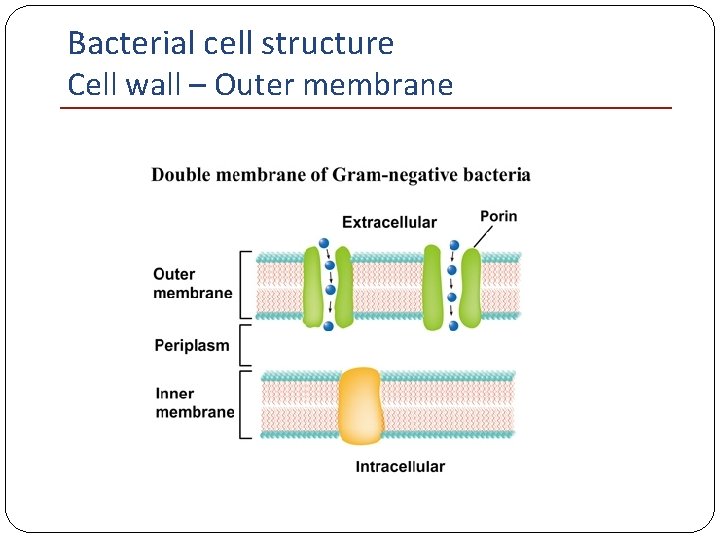
Bacterial cell structure Cell wall – Outer membrane

Bacterial cell structure Cell wall – Lipopolysaccharide �Lipopolysaccharide (LPS) Endotoxin of Gram-negative bacteria
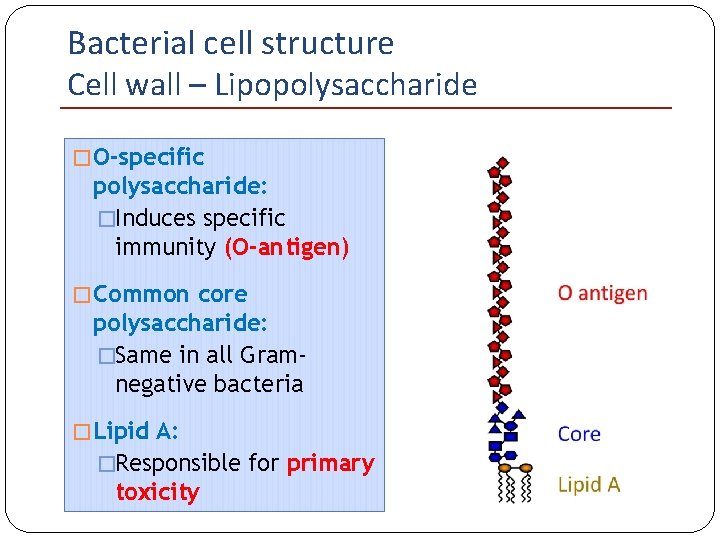
Bacterial cell structure Cell wall – Lipopolysaccharide � O-specific polysaccharide: �Induces specific immunity (O-antigen) � Common core polysaccharide: �Same in all Gramnegative bacteria � Lipid A: �Responsible for primary toxicity
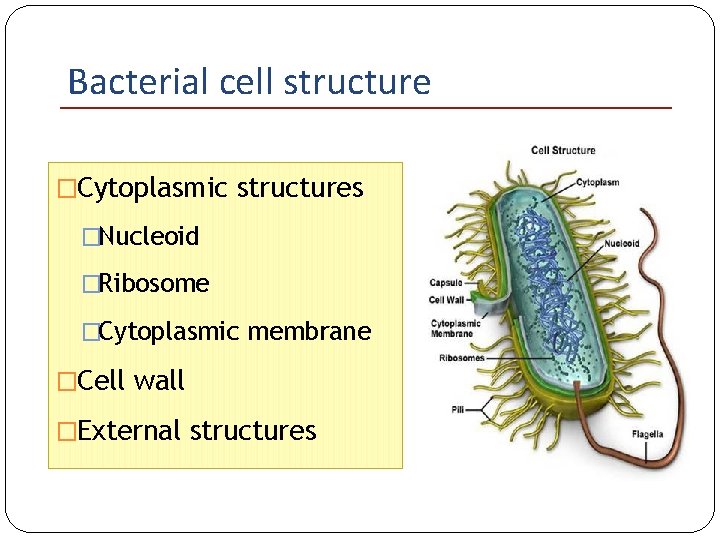
Bacterial cell structure �Cytoplasmic structures �Nucleoid �Ribosome �Cytoplasmic membrane �Cell wall �External structures

Bacterial cell structure External structures Capsule/slime layer Flagella Fimbriae (pili)

Bacterial cell structure External structures – Capsule/slime layer Distinct bacterial species Polysaccharide Capsule of Bacillus anthracis Polypeptide (poly-D-glutamic acid)

Bacterial cell structure External structures – Capsule/slime layer Capsule condensed layer; closely surrounds the bacterium Slime layer loosely adherent; nonuniform in density and thickness Capsule/slime layer: also called glycocalyx
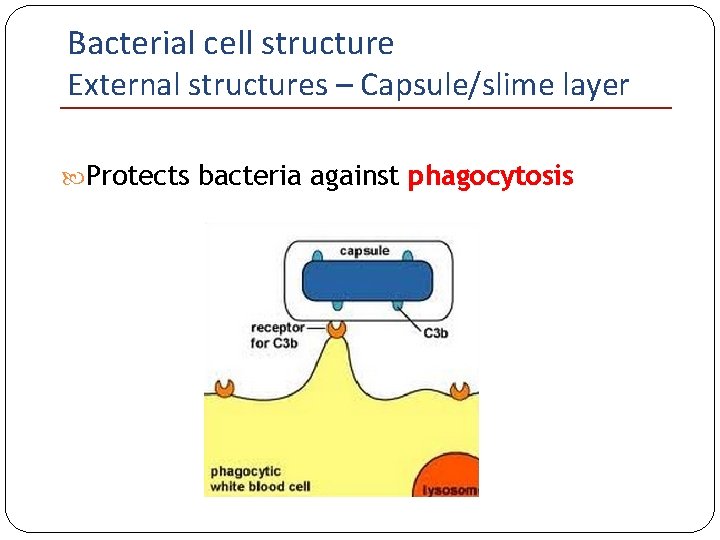
Bacterial cell structure External structures – Capsule/slime layer Protects bacteria against phagocytosis

Bacterial cell structure External structures – Capsule/slime layer Plays a role in adherence (biofilm formation) Artificial valves, catheters, …
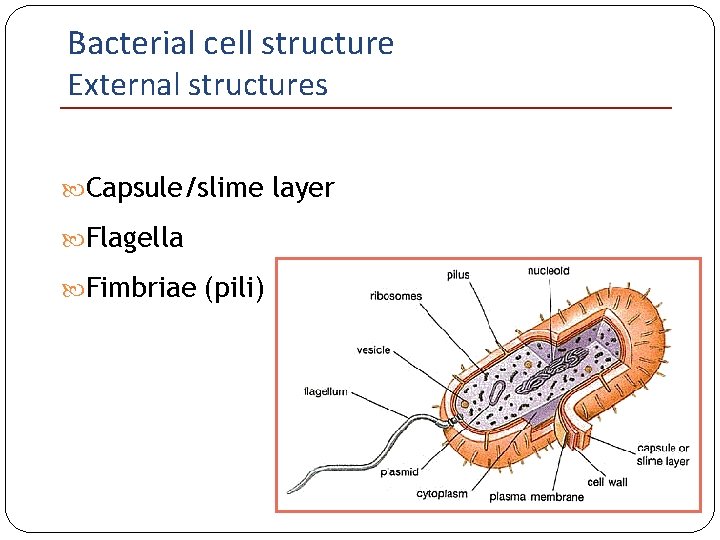
Bacterial cell structure External structures Capsule/slime layer Flagella Fimbriae (pili)

Bacterial cell structure External structures – Flagella

Bacterial cell structure External structures – Flagella �Thread-like appendages �Composed of protein subunits called flagellin �Subunits aggregate and form a helical structure �Highly antigenic (H antigens) �If removed by mechanical agitating; new flagella are rapidly formed and motility is rapidly restored

Bacterial cell structure External structures – Flagella Types of arrangements of flagella: Monotrichous single polar flagellum Amphitrichous single polar flagella – opposites Lophotrichous multiple polar flagella – opposites Peritrichous flagella distributed over the entire cell
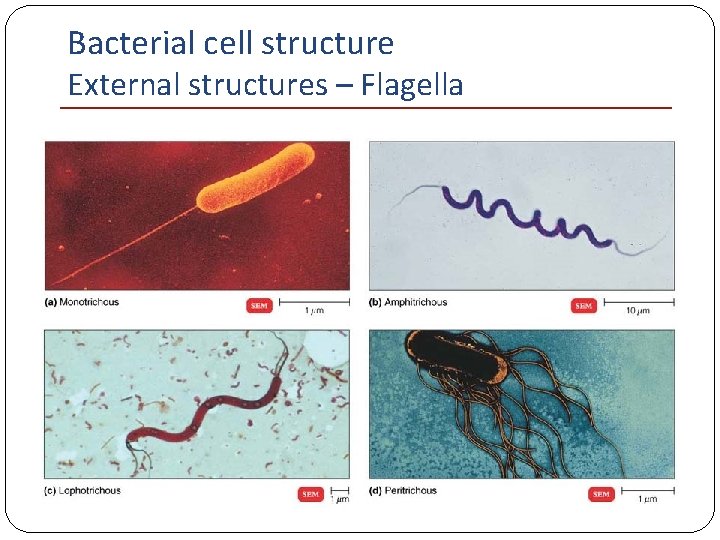
Bacterial cell structure External structures – Flagella
Bacterial cell structure External structures – Flagella Hook: Short curved structure Joint between the basal body and the flagellum Basal body: Set of rings 1 pair (M-S ring) in Gram- positive bacteria; 2 pairs (L-P, M-S) in Gramnegative bacteria
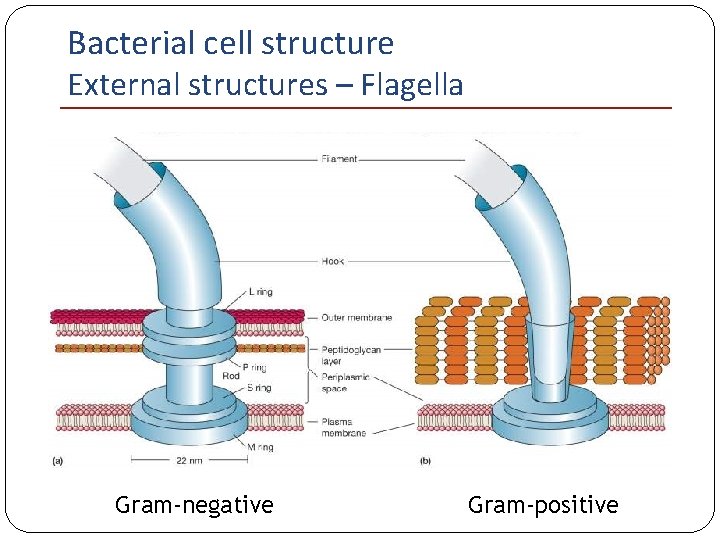
Bacterial cell structure External structures – Flagella Gram-negative Gram-positive

Bacterial cell structure External structures – Flagella
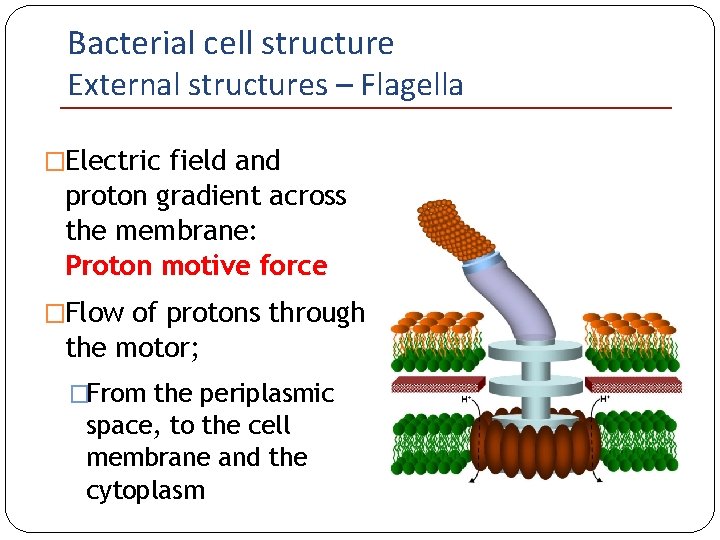
Bacterial cell structure External structures – Flagella �Electric field and proton gradient across the membrane: Proton motive force �Flow of protons through the motor; �From the periplasmic space, to the cell membrane and the cytoplasm
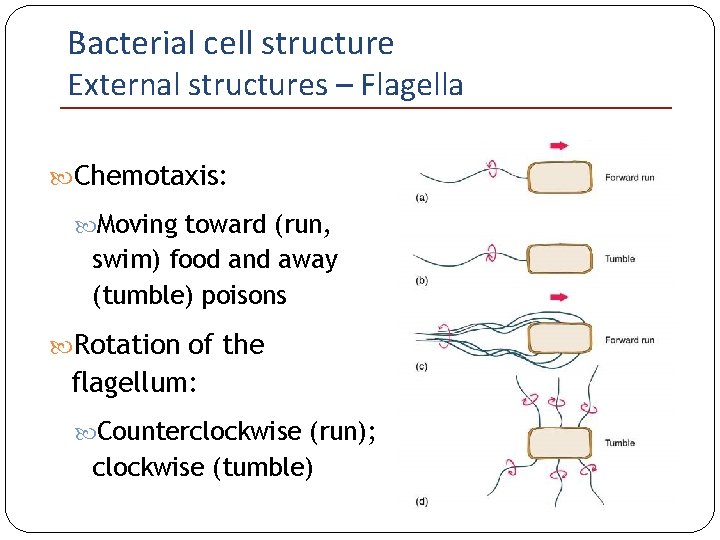
Bacterial cell structure External structures – Flagella Chemotaxis: Moving toward (run, swim) food and away (tumble) poisons Rotation of the flagellum: Counterclockwise (run); clockwise (tumble)
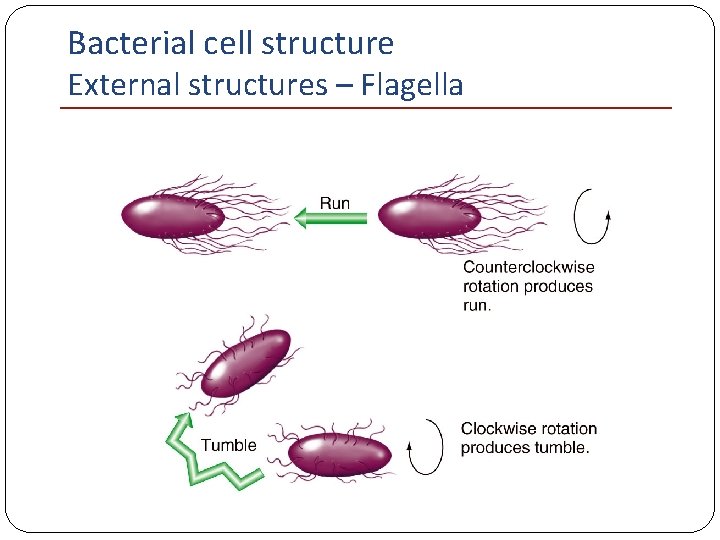
Bacterial cell structure External structures – Flagella

Bacterial cell structure External structures Capsule/slime layer Flagella Fimbriae (pili)
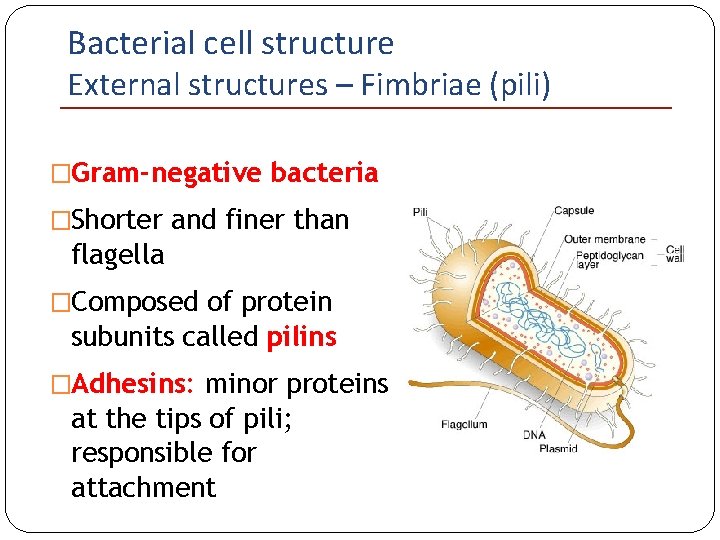
Bacterial cell structure External structures – Fimbriae (pili) �Gram-negative bacteria �Shorter and finer than flagella �Composed of protein subunits called pilins �Adhesins: minor proteins at the tips of pili; responsible for attachment
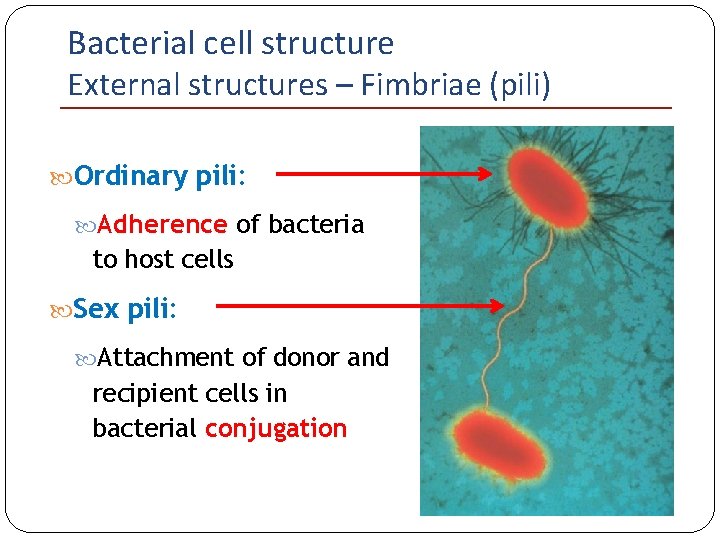
Bacterial cell structure External structures – Fimbriae (pili) Ordinary pili: Adherence of bacteria to host cells Sex pili: Attachment of donor and recipient cells in bacterial conjugation

Bacterial cell structure Endospores (spores) �Distinct bacterial genera; the most commons: �Bacillus (Gram-positive aerobic rod) �Clostridium (Gram- positive anaerobic rod) �Response to environmental conditions (depletion of nutrients)

Bacterial cell structure Endospores (spores) Cannot be stained with Gram (seen colourless) Bacillus Spore Clostridium
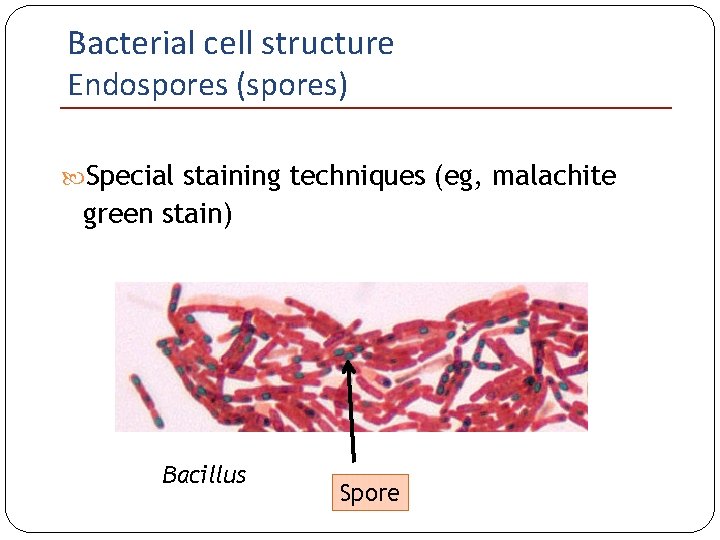
Bacterial cell structure Endospores (spores) Special staining techniques (eg, malachite green stain) Bacillus Spore

Bacterial cell structure Endospores (spores) �Sporulation formation of spore �Liberated when the mother cell undergoes autolysis �Resting cell; highly resistant to desiccation, heat, and chemical agents �Germination favorable nutritional conditions; activation of spore to produce a single vegetative cell
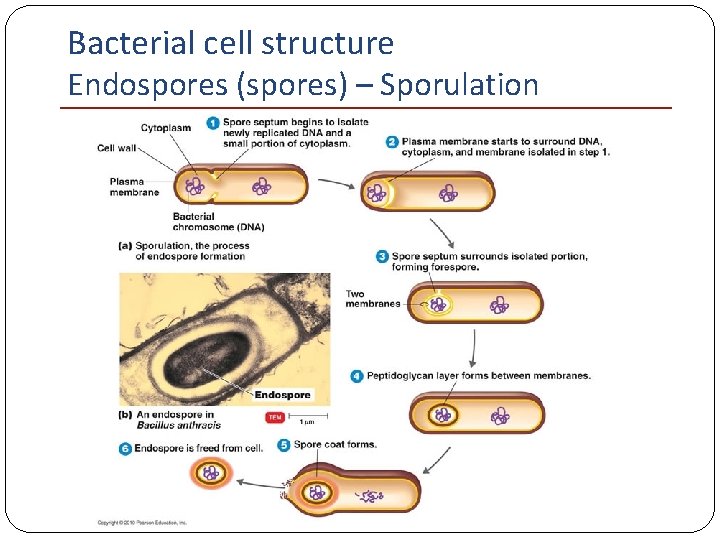
Bacterial cell structure Endospores (spores) – Sporulation

Bacterial cell structure Endospores (spores) Parts of spores (from in to outwards): Core Spore wall Cortex Coat Exosporium Core

Bacterial cell structure Endospores (spores) – Core The core: is the spore protoplast Contains chromosome Does not contain ATP (energy of germination is stored as 3 -phosphoglycerate) Heat resistance of spores; Dehydrated state Calcium dipicolinate found in the core

Bacterial cell structure Endospores (spores) – Spore wall: Contains normal peptidoglycan Becomes the cell wall of the germinating vegetative cell

Bacterial cell structure Endospores (spores) – Cortex: �The thickest layer of the spore envelope �Contains an unusual type of peptidoglycan �Fewer cross-links �Extremely sensitive to lysozyme �Autolysis plays a role in spore germination

Bacterial cell structure Endospores (spores) – Coat: Composed of a keratin-like protein Impermeable resistance of spores against antibacterial chemical agents

Bacterial cell structure Endospores (spores) – Exosporium: The outermost layer of spore envelope Composed of proteins, lipids, and carbohydrates Consists of a paracrystalline basal layer and a hairlike outer region

Bacterial cell structure Classification Gram-staining feature Gram-positive Gram-negative Morphology Coccus, bacillus, coccobacillus, …

Bacterial cell structure Classification – Gram staining Introduced by Hans Christian Joachim Gram Based on the differences of Gram-positive and Gram-negative cell wall

Bacterial cell structure Classification – Gram staining Basic principle: Crystal violet gets trapped in the thick and cross-linked peptidoglycan of Gram-positive bacteria Gram-negative bacteria (thin peptidoglycan) are easily decolorized by alcohol and do not retain crystal violet
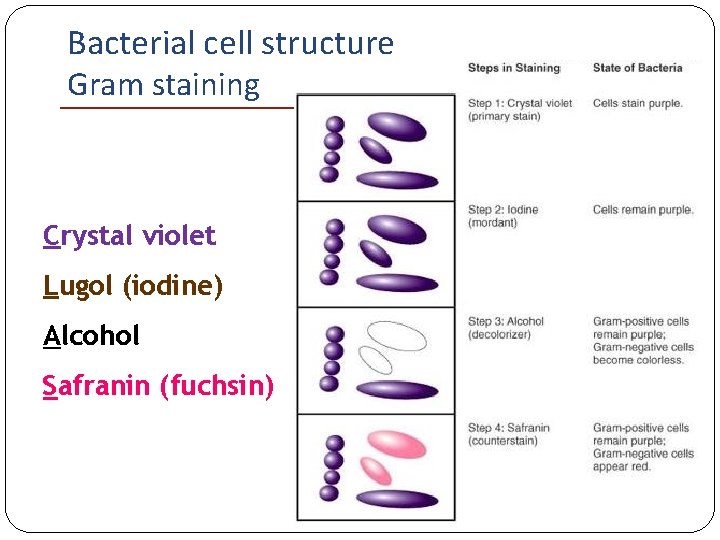
Bacterial cell structure Gram staining Crystal violet Lugol (iodine) Alcohol Safranin (fuchsin)
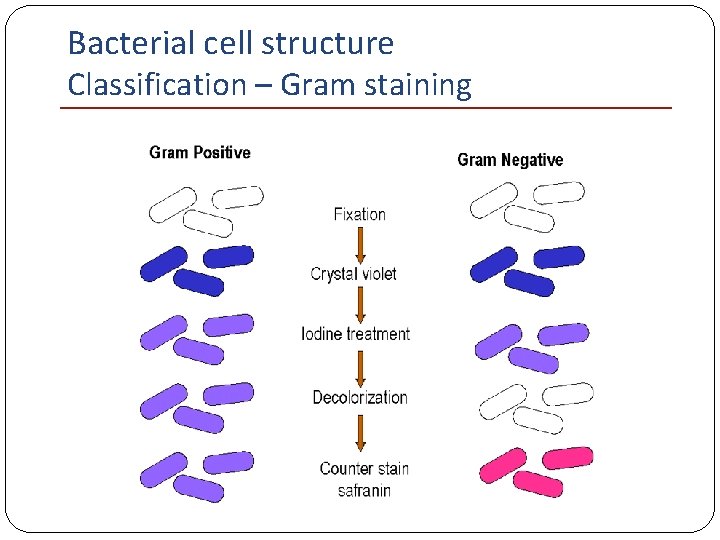
Bacterial cell structure Classification – Gram staining
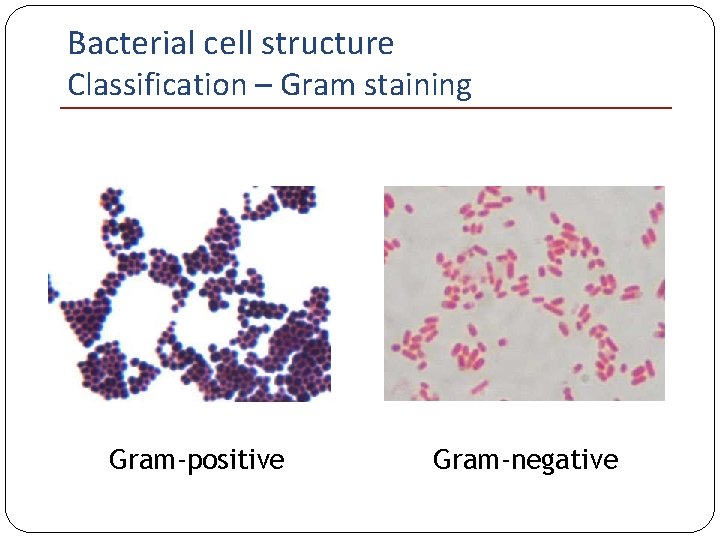
Bacterial cell structure Classification – Gram staining Gram-positive Gram-negative
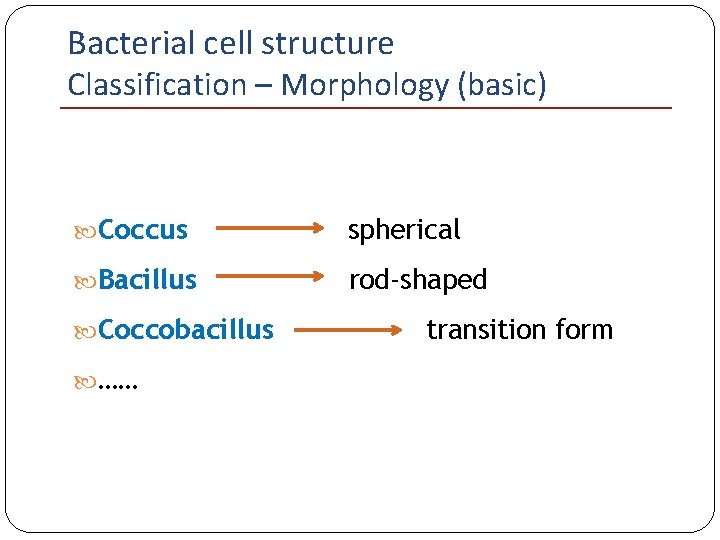
Bacterial cell structure Classification – Morphology (basic) Coccus spherical Bacillus rod-shaped Coccobacillus …… transition form
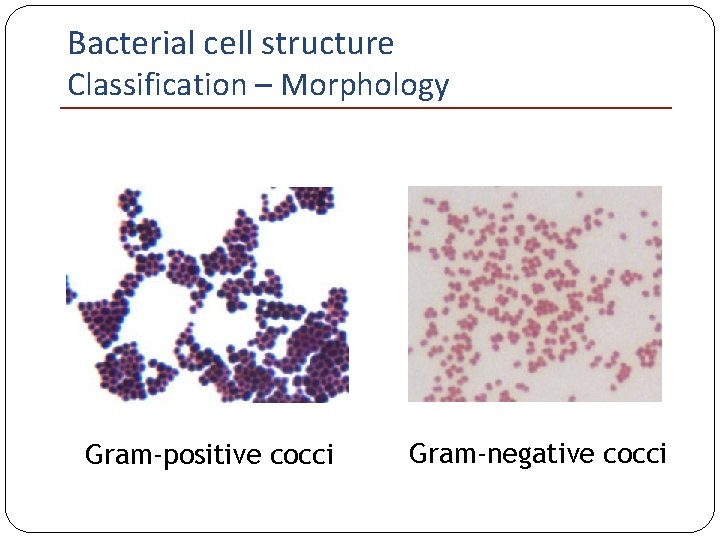
Bacterial cell structure Classification – Morphology Gram-positive cocci Gram-negative cocci

Bacterial cell structure Classification – Morphology Gram-positive bacilli Gram-negative bacilli
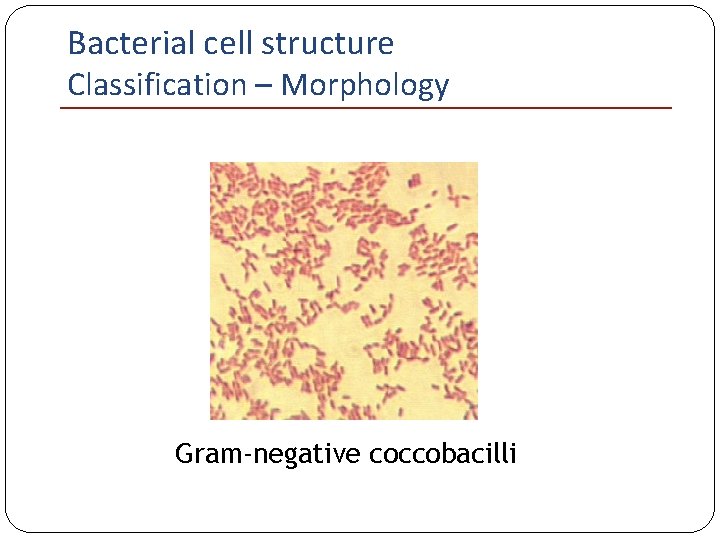
Bacterial cell structure Classification – Morphology Gram-negative coccobacilli
- Slides: 89