Axis of Rotation Crystal Structure 2 Axis of







- Slides: 34


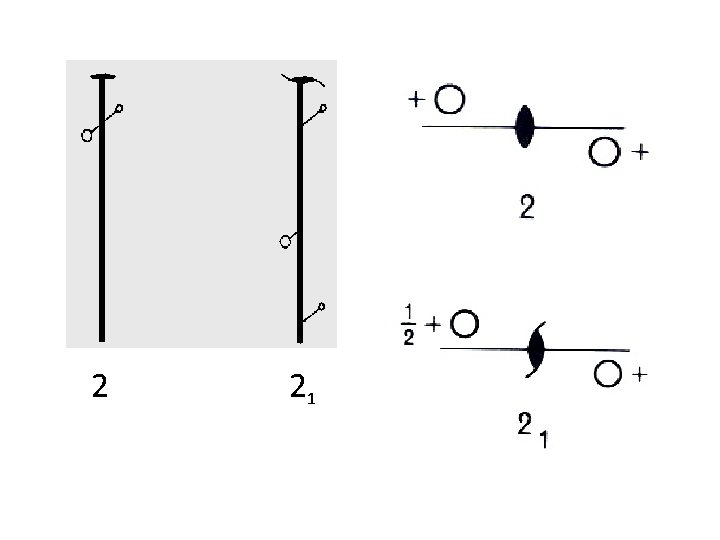
Axis of Rotation Crystal Structure 2

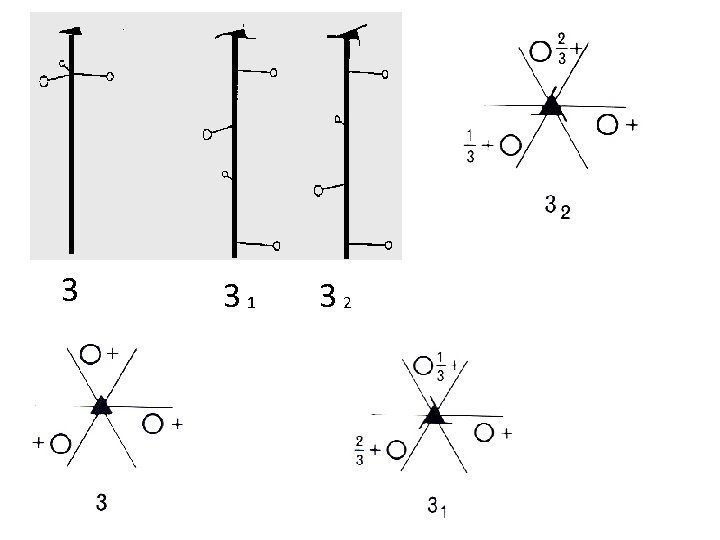
Axis of Rotation Crystal Structure 3



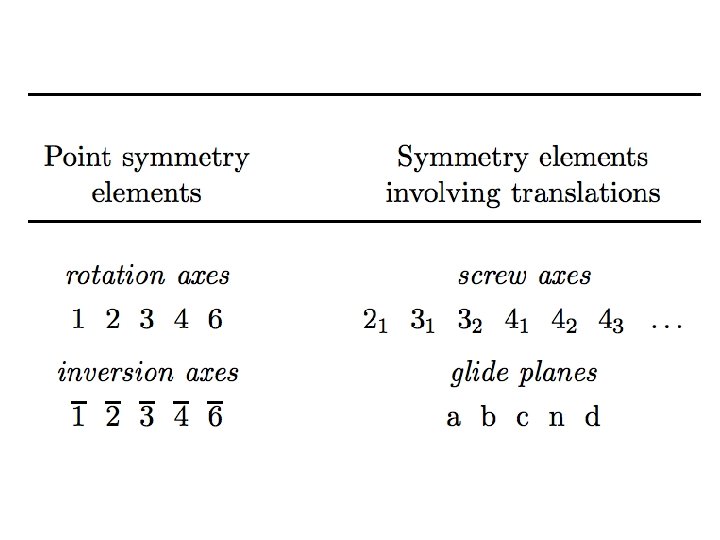
2 21

3 31 32

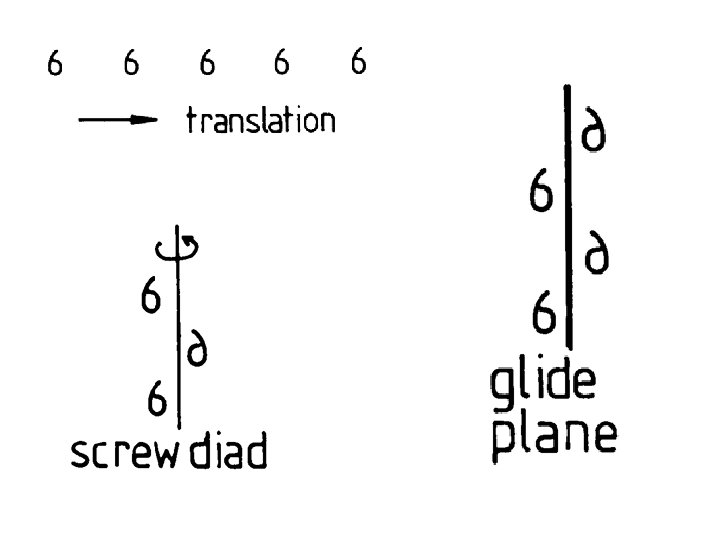
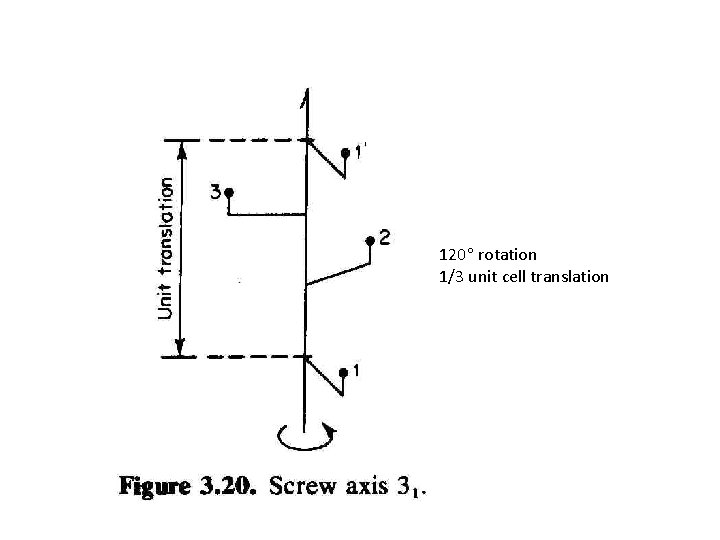
120 rotation 1/3 unit cell translation


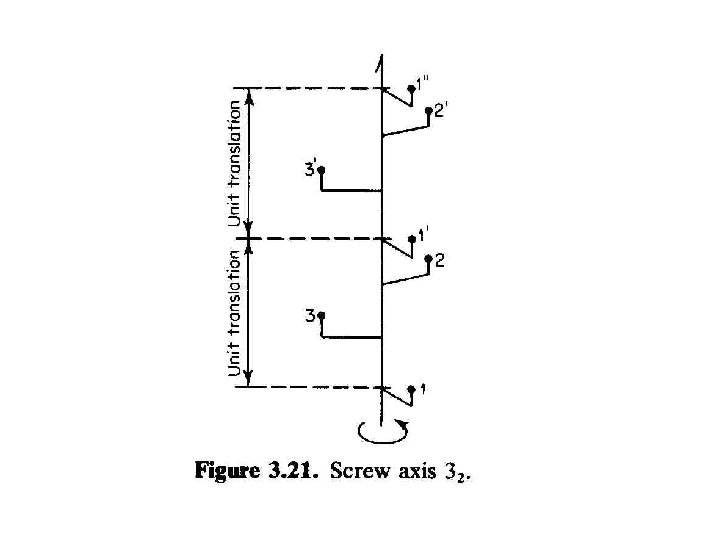
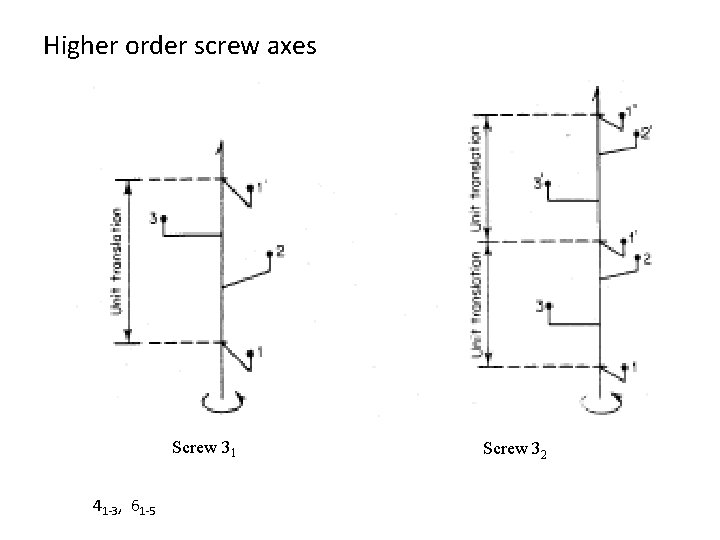
Higher order screw axes Screw 31 41 -3, 61 -5 Screw 32

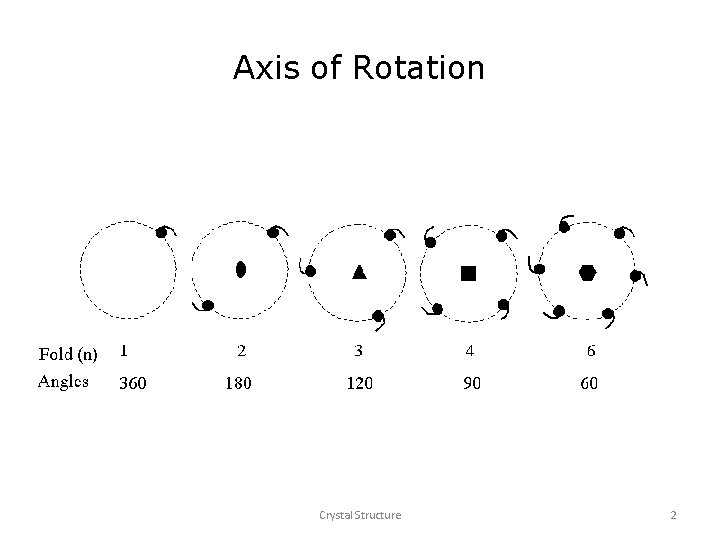
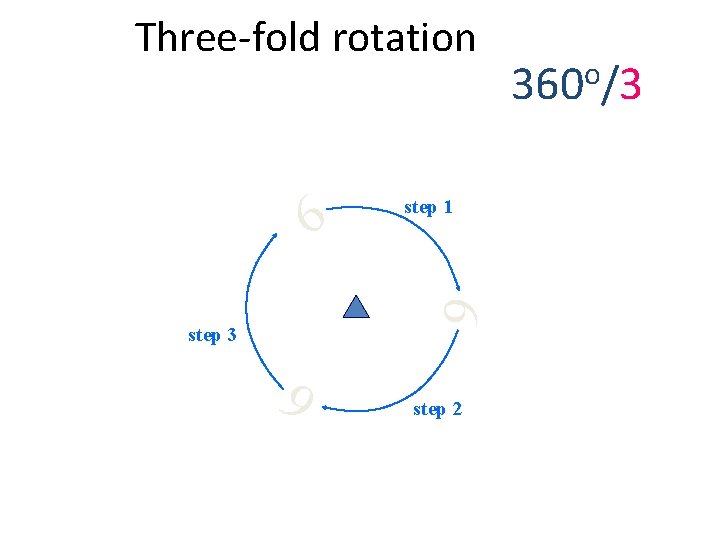
Three-fold rotation 6 step 1 6 step 3 step 2 360 o/3 6

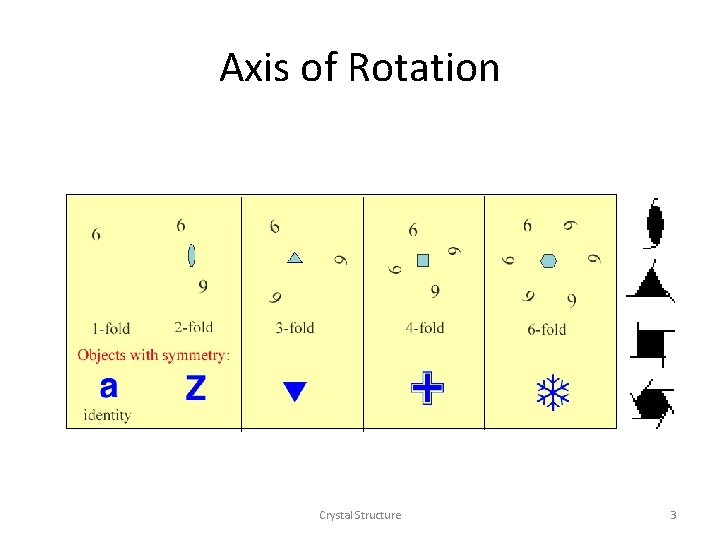
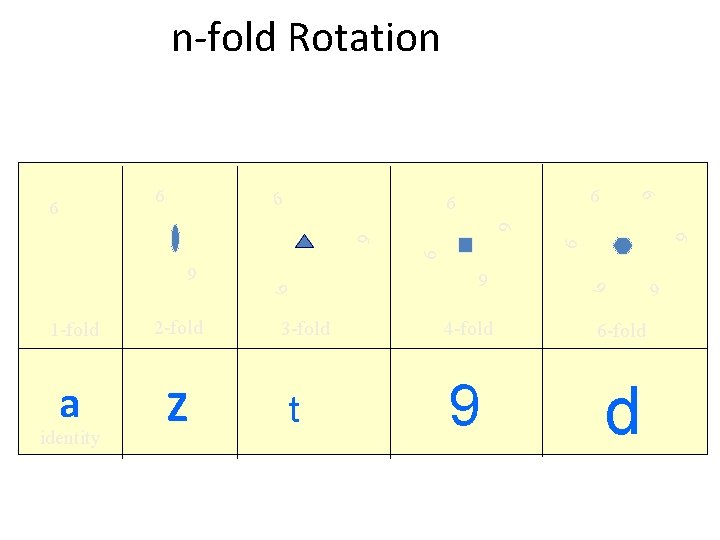
n-fold Rotation 6 6 6 Z identity t 6 a 3 -fold 4 -fold 6 -fold 9 d 6 6 2 -fold 6 1 -fold

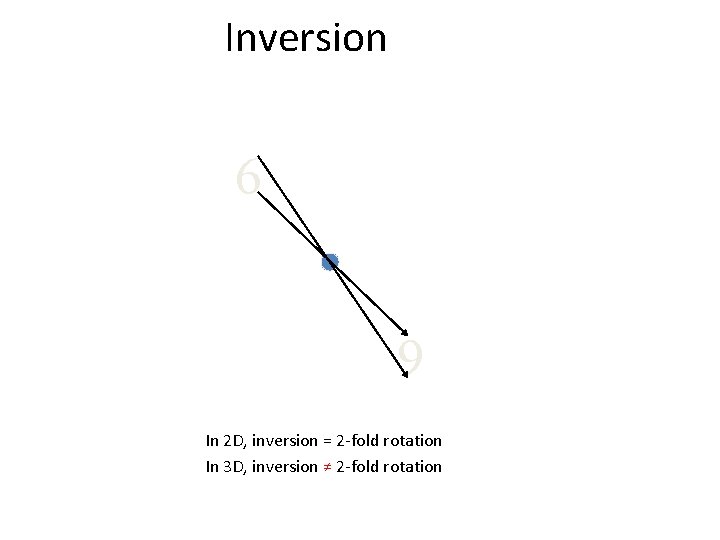
Inversion 6 6 In 2 D, inversion = 2 -fold rotation In 3 D, inversion ≠ 2 -fold rotation

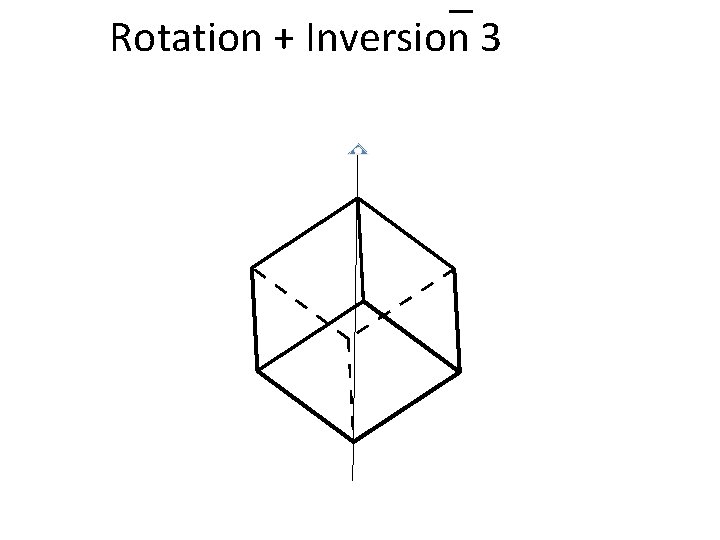
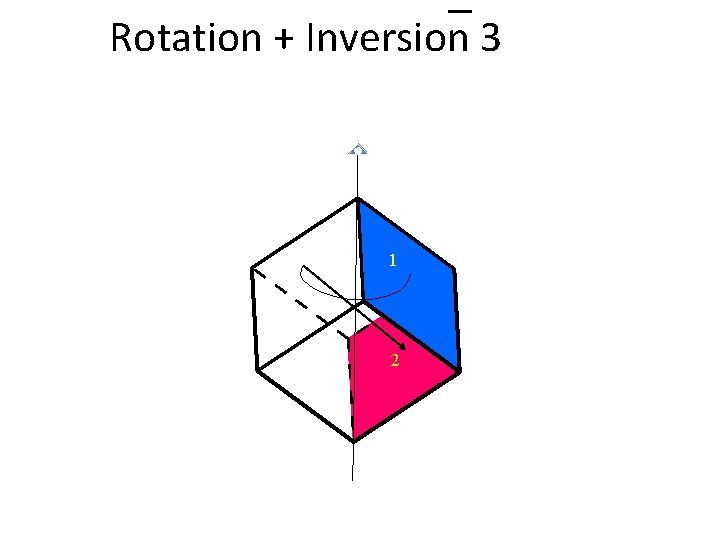
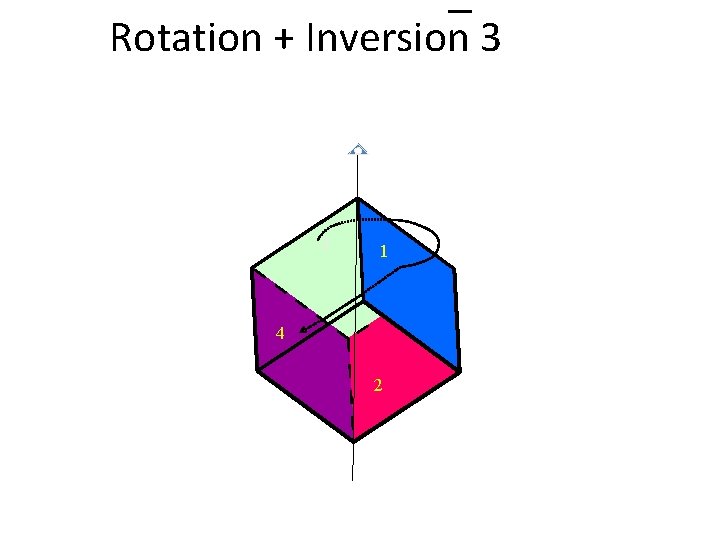
Rotation + Inversion 3

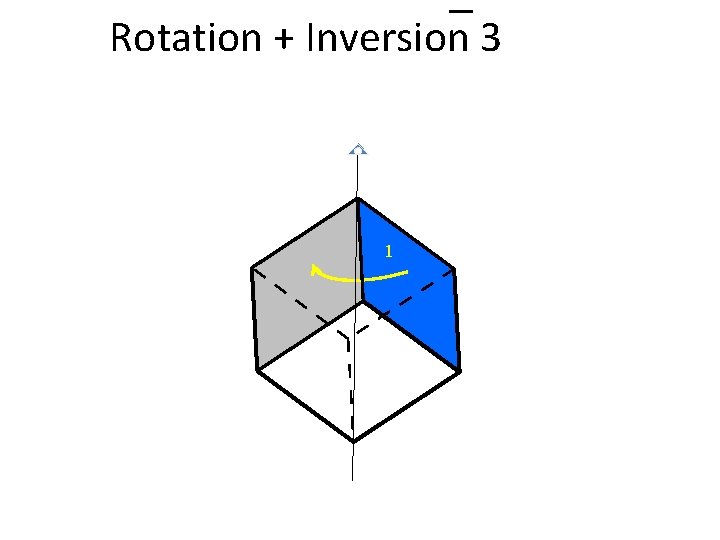
Rotation + Inversion 3 1

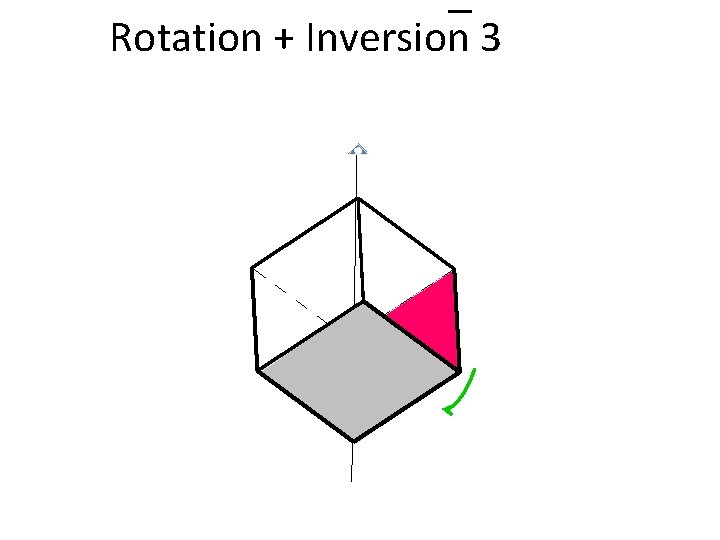
Rotation + Inversion 3

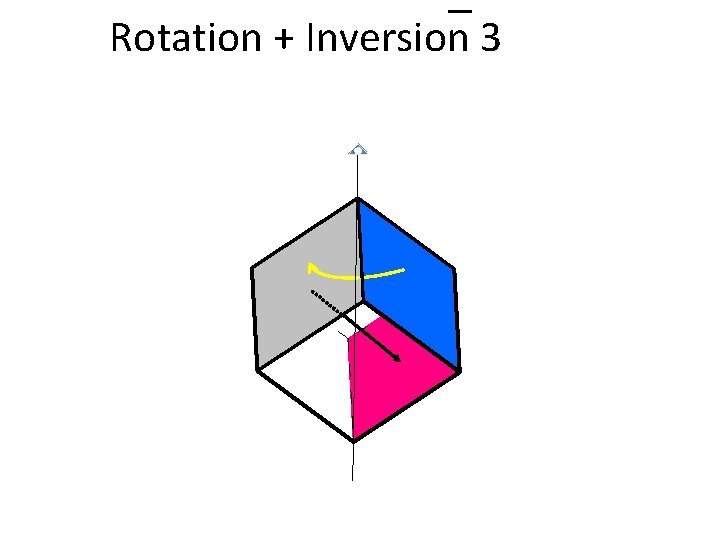
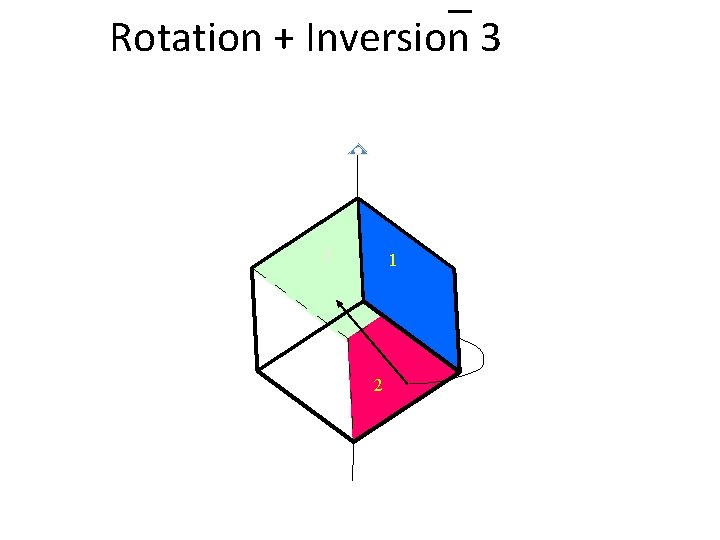
Rotation + Inversion 3 1 2

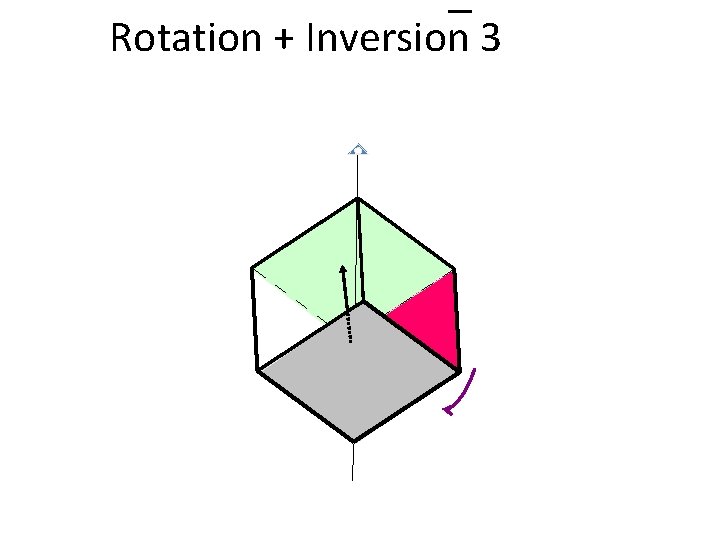
Rotation + Inversion 3

Rotation + Inversion 3

Rotation + Inversion 3 3 1 2

Rotation + Inversion 3 3 1 4 2

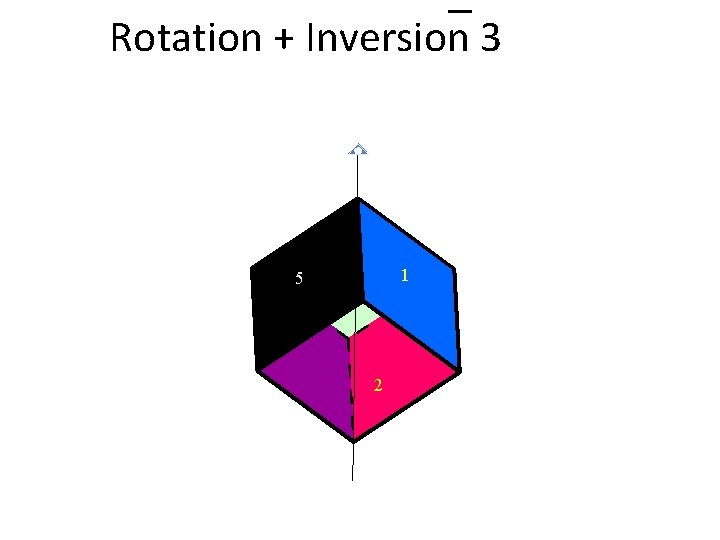
Rotation + Inversion 3 1 5 2

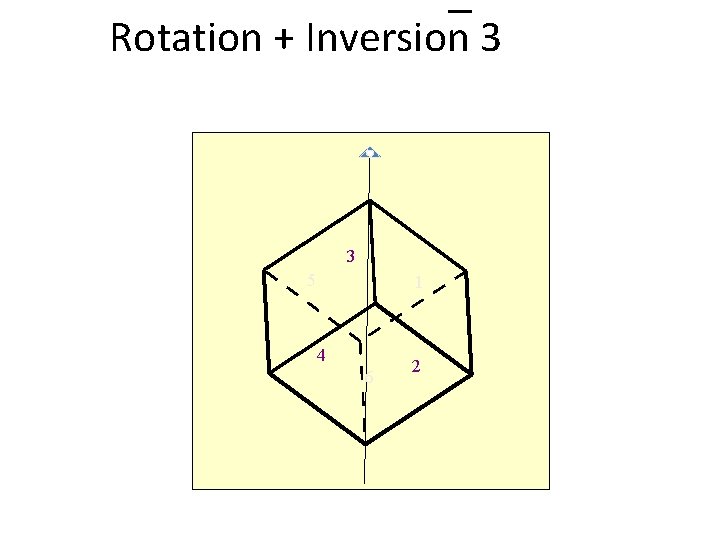
Rotation + Inversion 3 3 5 1 4 6 2

Crystal systems: length/angle relations Klein Fig. 5. 27, pg. 196

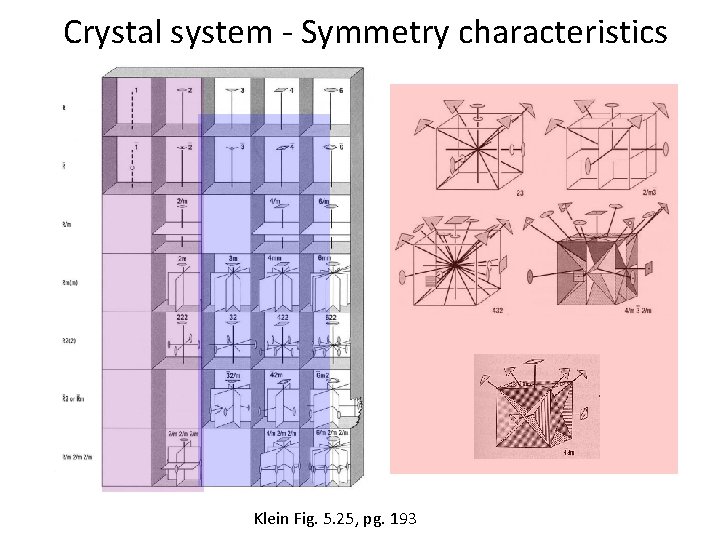
Crystal System - Symmetry Characteristics

Crystal system - Symmetry characteristics Klein Fig. 5. 25, pg. 193

Unit cell and asymmetric units Unit cell Asymmetric units Unique atoms Symmetry-related atoms • We must first find out the symmetry • •

Apply correct symmetry Too low symmetry wrong symmetry correct symmetry

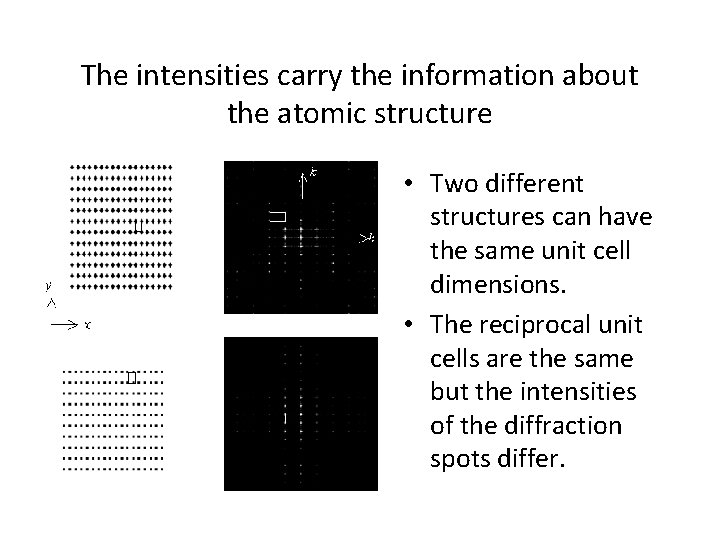
The intensities carry the information about the atomic structure • Two different structures can have the same unit cell dimensions. • The reciprocal unit cells are the same but the intensities of the diffraction spots differ.

Symmetry is best seen in reciprocal space • A square unit cell is necessary but not sufficient for the crystal having 4 -fold symmetry. • If the atoms in the unit cell are not arranged with 4 -fold symmetry (a), the diffraction pattern will not have 4 -fold symmetry (b). • A crystal with 4 -fold symmetry (c) gives rise to a diffraction pattern with 4 fold symmetry (d).

An example of symmetry correction PDB code: 1 yup spacegroup (PDB): P 1 8 molecules per a. u. spacegroup (true): P 21 4 molecules per a. u. Pseudo-symmetry spacegroup: (because of pseudo-translation) C 2 2 molecules per a. u.

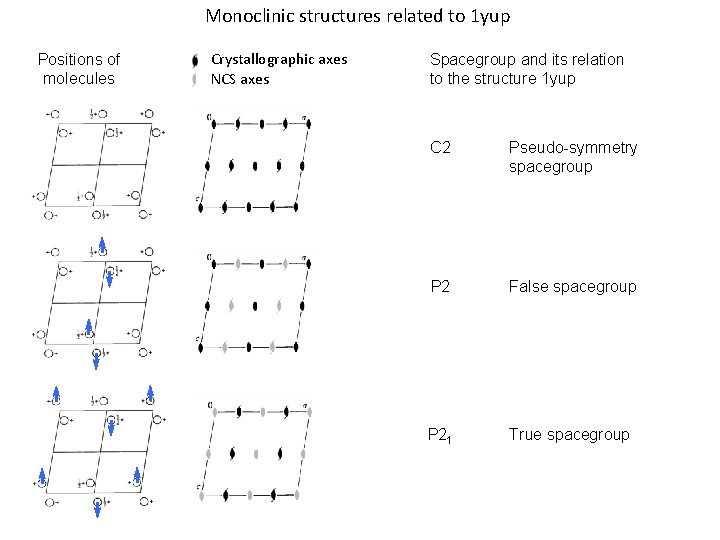
Monoclinic structures related to 1 yup Positions of molecules Crystallographic axes NCS axes Spacegroup and its relation to the structure 1 yup C 2 Pseudo-symmetry spacegroup P 2 False spacegroup P 21 True spacegroup

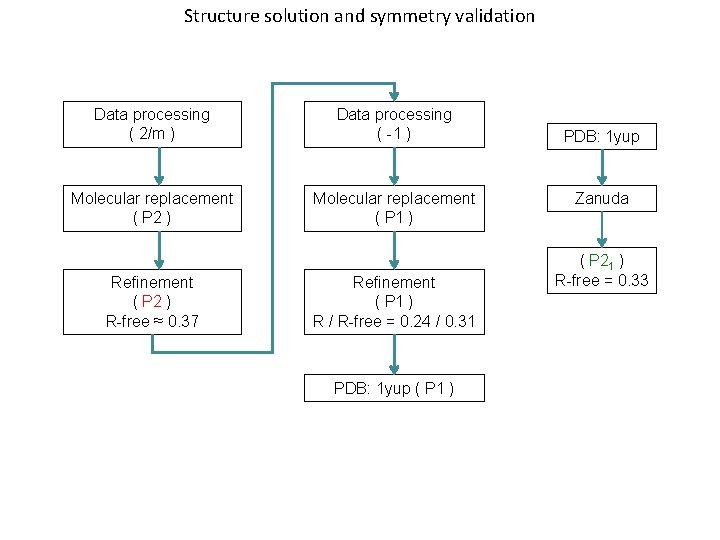
Structure solution and symmetry validation Data processing ( 2/m ) Data processing ( -1 ) Molecular replacement ( P 2 ) Molecular replacement ( P 1 ) Refinement ( P 2 ) R-free ≈ 0. 37 Refinement ( P 1 ) R / R-free = 0. 24 / 0. 31 PDB: 1 yup ( P 1 ) PDB: 1 yup Zanuda ( P 21 ) R-free = 0. 33