Bacillus Phage Annotation Phage Lab II Spring 2015

Bacillus Phage Annotation Phage Lab II, Spring 2015

Complete annotation of other phages, explore those genomes Claudi (Demetrius) Nemo (Sathya & Molly) Didn’t annotate yet Jugalone (Herleen) Zuko(Sailasya) DIGNKC (Ryan) +
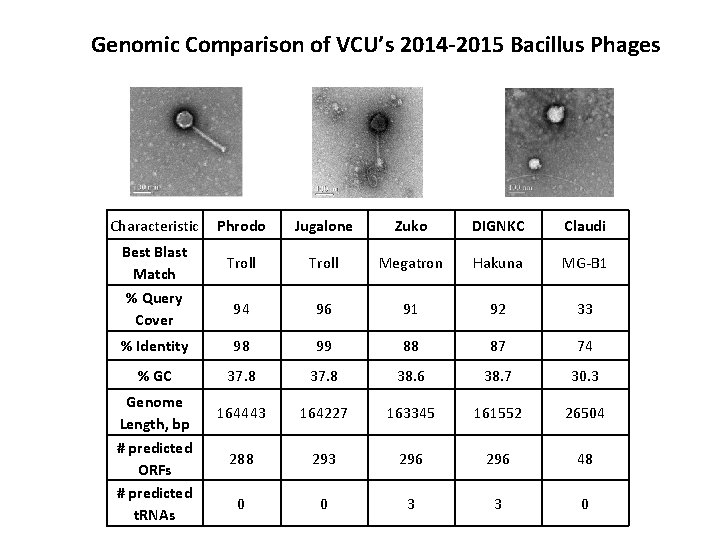
Genomic Comparison of VCU’s 2014 -2015 Bacillus Phages Characteristic Phrodo Jugalone Zuko DIGNKC Claudi Troll Megatron Hakuna MG-B 1 94 96 91 92 33 % Identity 98 99 88 87 74 % GC 37. 8 38. 6 38. 7 30. 3 164443 164227 163345 161552 26504 288 293 296 48 0 0 3 3 0 Best Blast Match % Query Cover Genome Length, bp # predicted ORFs # predicted t. RNAs
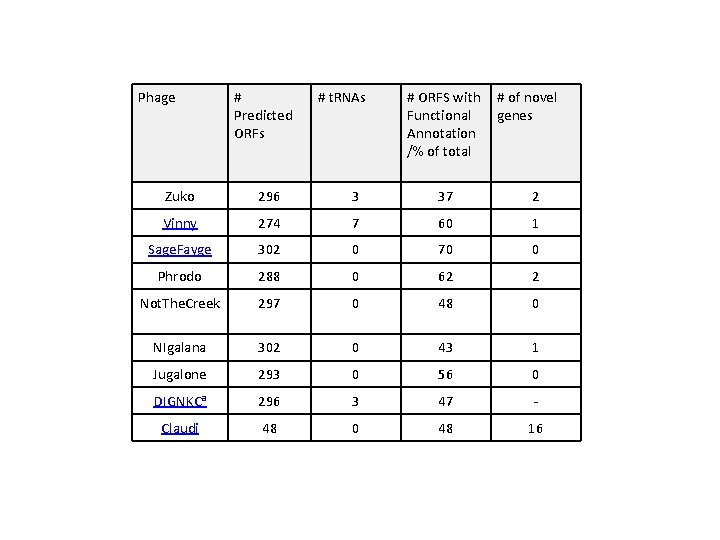
Phage # Predicted ORFs # t. RNAs # ORFS with Functional Annotation /% of total # of novel genes Zuko 296 3 37 2 Vinny 274 7 60 1 Sage. Fayge 302 0 70 0 Phrodo 288 0 62 2 Not. The. Creek 297 0 48 0 NIgalana 302 0 43 1 Jugalone 293 0 56 0 DIGNKCa 296 3 47 - Claudi 48 0 48 16
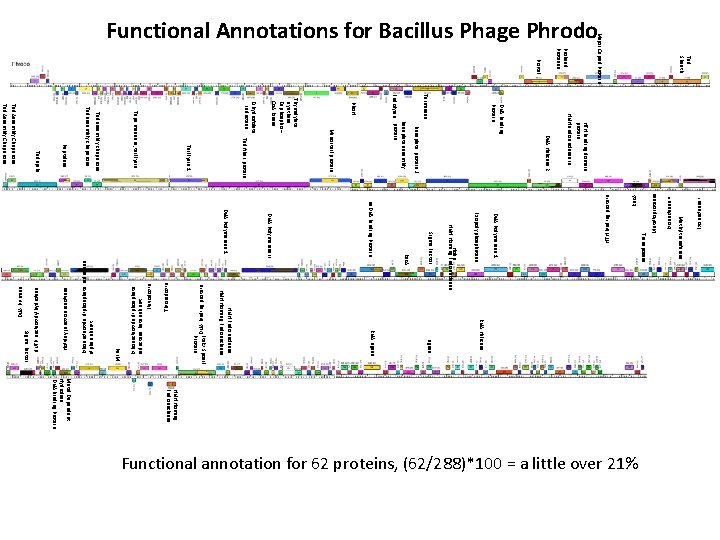
DNA Binding protein Thioredoxin Ribonucleotide diphosphate reductase beta subunit Flavodoxin Ribonucleotide diphosphate reductase alpha subunit Holiday junction resolvase Exonuclease I HTH binding protein Rec. A Metalloprotease Exonuclease II Methyltransferase RNA Helicase HNH Homing Endonuclease Metal Dependent Hydrolase DNA Binding Protein Functional annotation for 62 proteins, (62/288)*100 = a little over 21% Transposase DNA Polymerase 1 Exopolyphosphatase Ligase RNA Ligase HNH Endonuclease HNH Homing Endonuclease Holin HNH Homing Endonuclease Sigma Factor Rec. A ss. DNA Binding Protein DNA Polymerase II DNA Polymerase 1 Ftsk/ Spo. IIIE Protein Par. M Sigma Factor d. UTP nucleotidylhydrolase HTH binding domain protein HNH endonuclease III DNA Helicase 2 DNA Binding Protein Terminase Baseplate protein J Baseplate assembly Endolysin protein Pho. H Minor tail protein Thymidylate synthase Dephospho – Co. A kinase Dihydrofolate reductase Tail fiber protein Tail lysin 1 Tape measure, tail lysin Tail assembly chaperone Peptidase Tail spike Tail Assembly Chaperone DNA Primase Tail Sheath Portal Prohead Protease Major Capsid Protein Functional Annotations for Bacillus Phage Phrodo
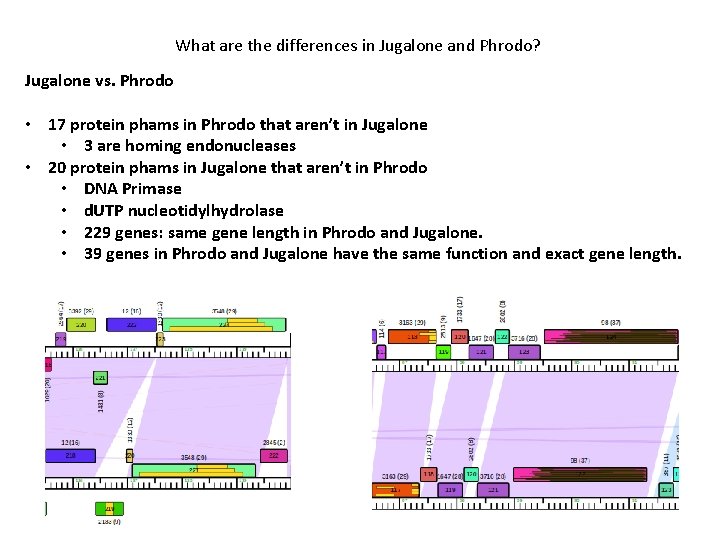
What are the differences in Jugalone and Phrodo? Jugalone vs. Phrodo • 17 protein phams in Phrodo that aren’t in Jugalone • 3 are homing endonucleases • 20 protein phams in Jugalone that aren’t in Phrodo • DNA Primase • d. UTP nucleotidylhydrolase • 229 genes: same gene length in Phrodo and Jugalone. • 39 genes in Phrodo and Jugalone have the same function and exact gene length.
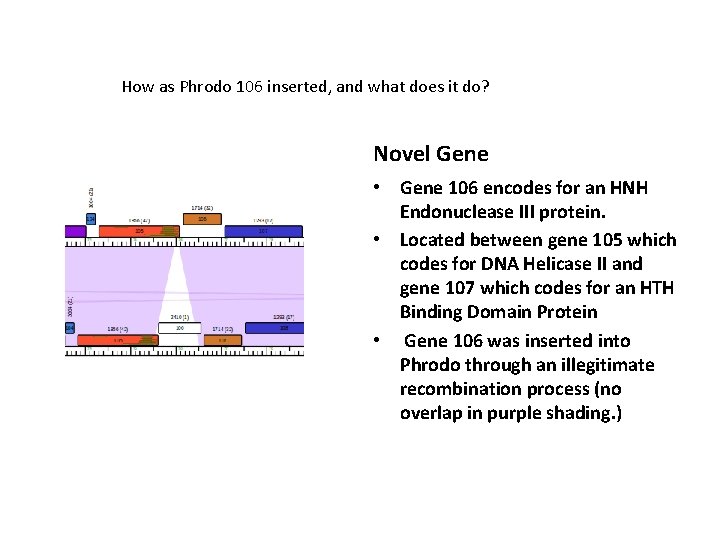
How as Phrodo 106 inserted, and what does it do? Novel Gene • Gene 106 encodes for an HNH Endonuclease III protein. • Located between gene 105 which codes for DNA Helicase II and gene 107 which codes for an HTH Binding Domain Protein • Gene 106 was inserted into Phrodo through an illegitimate recombination process (no overlap in purple shading. )
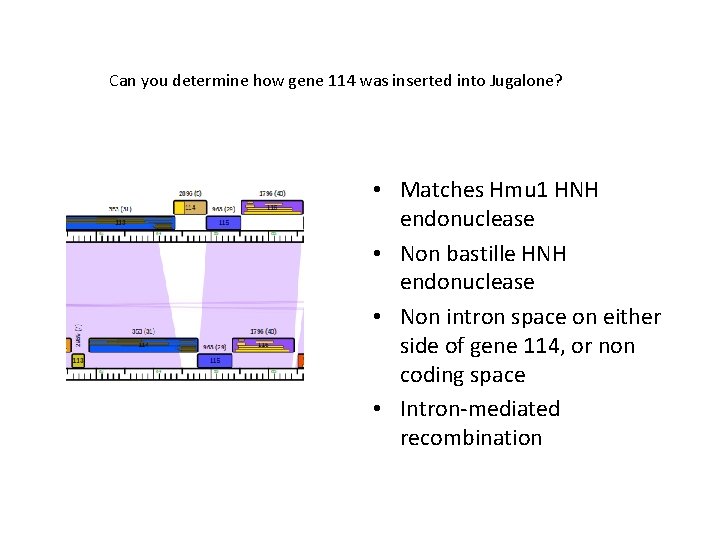
Can you determine how gene 114 was inserted into Jugalone? • Matches Hmu 1 HNH endonuclease • Non bastille HNH endonuclease • Non intron space on either side of gene 114, or non coding space • Intron-mediated recombination
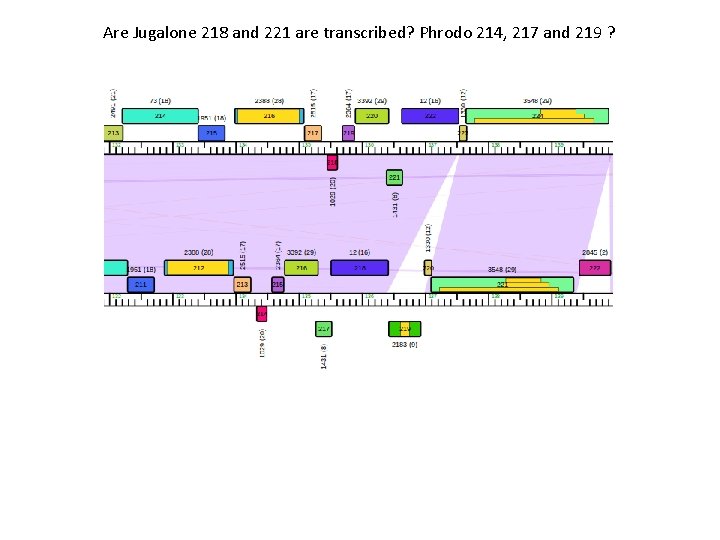
Are Jugalone 218 and 221 are transcribed? Phrodo 214, 217 and 219 ?
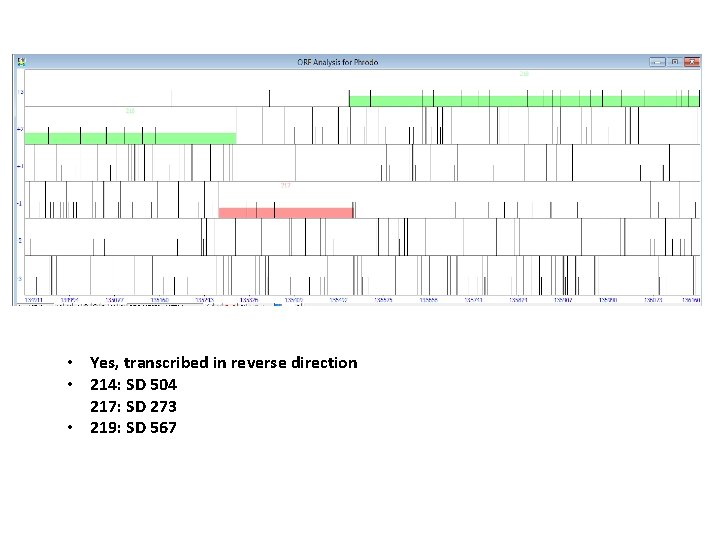
• Yes, transcribed in reverse direction • 214: SD 504 217: SD 273 • 219: SD 567
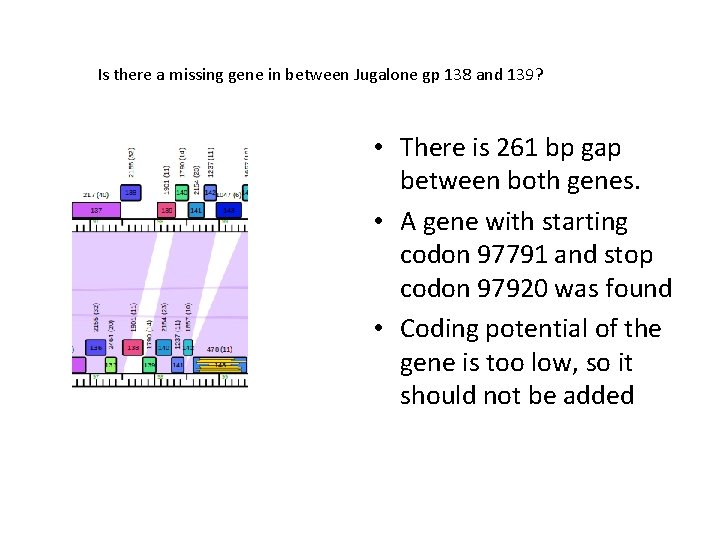
Is there a missing gene in between Jugalone gp 138 and 139? • There is 261 bp gap between both genes. • A gene with starting codon 97791 and stop codon 97920 was found • Coding potential of the gene is too low, so it should not be added
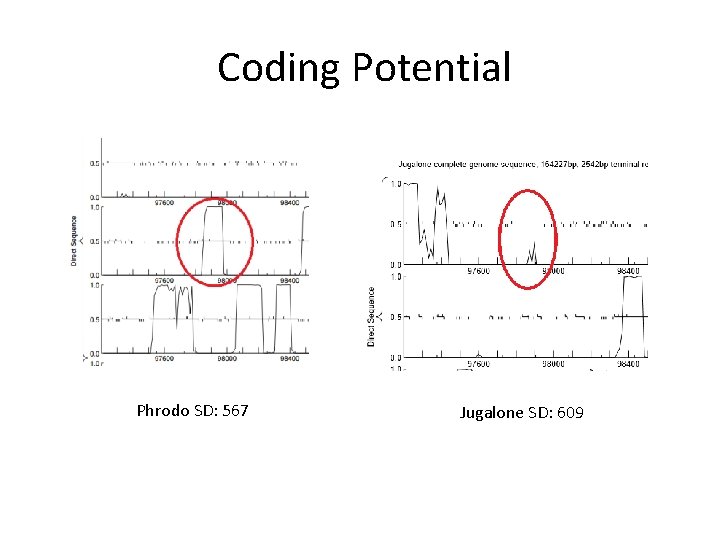
Coding Potential Phrodo SD: 567 Jugalone SD: 609
- Slides: 12