Backcross Breeding History of Backcrossing Harlan and Pope

Backcross Breeding

History of Backcrossing • Harlan and Pope, 1922 • Wanted the smooth awns from European barleys in the domestic barleys • Crosses with European types were not fruitful • Decided to backcross smooth awn • After 1 BC, progeny resembled Manchuria and they were able to recover high yielding smooth awn types

Terminology • Recurrent parent (RP) - parent you are transferring trait to • Donor or nonrecurrent parent (DP) source of desirable trait • Progeny test - when trait is recessive
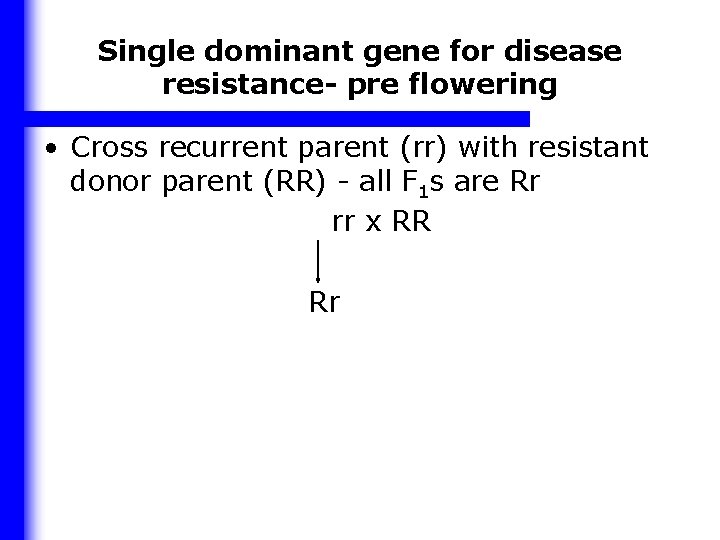
Single dominant gene for disease resistance- pre flowering • Cross recurrent parent (rr) with resistant donor parent (RR) - all F 1 s are Rr rr x RR Rr
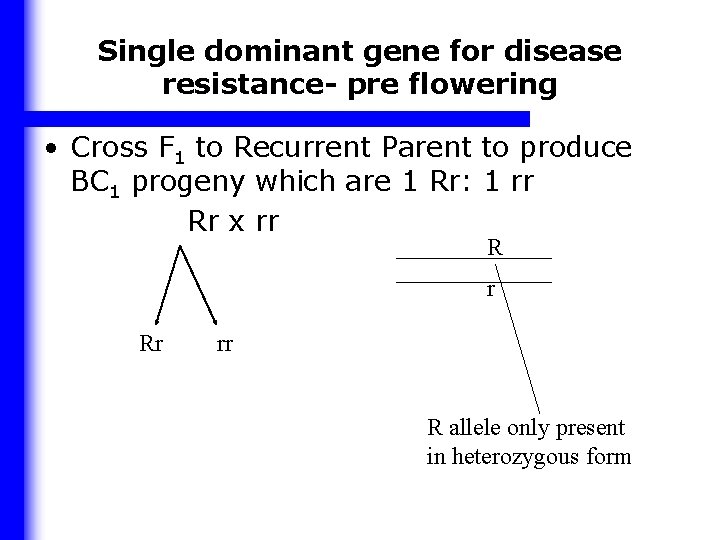
Single dominant gene for disease resistance- pre flowering • Cross F 1 to Recurrent Parent to produce BC 1 progeny which are 1 Rr: 1 rr Rr x rr R r Rr rr R allele only present in heterozygous form

Single dominant gene for disease resistance- pre flowering • Evaluate BC 1 s before flowering and discard rr plants; cross Rr plants to Recurrent Parent Rr – keep rr - discard

Single dominant gene for disease resistance- pre flowering • BC 2 F 1 plants evaluated, rr plants discarded, Rr plants crossed to Recurrent Parent • BC 4 F 1 plants evauated, rr plants discarded, Rr plants selfed to produce BC 4 F 2 seeds, which are 1 RR: 2 Rr: 1 rr

Single dominant gene for disease resistance- pre flowering • BC 4 F 2 plants evaluated before flowering, rr discarded, R_ selfed and harvested by plant, then progeny tested. Segregating rows discarded, homozygous RR rows kept and tested.


Single dominant gene - post flowering • Cross susceptible Recurrent Parent (rr) with resistant Donor Parent (RR) - all F 1 s are Rr rr x RR Rr X rr rr; Rr BC 1

Single dominant gene - post flowering • Cross F 1 to Recurrent Parent to produce BC 1 progeny which are 1 Rr: 1 rr • Because we can’t evaluate the trait before flowering, a number of BC 1 F 1 plants must be crossed to Recurrent Parent, then the trait is evaluated and susceptible plants discarded • This procedure is therefore less efficient than the pre-flowering trait because we have made crosses that we cannot use

Single dominant gene - post flowering • BC 2 F 1 plants (1 Rr: 1 rr) are crossed to RP, trait evaluated before harvest, susceptible plants discarded

Single dominant gene - post flowering • Procedure followed through BC 4 • Seeds from each BC 4 F 2 individual are harvested by plant and planted in rows • Segregating rows are discarded, homozygous RR rows are maintained, harvested and tested further


Single recessive allele progeny test in same season • Cross susceptible (RR) Recurrent Parent to resistant (rr) Donor Parent • F 1 plants crossed to Recurrent Parent, BC 1 seeds are 1 RR: 1 Rr • Note now that all BC 1 plants are susceptible; we are interested only in those plants which carry the resistant “r” allele • All BC 1 plants crossed to Recurrent Parent and selfed to provide seeds for progeny test

Single recessive allele progeny test in same season • Screen BC 1 F 2 plants before BC 2 F 1 plants flower. BC 1 F 1 plants that are RR will have only RR progeny. BC 1 F 1 plants that are Rr will produce BC 1 F 2 progeny that segregate for resistance.

Single recessive allele progeny test in same season • BC 2 F 1 plants from heterozygous (Rr) BC 1 plants are crossed to RP; those from susceptible (RR) BC 1 plants are discarded • BC 2 F 2 selfed seed is harvested for progeny testing • Progeny tests are conducted before BC 3 F 1 plants flower. Only plants from (Rr) BC 2 plants are crossed to Recurrent Parent

Single recessive allele progeny test in same season • Each BC 4 F 1 plant is progeny tested. Progeny from susceptible BC 3 plants are all susceptible and family is discarded • If progeny test completed before flowering, only homozygous resistant (rr) plants are selfed. Otherwise, all plants selfed and only seed from (rr) plants harvested. • Additional testing of resistant families required.


Single recessive allele - progeny test in different season • Cross susceptible (RR) Recurrent Parent to resistant (rr) Donor Parent • F 1 plants crossed to RP, seeds are 1 RR: 1 Rr • Again, we are interested in plants carrying the resistant “r” allele – we can’t distinguish them yet from RR types

Single recessive allele - progeny test in different season • The difference is now that we cannot do the progeny test in the same season because the resistance is expressed late in plant’s life. • BC 1 plants selfed, seed harvested by plant • BC 1 F 2 plants grown in progeny rows, evaluated, seed from resistant (rr) rows is harvested. BC 1 F 3 progeny crossed to Recurrent Parent to produce BC 2 F 1 seeds.

Single recessive allele - progeny test in different season • BC 2 F 1 plants crossed to Recurrent Parent to obtain BC 3 F 1 seeds which are 1 Rr: 1 RR • BC 3 F 1 plants are selfed, and progeny are planted in rows • BC 3 F 2 seeds are harvested from resistant (rr) progeny rows • Resistant BC 3 F 3 plants crossed to RP to produce BC 4 F 1 seeds

Single recessive allele - progeny test in different season • BC 4 F 1 plants selfed and produce 1 RR: 2 Rr: 1 rr progeny • BC 4 F 2 plants selfed and resistant ones harvested by plant • Resistant families tested further


Importance of cytoplasm • For certain traits (e. g. male sterility) it is important that a certain cytoplasm be retained • In wheat, to convert a line to a male sterile version the first cross should be made as follows: Triticum timopheevi (male sterile) x male fertile wheat line. From that point on, the recurrent parent should always be used as the male.

Cytoplasmic male sterility in Wheat Triticum timopheevi x Elite breeding line (Male sterile) (Male fertile) F 1 (female) x RP (male) Carry out for 4 BC; use male-sterile version of elite breeding line as female parent in hybrid

Probability of transferring genes • How many backcross progeny should be evaluated? • Consult table in Fehr, p. 367; for example in backcrossing a recessive gene, to have a 95% probability of recovering at least 1 Rr plant, you need to grow 5 backcross progeny.

Probability of transferring genes • To increase the probability to 99% and the number of Rr plants to 3, you must grow 14 progeny • If germination is only 80%, you must grow 14/0. 8 = 18 progeny

Recovery of genes from RP • Ave. recovery of RP = 1 -(1/2)n+1, where n is the number of backcrosses to RP • The percentage recovery of RP varies among the backcross progeny • For example, in the BC 3, if the Donor Parent and Recurrent Parent differ by 10 loci, 26% of the plants will be homozygous for the 10 alleles of the Recurrent Parent; remainder will vary.

Recovery of genes from Recurrent Parent • Selection for the Recurrent Parent phenotype can hasten the recovery of the Recurrent Parent • If the number of BC progeny is increased, selection for Recurrent Parent can be effective

Linkage Drag • When backcrossing, we often get more than one gene from the donor parent • The additional genes may be undesirable, hence the term linkage drag • Backcrossing provides opportunity for recombination between the favorable gene and the linked unfavorable genes

Linkage Drag • Recombination fraction has a profound impact: with c=0. 5, probability that undesirable gene will be eliminated with 5 BC is 0. 98 • with c=0. 02, probability that undesirable gene will be eliminated with 5 BC is 0. 11

Backcrossing for Quantitative Characters • Choose Donor Parent that differs greatly from Recurrent Parent to increase the likelihood of recovery of desired trait (earliness for example) • Effect of environment on expression of trait can be a problem in BC quantitative traits

Backcrossing for Quantitative Characters • Consider selfing after each BC • Expression of differences among plants will be greater • May be possible to practice selection • Single plant progeny test will not be worthwhile; must use replicated plots

Other Considerations • Marker assisted backcrossing • Assume that you have a saturated genetic map • Make cross and backcross • To hasten the backcrossing process, select against the donor genotype (except for the marker(s) linked to the gene of interest) in backcross progeny

Marker-Assisted Backcrossing • May improve efficiency in three ways: – 1) If phenotyping is difficult – 2) Markers can be used to select against the donor parent in the region outside the target – 3) Markers can be used to select rare progeny that result from recombinations near the target gene

Model Two alleles at marker locus: M 1 and M 2 Two alleles at target gene: Q 1 and Q 2 M 1 M 2 Q 1 R=0. 10 Q 2 is the target allele we want to backcross into recurrent parent, which has Q 1 to begin with.

Recombination • Assume recombination between marker and QTL=10% • Select one plant based on marker genotype alone, 10% chance of losing target gene • Probability of not losing gene=(1 -r) • For t generations, P=1 -( 1 -r )t • For 5 BC generations, probability of losing the target gene is P=1 -(. 9)5=0. 41

Flanking Markers Best way to avoid losing the target gene is to have marker loci flanking it MA 1 MA 2 r. A Q 1 r. B MB 1 MB 2

Flanking Markers Probabilityof losing the target gene after selecting On flanking markers: Example: If the flanking markers have 10% recombination frequency with the target gene: , the probability of losing the gene after 1 generation is P=0. 024. The probability of losing the gene after 5 generations is P=0. 1182

Other Considerations • Backcross breeding is viewed as a conservative approach • The goal is to improve an existing cultivar • Meanwhile, the competition moves past

Backcross Populations • May be used as breeding populations instead of F 2, for example • Studies have shown that the variance in a backcross population can exceed that of an F 2 • Many breeders use 3 -way crosses, which are similar to backcrosses
Marker Assisted BC
- Slides: 43