Autophagy n n n Autophagosome A double membrane


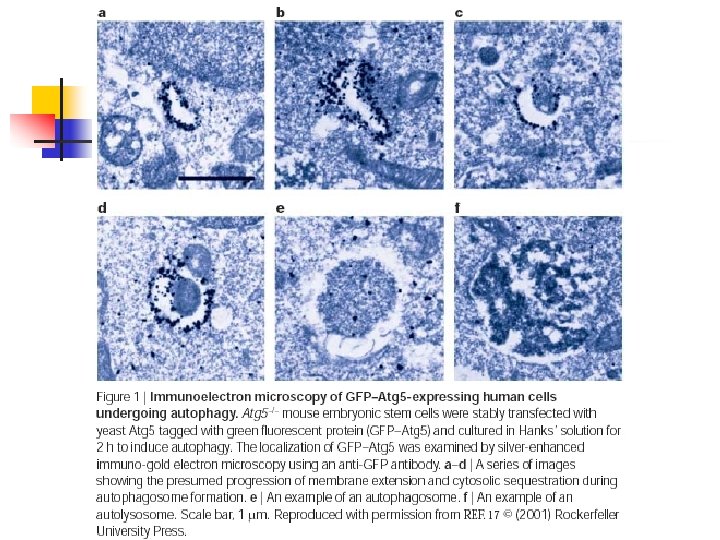
Autophagy n n n Autophagosome: A double membrane bound compartment that engulfs cytosol and degrades the cytoplasmic contents. Large: 400 -1500 nm May originate from ER or from fusion of lipid -containing vesicles that form ‘sequestration crescent’.

Autophagic pathways n Plants and animals n Bacteria, viruses, parasites n Highly regulated processes n Stages: induction, execution, maturation

Autophagy genes n Based on S. cerevisiae yeast studies: n ATG in yeast, homologs in other spp. n Involved in: n n n Tumor suppression Starvation responses Preventing premature cell senescence

Role of autophagy n n n Does autophagy benefit the host as a defense mechanism? Does autophagy benefit the microbe by facilitating survival and replication? Or both?

Role of autophagy n n A host response to degrade intracellular pathogens? Hepatic pathogen purging n n 10 weeks: 75% of chimpanzee liver cells contained hepatitis B viral proteins. 20 weeks: chimpanzees were virus free, with no evidence of extensive cell death!

Role of autophagy n n A microbe strategy for enhanced persistence and intracellular replication? Murine hepatitis virus (MHV): n Cellular markers on viral membranes were consistent with autophagosomal markers. n ATG 5 mutant had 1000 -fold decrease in viral yield. n Plasmid expressing ATG 5 restored viral yield.

Induction n TOR (Target Of Rapamycin) kinase n Inhibits autophagy when phosphorylated. n Also regulates protein and amino acid synthesis. n n Rapamycin and nutrient starvation dephosphorylate TOR, inducing autophagy. Tamoxifen induces autophagy too (TOR? )

Induction n Trimeric G proteins n n High amino acids inactivate G proteins, so aa depletion may induce autophagy via G proteins. PI 3 Ks (Phosphatidylinositol-3 -kinases) n n Essential for starvation induced autophagy. 3 -MA, wortmannin, LY 294002 target PI 3 Ks, and result in inhibition of autophagy.
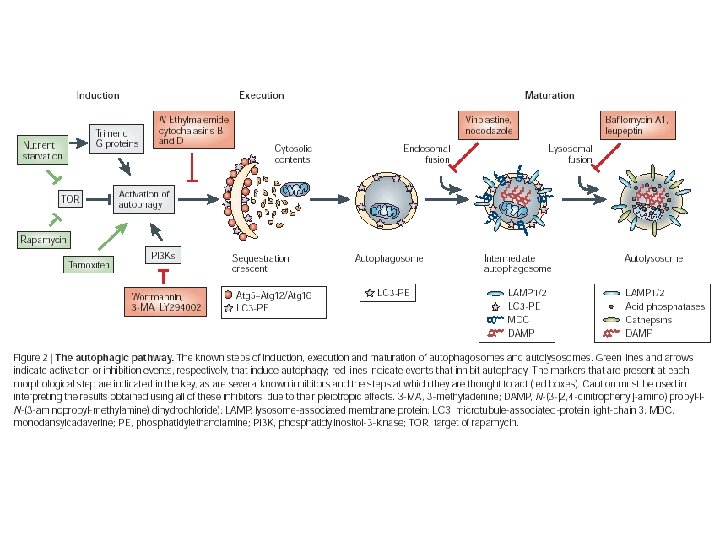

Execution n Covalent linkage of Atg 5 and Atg 12 n Covalent lipidation of Atg 8 n Enzymes Atg 3, Atg 7, and Atg 10 are homologs of ubiquitylation enzymes but are used to modify pathway components instead of labeling them for degradation.
Maturation n GTPases (Rab 24) mediate vesicle fusion. n Intermediate autophagosomes n n n Fuse with endosomal vesicles. Acquire LAMP, accumulate DAMP proteins. Mature autolysosomes n n Fuse with lysosomes. Acquire cathepsins and acid phosphatases.
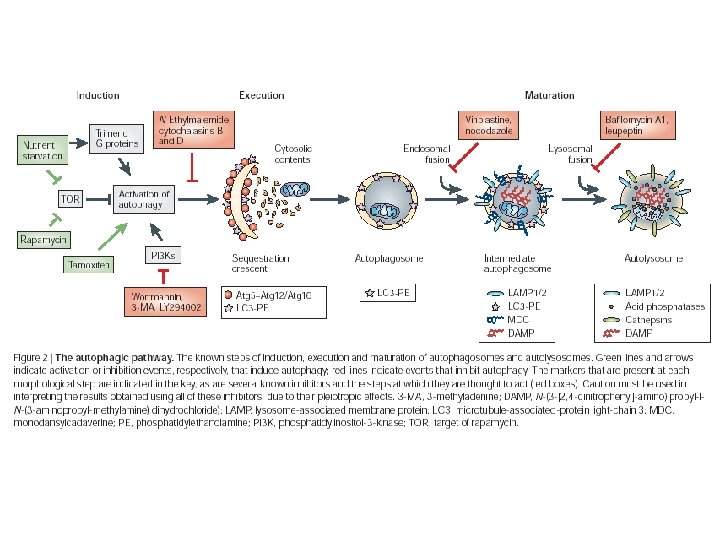

Detection methods n Microscopy n n Biochemistry n n n Electron microscopy – ultrastructure, morphology, volumetrics, staining. Enzyme activity assays. Radioactive degradation studies. Marker studies n n Autophagosomal or organelle markers. Fluorescence or immunodetection.

Bacterial susceptibility n n n Astragalus-Mesorhizobium: Bacteria differentiate within membrane compartments until they can fix nitrogen and establish symbiosis. Nutrient starvation: bacterial degradation observed. Autophagy? http: //www. bioscience. drexel. edu/Homepage/immunology/presentations/group 6/Aintro. htm

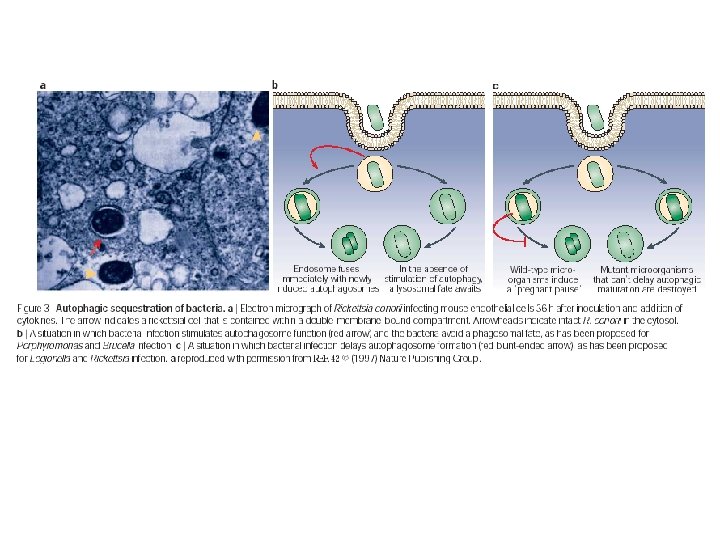
Bacterial susceptibility n Rickettsiae conorii n n n Sensitive to NO produced by IFN, TNF-α. Correlations between autophagosome-like structures and bacterial degradation. Are autophagosomes destroying bacteria or just cleaning up after bacteria are killed?

Bacterial susceptibility n Listeria monocytogenes n n Enter host cells by phagocytosis, escape from phagosomes, multiply in cytoplasm. Act. A (actin) mutants are engulfed in autophagosome-like compartments. Wortmannin reduces bacterial entry into autophagosomes. Nutrient depletion increases bacterial entry into autophagosomes. http: //www. rapidmicrobiology. com/news/603 h 48. php
Bacterial subversion n Porphyromonas gingivalis n Infects human coronary artery endothelial cells. n Localizes to autophagosome-like compartments. n Wortmannin: compartments resembled lysosomes and acquired cathepsin earlier. Bacterial survival decreased. http: //www. pgingivalis. org/ATCC 33277(1). htm

Bacterial subversion n Brucella abortus n n n Endosomal uptake, then autophagosomes. Wortmannin reduced survival, cell starvation increased survival. Vir. B mutants have Type IV secretion mutation that inhibits intracellular transport and growth. Mutants are localized to membrane compartments that acquire cathepsin earlier, resembling lysosomes.

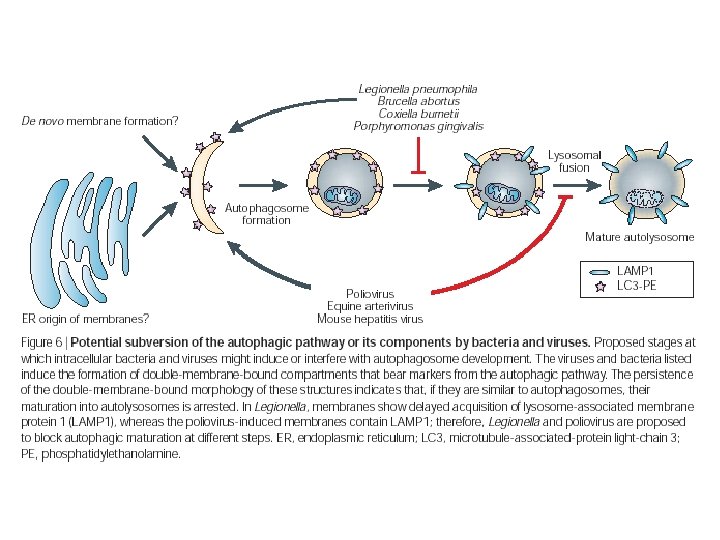
Bacterial subversion n Legionella pneumophila n n n Replicates in autophagosome-like compartments in macrophages. Dot/icm mutants are defective in Type IV secretion involved with organelle trafficking or intracellular multiplication. Mutants localized to lysosomal-like vesicles, not autophagosomes. ‘Pregnant pause’ model says that autophagosome maturation is delayed to allow for pathogen development.

Bacterial subversion n Coxiella burnetti n n n Replicates within autophagosome-like acidic vesicles. Rab 7 mutants had altered size and numbers of vesicles containing bacteria. Dot/icm homologs in Coxiella were able to return Dot/icm deficient Legionella to a wild phenotype.

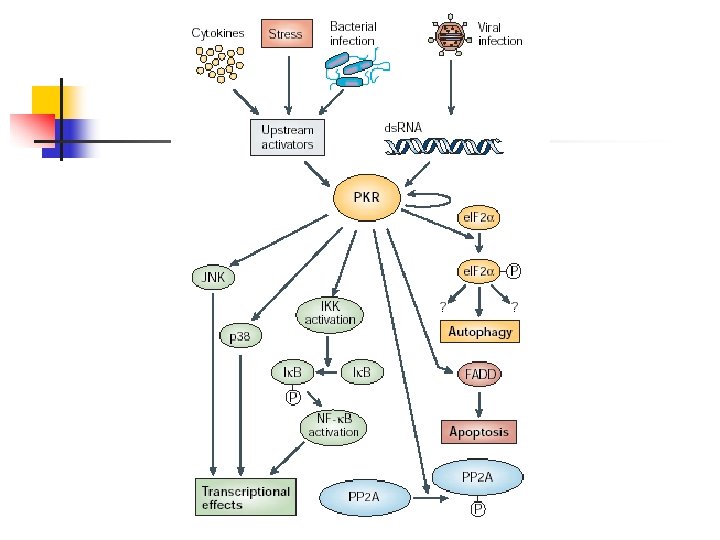
Viral susceptibility n Herpes-virus n n n PKR kinase – phosphorylates eukaryotic translation -initiation factor e. IF 2α to inhibit and deregulate cellular translation. PKR can also induce autophagy, apoptosis, and activate NF-KB. ICP 34. 5 – produced by herpes simplex virus 1 to antagonize PKR function by dephosphorylating e. IF 2α.

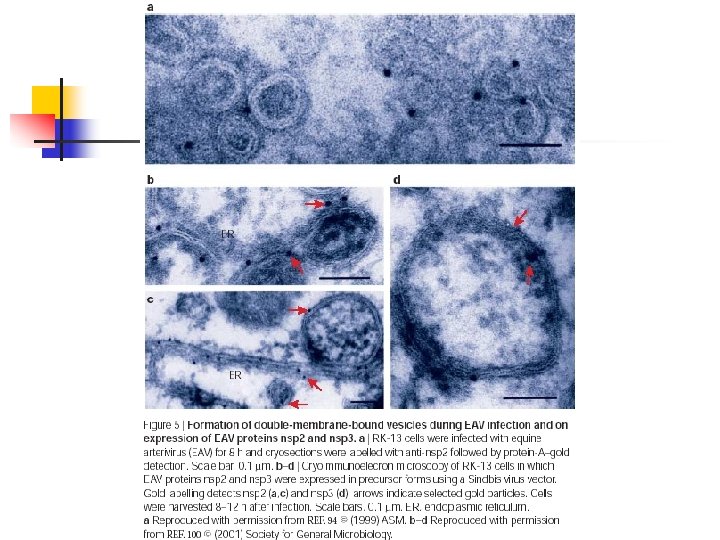
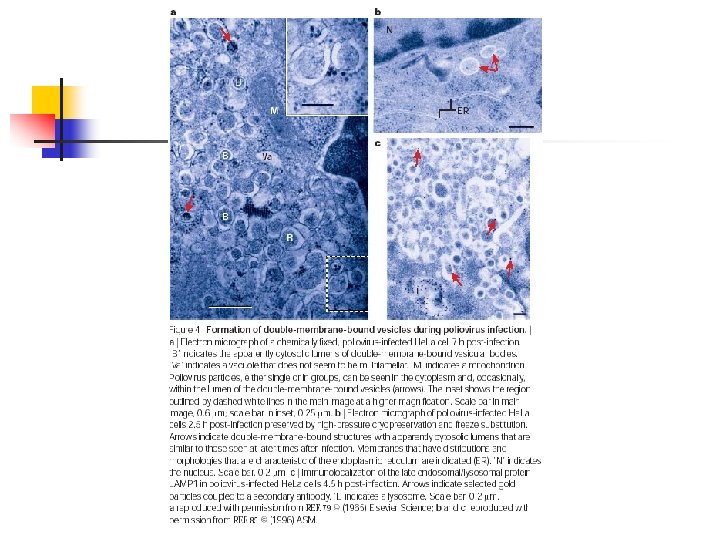
Viral subversion n Positive-strand RNA viruses n n n Poliovirus, murine hepatitis virus (MHV), equine arterivirus (EAV), SARS human corona virus. Require membranes for replication. Autophagosomes are induced during infection, but are they part of the viral replication process or the host response to eliminate pathogens?

TIP OF THE ICEBERG?
- Slides: 30