Authors David Ginsburg 2009 License Unless otherwise noted

Author(s): David Ginsburg, 2009 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution–Non-commercial–Share Alike 3. 0 License: http: //creativecommons. org/licenses/by-nc-sa/3. 0/ We have reviewed this material in accordance with U. S. Copyright Law and have tried to maximize your ability to use, share, and adapt it. The citation key on the following slide provides information about how you may share and adapt this material. Copyright holders of content included in this material should contact open. michigan@umich. edu with any questions, corrections, or clarification regarding the use of content. For more information about how to cite these materials visit http: //open. umich. edu/education/about/terms-of-use. Any medical information in this material is intended to inform and educate and is not a tool for self-diagnosis or a replacement for medical evaluation, advice, diagnosis or treatment by a healthcare professional. Please speak to your physician if you have questions about your medical condition. Viewer discretion is advised: Some medical content is graphic and may not be suitable for all viewers.

Citation Key for more information see: http: //open. umich. edu/wiki/Citation. Policy Use + Share + Adapt { Content the copyright holder, author, or law permits you to use, share and adapt. } Public Domain – Government: Works that are produced by the U. S. Government. (USC 17 § 105) Public Domain – Expired: Works that are no longer protected due to an expired copyright term. Public Domain – Self Dedicated: Works that a copyright holder has dedicated to the public domain. Creative Commons – Zero Waiver Creative Commons – Attribution License Creative Commons – Attribution Share Alike License Creative Commons – Attribution Noncommercial Share Alike License GNU – Free Documentation License Make Your Own Assessment { Content Open. Michigan believes can be used, shared, and adapted because it is ineligible for copyright. } Public Domain – Ineligible: Works that are ineligible for copyright protection in the U. S. (USC 17 § 102(b)) *laws in your jurisdiction may differ { Content Open. Michigan has used under a Fair Use determination. } Fair Use: Use of works that is determined to be Fair consistent with the U. S. Copyright Act. (USC 17 § 107) *laws in your jurisdiction may differ Our determination DOES NOT mean that all uses of this 3 rd-party content are Fair Uses and we DO NOT guarantee that your use of the content is Fair. To use this content you should do your own independent analysis to determine whether or not your use will be Fair.

The Human Genome M 1 Patients and Populations David Ginsburg, MD August 22, 2008 Fall 2008

Learning Objectives UNDERSTAND: • The basic anatomy of the human genome [eg. 3 X 109 bp (haploid genome); 1 -2% coding sequence (~20, 000 genes); types and extent of DNA sequence variation]. • Recombination and how it allows genes to be mapped • Genetic data for a pedigree, assigning phase, defining haplotypes • Linkage: Use of markers linked to a disease gene for genetic diagnosis and to identify disease genes (positional cloning) • Distinction between a linked marker and the disease causing mutation itself • Linkage disequilibrium and haplotype blocks • Genome wide association studies (GWAS) to identify gene variants contributing to complex diseases/traits

DNA Sequence Variation • DNA Sequence Variation: – Human to human: ~0. 1% (1: 1000 bp) • Human genome = 3 X 109 bp X 0. 1% =~3 X 106 DNA common variants – Human to chimp: ~1 -2% – More common in “junk” DNA: introns, intergenic regions • poly·mor·phism Pronunciation: "päl-i-'mor-"fiz-&m Function: noun : the quality or state of existing in or assuming different forms: as a (1) : existence of a species in several forms independent of the variations of sex (2) : existence of a gene in several allelic forms (3) : existence of a molecule (as an enzyme) in several forms in a single species
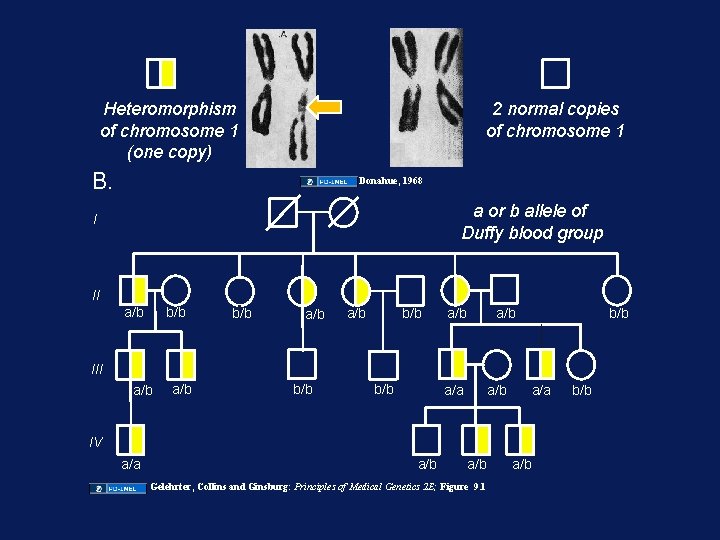
Heteromorphism of chromosome 1 (one copy) 2 normal copies of chromosome 1 B. Donahue, 1968 a or b allele of Duffy blood group I II a/b b/b a/b a/b b/b III a/b b/b a/a a/b a/a IV a/a a/b Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 1 a/b b/b
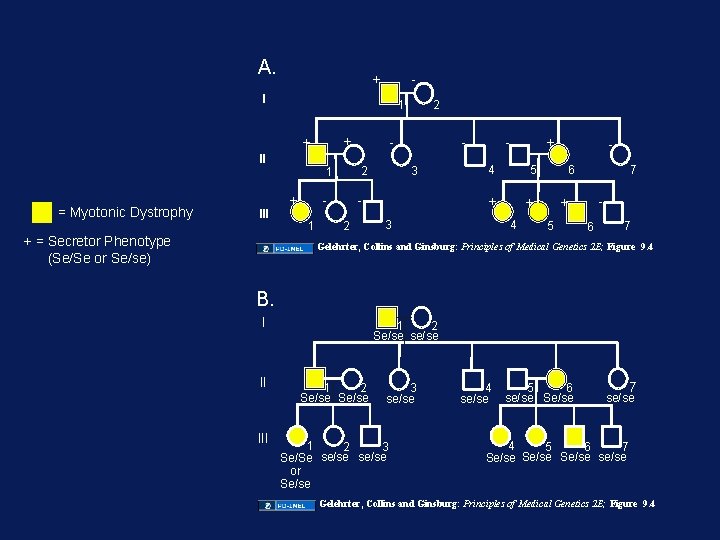
A. + - I + + II = Myotonic Dystrophy III + = Secretor Phenotype (Se/Se or Se/se) 2 1 + 1 - 1 2 - 3 4 - 5 + 3 + + 4 7 6 - + 5 6 7 Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 4 B. I II III 1 2 Se/se se/se 1 2 Se/se 3 se/se 1 2 3 Se/Se se/se or Se/se 4 se/se 5 6 se/se Se/se 7 se/se 4 6 5 7 Se/se se/se Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 4
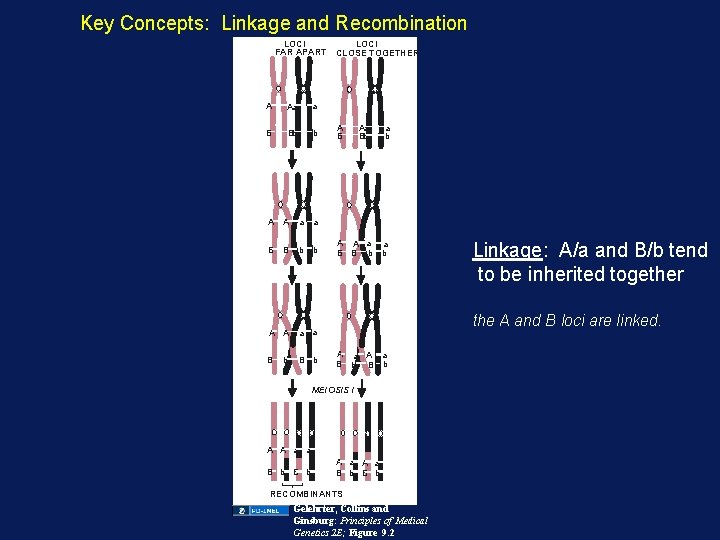
Key Concepts: Linkage and Recombination LOCI FAR APART A Aa a B Bb b A A a a B B b b LOCI CLOSE TOGETHER A B Aa Bb A A a B B b a b the A and B loci are linked. A a B b MEIOSIS I A A a a B b Linkage: A/a and B/b tend to be inherited together A a B b RECOMBINANTS Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 2
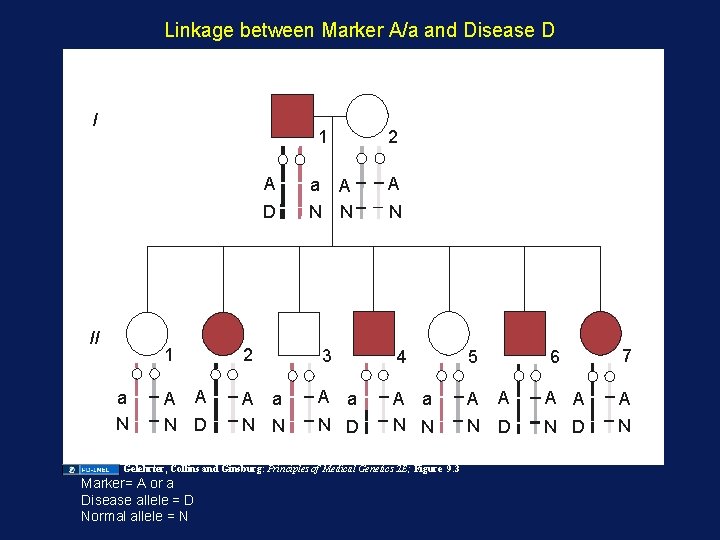
Linkage between Marker A/a and Disease D I 1 A D II a N a A N N 2 A N 1 2 3 4 5 A A N D A a N N A a N D A a N N A Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 3 Marker= A or a Disease allele = D Normal allele = N 6 7 A A N D N

Linkage between NF and RFLP marker I 1 II 1 2 3 2 recombinant 4 5 6 7 8 III 9 1 2 3 4 5 6 7 8 4. 7 kb 3. 0 kb Eco. RI 3. 0 kb PROBE Eco. RI 1. 7 kb Eco. RI * Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 5 = NEUROFIBROMATOSIS (NF 1) 9
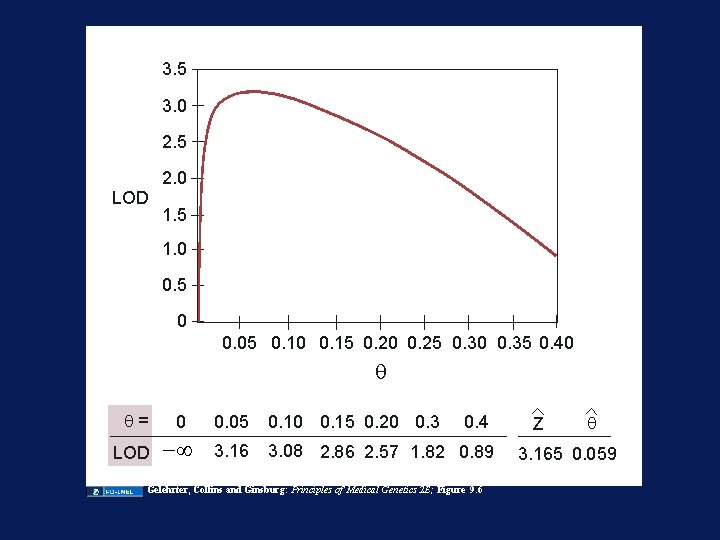
3. 5 3. 0 2. 5 2. 0 LOD 1. 5 1. 0 0. 5 0 0. 05 0. 10 0. 15 0. 20 0. 25 0. 30 0. 35 0. 40 q q= LOD 0 0. 05 0. 10 0. 15 0. 20 0. 3 -¥ 3. 16 3. 08 2. 86 2. 57 1. 82 0. 89 0. 4 Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 6 Z q 3. 165 0. 059
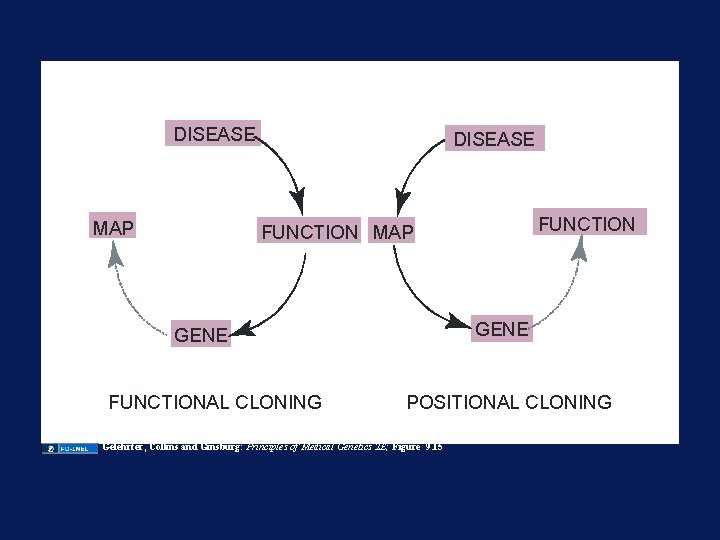
DISEASE MAP DISEASE FUNCTION MAP GENE FUNCTIONAL CLONING POSITIONAL CLONING Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 15
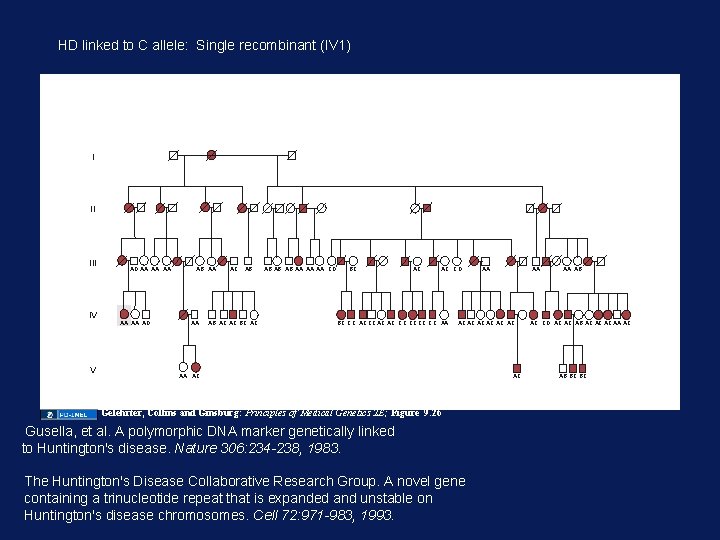
HD linked to C allele: Single recombinant (IV 1) I II IV V AD AA AA AA AD AB AA AA AC AB AB AC AC BC AC AB AB AB AA AA AA CD BC AC AC CD BC CC AC AC CC CC AA AA AA AC AC AC AA AC Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 26 Gusella, et al. A polymorphic DNA marker genetically linked to Huntington's disease. Nature 306: 234 -238, 1983. The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72: 971 -983, 1993. AC AA AB AC CD AC AC AB AC AC AC AA AC AB BC BC
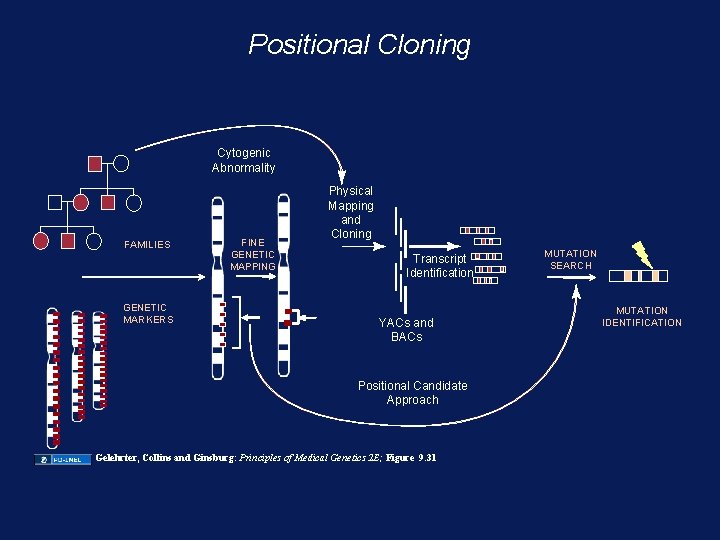
Positional Cloning Cytogenic Abnormality FAMILIES GENETIC MARKERS FINE GENETIC MAPPING Physical Mapping and Cloning Transcript Identification YACs and BACs Positional Candidate Approach Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 31 MUTATION SEARCH MUTATION IDENTIFICATION

Image of Cystic Fibrosis carrier screening advertisement removed

Positional Cloning Approach Image of positional cloning procedure removed Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E

Types of DNA Sequence Variation • RFLP: Restriction Fragment Length Polymorphism • VNTR: Variable Number of Tandem Repeats – or minisatellite – ~10 -100 bp core unit • SSR : Simple Sequence Repeat – or STR (simple tandem repeat) – or microsatellite – ~1 -5 bp core unit • SNP: Single Nucleotide Polymorphism – Commonly used to also include rare variants • Insertions or deletions – INDEL – small (few nucleotides) insertion or deletion • Rearrangement (inversion, duplication, complex rearrangement) • CNV: Copy Number Variation
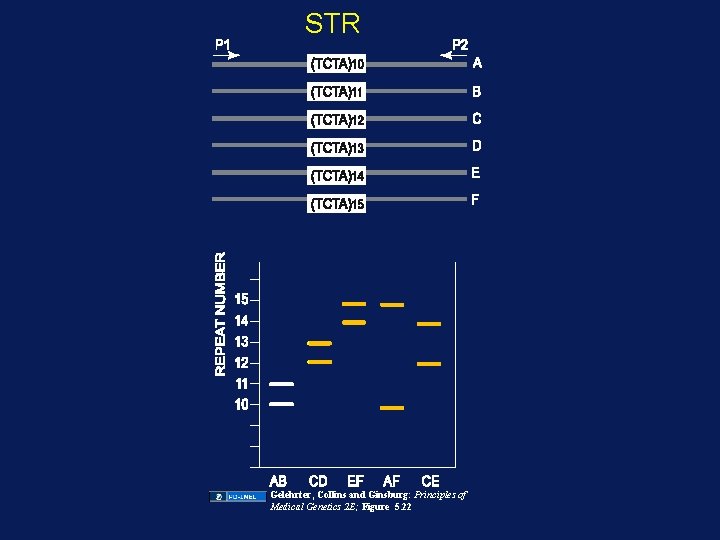
STR Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 5. 22

SNP Allele 1 A U G A A G U U U G G C A U U G C A A Allele 2 A U G A A G U U U G G U G C A U U G C A A Adapted from ASCO teaching slides • • • Most are “silent” Intragenic Promoters and other regulatory sequences Introns Exons – 5’ and 3’ untranslated regions – Coding sequence (~1 -2% of genome)
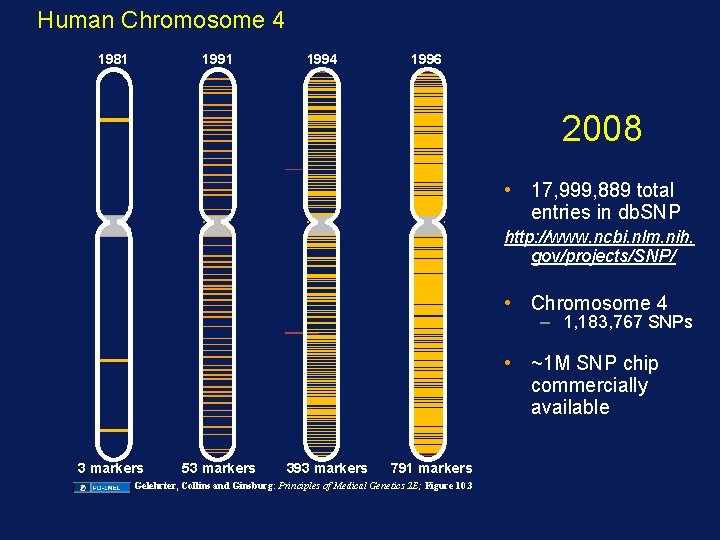
Human Chromosome 4 1981 1994 1996 2008 • 17, 999, 889 total entries in db. SNP http: //www. ncbi. nlm. nih. gov/projects/SNP/ • Chromosome 4 – 1, 183, 767 SNPs • ~1 M SNP chip commercially available 3 markers 53 markers 393 markers 791 markers Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 10. 3
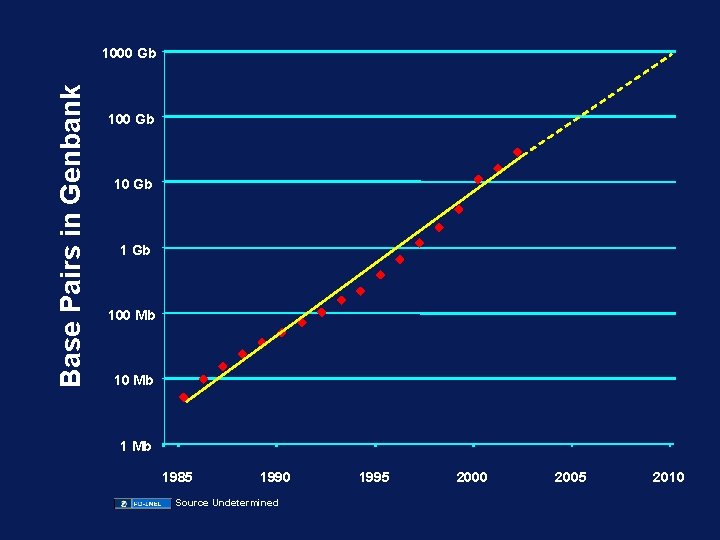
Base Pairs in Genbank 1000 Gb 100 Mb 1985 1990 Source Undetermined 1995 Year 2000 2005 2010
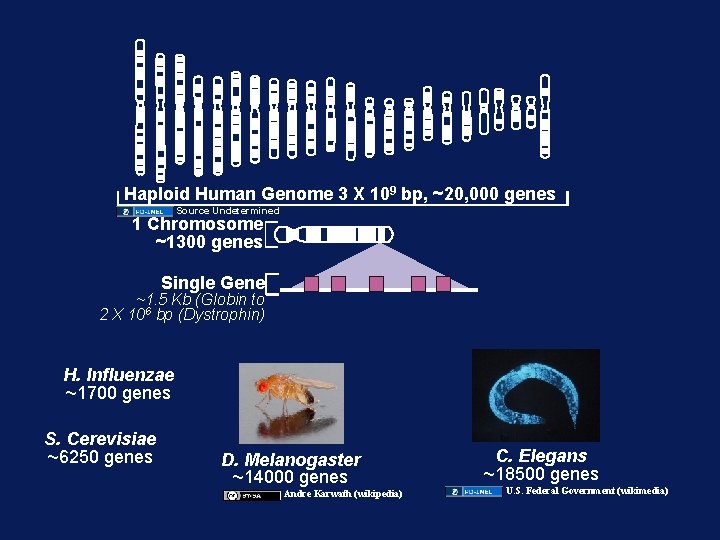
Haploid Human Genome 3 X 109 bp, ~20, 000 genes Source Undetermined 1 Chromosome ~1300 genes Single Gene ~1. 5 Kb (Globin to 2 X 106 bp (Dystrophin) H. Influenzae ~1700 genes S. Cerevisiae ~6250 genes D. Melanogaster ~14000 genes Andre Karwath (wikipedia) C. Elegans ~18500 genes U. S. Federal Government (wikimedia)

Genomes • >1000 viruses • >100 microbes • Plants (arabidopsis, oat, soybean, barley, wheat, rice, tomato, corn) • Yeast, fly, worm, human, mouse, rat, zebrafish, mosquito, malaria, ciona • Cow, pig, frog, chimp, gorilla, dog, chicken, cat, bee
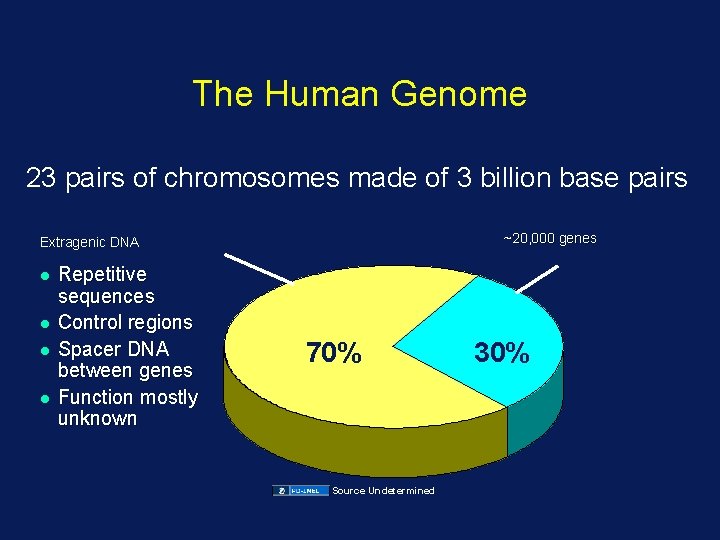
The Human Genome 23 pairs of chromosomes made of 3 billion base pairs ~20, 000 genes Extragenic DNA l l Repetitive sequences Control regions Spacer DNA between genes Function mostly unknown 70% Source Undetermined 30%

Characteristics of the Human Genome Sequence • • • 99% of euchromatin is covered, 2. 85 Gb Error rate: <<1: 100, 000 bp <350 unclonable gaps All data is freely accessible without restriction Humans have fewer genes than expected – ~20, 000 from prev. estimates of 100, 000) – ? human genes make more proteins • ~1 -2% of genome = coding sequences • ~1% = highly conserved noncoding sequences
National Center for Biotechnology

Positional Cloning Approach Image of positional cloning procedure removed Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E

University Of California Santa Cruz
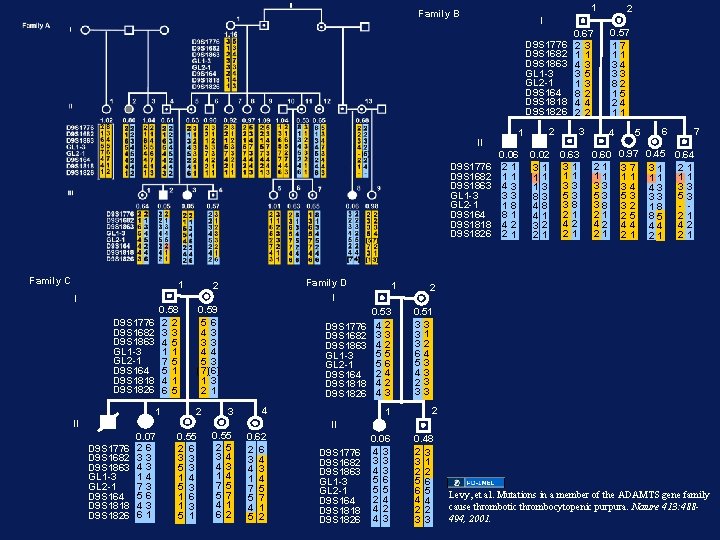
1 Family B D 9 S 1776 D 9 S 1682 D 9 S 1863 GL 1 -3 GL 2 -1 D 9 S 164 D 9 S 1818 D 9 S 1826 II D 9 S 1776 D 9 S 1682 D 9 S 1863 GL 1 -3 GL 2 -1 D 9 S 164 D 9 S 1818 D 9 S 1826 Family C 1 Family D 2 1 I I 0. 59 5 6 4 3 3 3 4 4 5 3 (7)(6) 1 3 2 1 0. 58 2 2 3 3 4 5 1 1 7 5 5 1 4 1 6 5 D 9 S 1776 D 9 S 1682 D 9 S 1863 GL 1 -3 GL 2 -1 D 9 S 164 D 9 S 1818 D 9 S 1826 1 2 D 9 S 1776 D 9 S 1682 D 9 S 1863 GL 1 -3 GL 2 -1 D 9 S 164 D 9 S 1818 D 9 S 1826 3 4 0. 55 2 5 3 4 4 3 1 4 7 5 5 7 4 1 6 2 0. 62 2 6 3 4 4 3 1 4 7 5 5 7 4 1 5 2 II D 9 S 1776 D 9 S 1682 D 9 S 1863 GL 1 -3 GL 2 -1 D 9 S 164 D 9 S 1818 D 9 S 1826 0. 07 26 33 43 14 73 56 43 61 0. 55 2 6 3 3 5 3 1 4 5 3 1 6 1 3 5 1 0. 53 42 33 42 55 56 24 42 43 1 2 I 1 0. 06 21 11 43 33 18 81 42 21 2 0. 02 31 11 13 83 48 41 32 21 0. 67 23 11 43 35 13 82 44 22 3 0. 63 31 11 33 53 38 21 42 21 0. 57 17 11 34 33 82 15 24 11 4 0. 60 21 11 33 53 38 21 42 21 5 6 7 0. 97 0. 45 0. 64 37 31 21 11 11 11 34 43 33 53 32 18 - 25 85 21 44 44 42 21 21 21 2 0. 51 33 31 32 64 53 43 23 33 2 II D 9 S 1776 D 9 S 1682 D 9 S 1863 GL 1 -3 GL 2 -1 D 9 S 164 D 9 S 1818 D 9 S 1826 0. 06 4 3 3 3 4 3 5 6 5 5 2 4 4 2 4 3 0. 48 2 3 3 1 2 2 5 6 6 5 4 4 2 2 3 3 Levy, et al. Mutations in a member of the ADAMTS gene family cause thrombotic thrombocytopenic purpura. Nature 413: 488494, 2001.
University Of California Santa Cruz
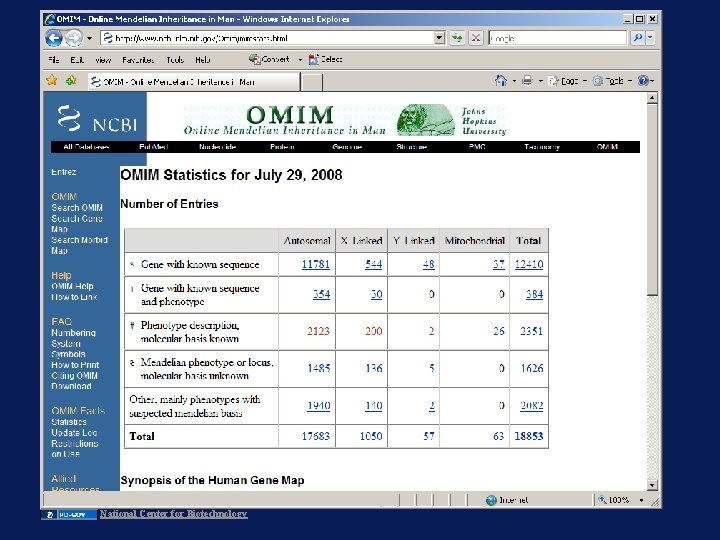
National Center for Biotechnology

Learning Objectives UNDERSTAND: • The basic anatomy of the human genome [eg. 3 X 109 bp (haploid genome); 1 -2% coding sequence (~20, 000 genes); types and extent of DNA sequence variation]. • Recombination and how it allows genes to be mapped • Linkage: Use of markers linked to a disease gene for genetic diagnosis and to identify disease genes (positional cloning) • Distinction between a linked marker and the disease causing mutation itself • Genetic data for a pedigree, assigning phase, defining haplotypes • Linkage disequilibrium and haplotype blocks • Genome wide association studies (GWAS) to identify gene variants contributing to complex diseases/traits
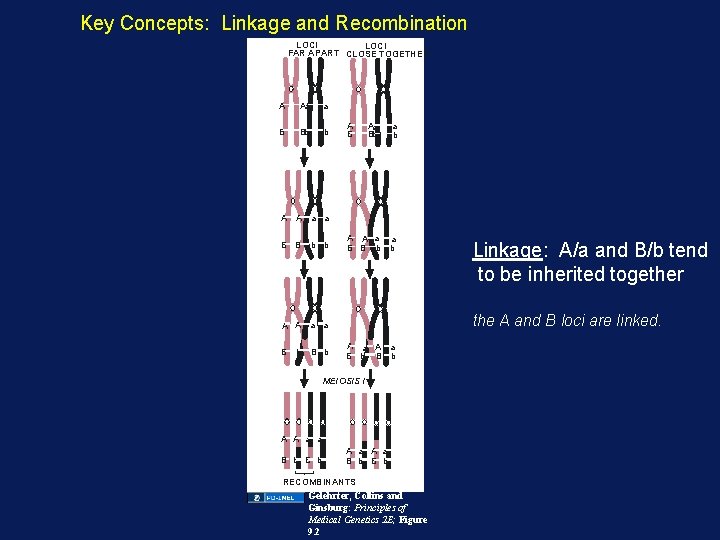
Key Concepts: Linkage and Recombination LOCI FAR APART CLOSE TOGETHER A Aa a B Bb b A A a a B B b b A B Aa Bb a b A A a a B B b b the A and B loci are linked. A A a a B b A a B b MEIOSIS I A A a a B b Linkage: A/a and B/b tend to be inherited together A a B b RECOMBINANTS Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 2
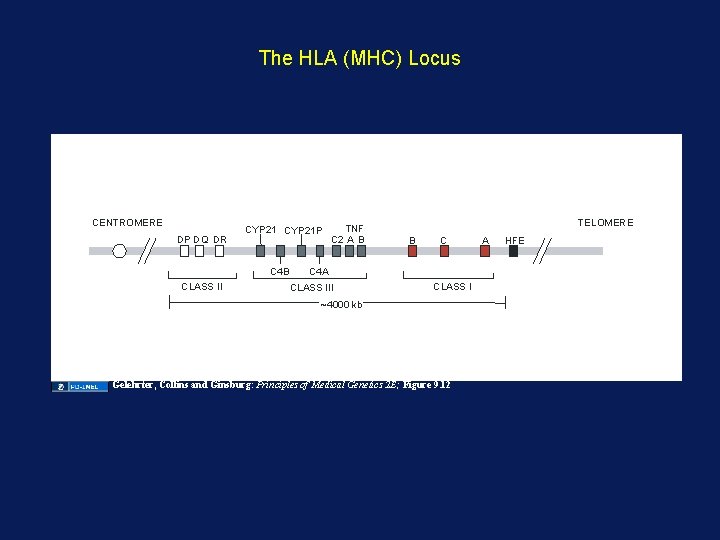
The HLA (MHC) Locus CENTROMERE DP DQ DR CYP 21 P C 4 B CLASS II TNF C 2 A B TELOMERE B C C 4 A CLASS III CLASS I ~4000 kb Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 12 A HFE
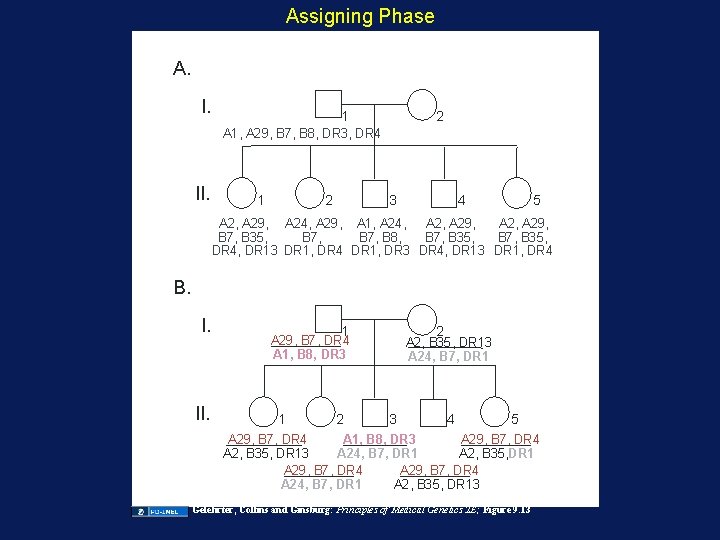
Assigning Phase A. I. 1 A 1, A 29, B 7, B 8, DR 3, DR 4 II. 1 2 A 2, A 24, 2 B 7, B 35, DR 13 3 4 5 A 2, A 29, A 24, A 29, A 1, A 24, A 2, A 29, B 7, B 35, B 7, B 8, B 7, B 35, DR 4, DR 13 DR 1, DR 4 DR 1, DR 3 DR 4, DR 13 DR 1, DR 4 B. I. II. 1 2 A 29, B 7, DR 4 A 1, B 8, DR 3 1 2 A 2, B 35, DR 13 A 24, B 7, DR 1 3 4 5 A 29, B 7, DR 4 A 1, B 8, DR 3 A 29, B 7, DR 4 A 2, B 35, DR 13 A 24, B 7, DR 1 A 2, B 35, DR 1 A 29, B 7, DR 4 A 24, B 7, DR 1 A 2, B 35, DR 13 Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 13
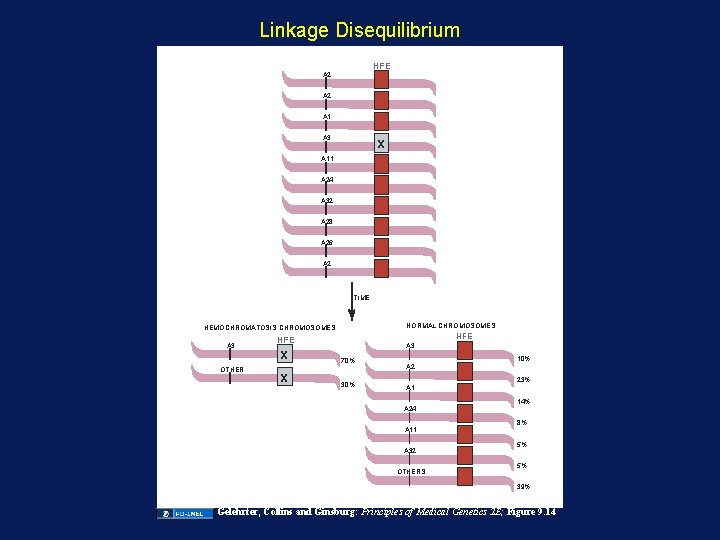
Linkage Disequilibrium HFE A 2 A 1 A 3 X A 11 A 24 A 32 A 28 A 26 A 2 TIME NORMAL CHROMOSOMES HEMOCHROMATOSIS CHROMOSOMES A 3 OTHER HFE X X A 3 70% 30% 10% A 2 A 1 A 24 A 11 A 32 OTHERS 23% 14% 8% 5% 5% 39% Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 9. 14

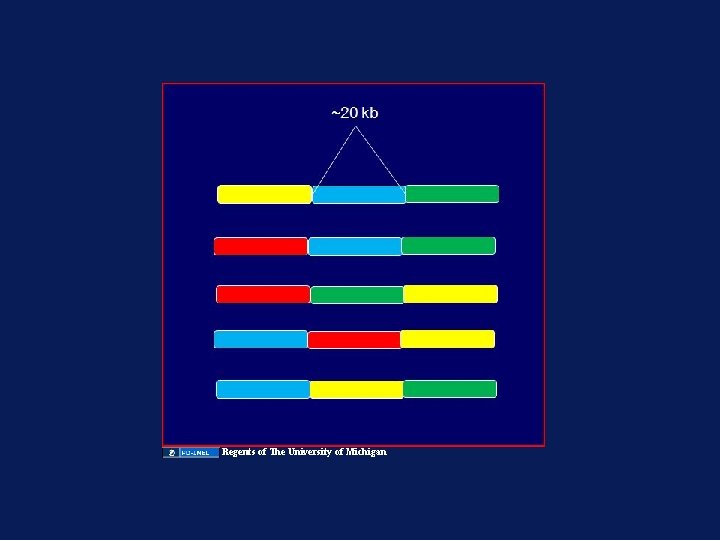
These three SNPs could theoretically occur in 8 different haplotypes …C…A…A… …C…A…G… …C…C…A… …C…C…G… …T…A…A… …T…A…G… …T…C…A… …T…C…G…

But in practice, only two are observed …C…A…A… …C…A…G… …C…C…A… …C…C…G… …T…A…A… …T…A…G… …T…C…A… …T…C…G…

These three variants are said to be in linkage disequilibrium …C…A…A… …C…A…G… …C…C…A… …C…C…G… …T…A…A… …T…A…G… …T…C…A… …T…C…G…
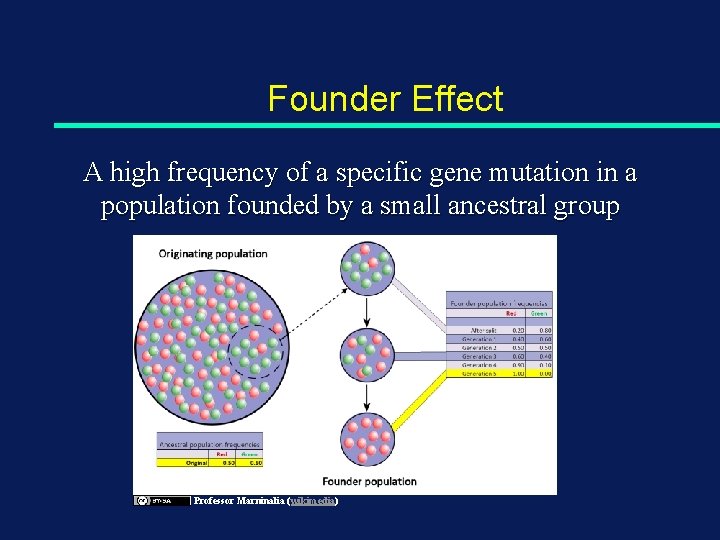
Founder Effect A high frequency of a specific gene mutation in a population founded by a small ancestral group Professor Marninalia (wikimedia)
Regents of The University of Michigan
Regents of The University of Michigan

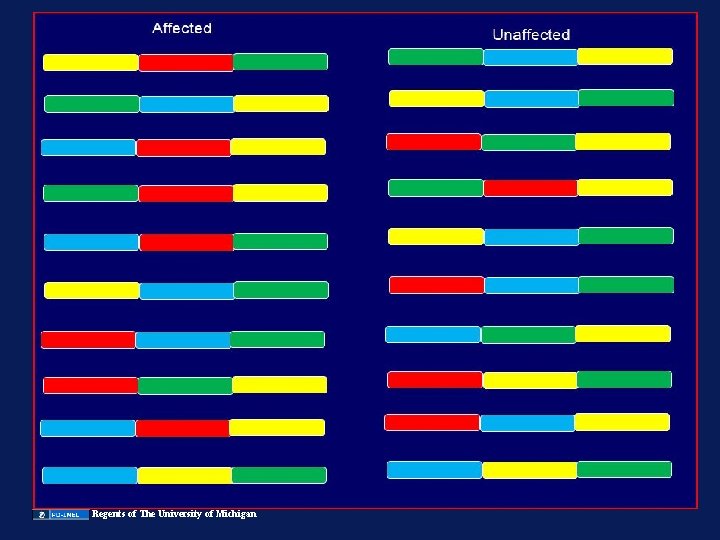
Image of genetic make up of Benin, Bantu, Senegal and Arab-India populations removed Hb S only occurs on 4 haplotypes…only occurred 4 times in history Could we use this approach to find human disease genes (identify specific haplotypes present more often in patients than in controls)?

Complex Diseases • • • Hypertension Coronary artery disease Diabetes Obesity Cancer … Difficult to map in large pedigrees by conventional linkage (multiple genes, variable effects)

• Candidate gene association study – Test a SNP (or SNPs) surrounding your favorite gene for association with disease (more common in patients than controls) • Publication bias • Multiple observations • Population substructure • “looking under the streetlamp” vs. • Genome-wide association study (GWAS) – Unbiased – No prior assumptions
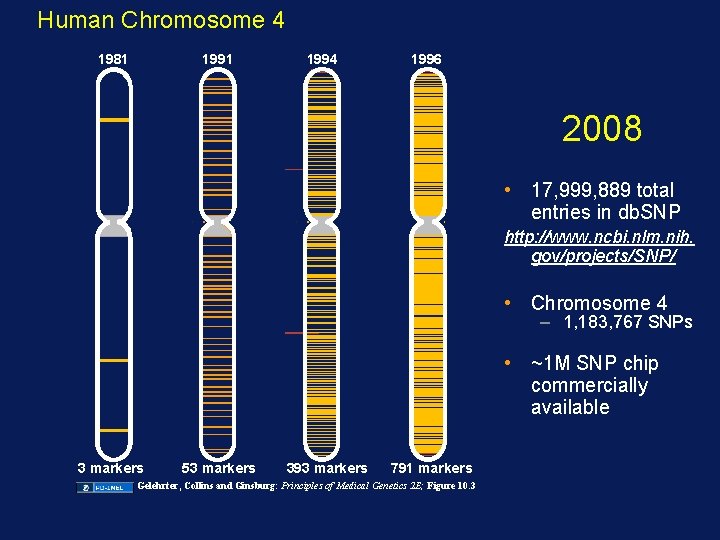
Human Chromosome 4 1981 1994 1996 2008 • 17, 999, 889 total entries in db. SNP http: //www. ncbi. nlm. nih. gov/projects/SNP/ • Chromosome 4 – 1, 183, 767 SNPs • ~1 M SNP chip commercially available 3 markers 53 markers 393 markers 791 markers Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E; Figure 10. 3

Hybridization array: gene chip DNA chip v 6. 0: ~1 million SNPs RNA expression: All ~ 20, 000 genes Ricardipus (flickr) CGH: Survey whole genome for large deletions, insertions, rearrangements

The Multiple observations problem • Roll the dice once – Probability of rolling two 6’s = 2. 8% (1: 36=1/6 X 1/6) • Roll the dice twice – Probability of rolling two 6’s at least once = 5. 5% • Roll the dice 100 times – Probability of rolling two 6’s at least once = 94% Test 1 million SNPs, 100 phenotypes. . .

• Age-related macular degeneration (AMD) – > 10 million cases in the US – Leading cause of blindness among the elderly • Common variant in complement factor H (CFH) gene – – – Tyr 402 His allele = 2 -4 X increase risk of AMD Accounts for 20 -50% of AMD risk

Image of journal article on variant of transcription factor 7 like 2 removed Nature Genetics, 2006, 38: 320 -323. • Odds ratio 1. 7 • Replicated multiple times

Images of journal articles about genome -wide association studies removed
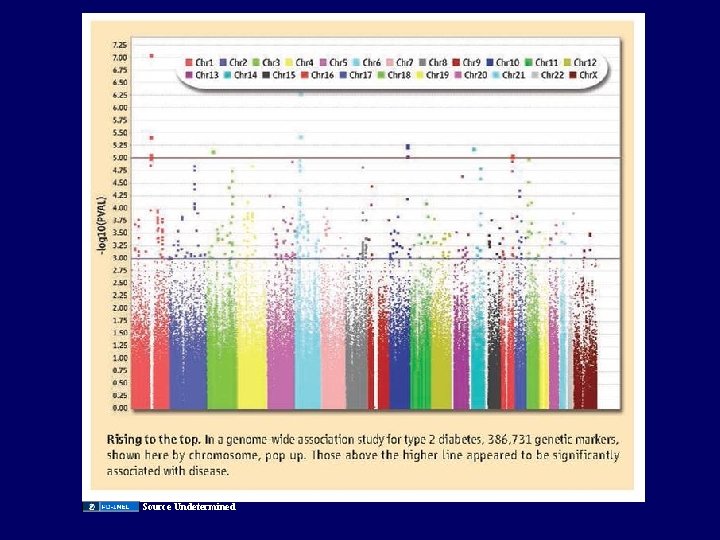
Source Undetermined

How do we distinguish a disease causing mutation from a silent sequence variation? • Obvious disruption of gene – large deletion or rearrangement – frameshift – nonsense mutation • Functional analysis of gene product – expression of recombinant protein – transgenic mice • New mutation by phenotype and genotype • Computer predictions • Disease-specific mutation databases – Same/similar mutation in other patients, not in controls • Rare disease-causing mutation vs. private “polymorphism” (rare variant) X

Other Genome Wide Association Studies (GWAS) • Heart disease – MI, AF, QT prolongation, CAD, lipids • Inflammatory bowel disease • Asthma • Neuropsychiatric disorders – ALS, MS, Alzheimer, schizophrenia, bipolar disorder • Rheumatologic disorders – RA, SLE • Cancer risk – breast, prostate, colon • Common traits – BMA, height, hair/eye/skin color

Lessons from GWAS • Most (nearly all) previous “candidate” gene association studies are wrong • Most common variants have only modest effects on risk (<< 2 fold) • For type II diabetes, variants identified to date only account for ~5% of overall risk– NOT useful clinically (yet) • Other diseases may be different – AMD – Thrombosis • For the future: – ? More predictive, combinatorial analyses – ? Identify new drug targets

GWAS Lessons (continued) • common alleles with small effects: – ? Clinical diagnostic utility – New biologic pathways- ? significance – New drug targets- ? value • Should we be looking for large effect, rare variants instead? – Deep resequencing of extremes of the distribution – New technologies (massively parallel sequencing)

Personalized Medicine: Potential Impact of Genomics • Establish/confirm diagnosis – Subclassification – prognosis • Predisposition testing/prevention • Guiding therapy (pharmacogenomics)

Image of decodme. com homepage removed

Image of DNAdirect. com homepage removed

Image of 23 andme. com homepage removed
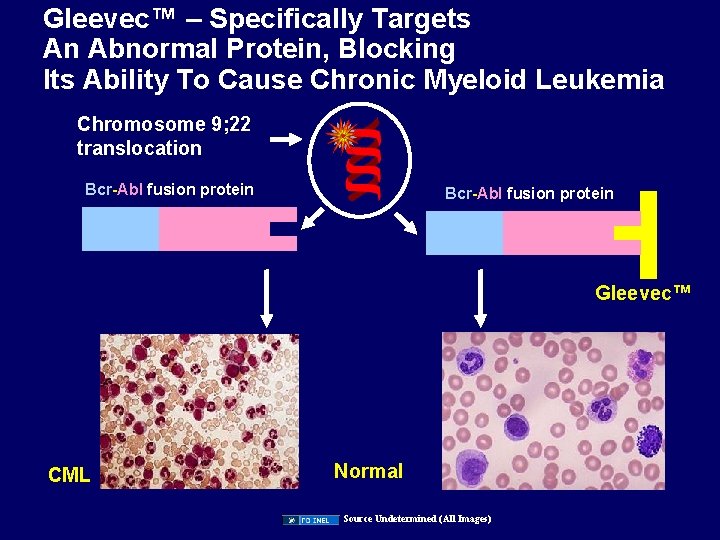
Gleevec™ – Specifically Targets An Abnormal Protein, Blocking Its Ability To Cause Chronic Myeloid Leukemia Chromosome 9; 22 translocation Bcr-Abl fusion protein Gleevec™ CML Normal Source Undetermined (All Images)

Pharmacogenomics today • Thiopurine methyltransferase (TPMT) – 6 -MP, azathioprine therapy dosing • UGT 1 A 1 – irinotecan • Dihydropyrimidine dehydrogenase – 5 -FU • Cytochrome P 450 – warfarin – Multiple other drugs: • antidepressants, tamoxifen, PPIs • VKORC 1 – warfarin

Image of Warfarin mechanism of action removed Sadler, Nature 427: 493, 2004.
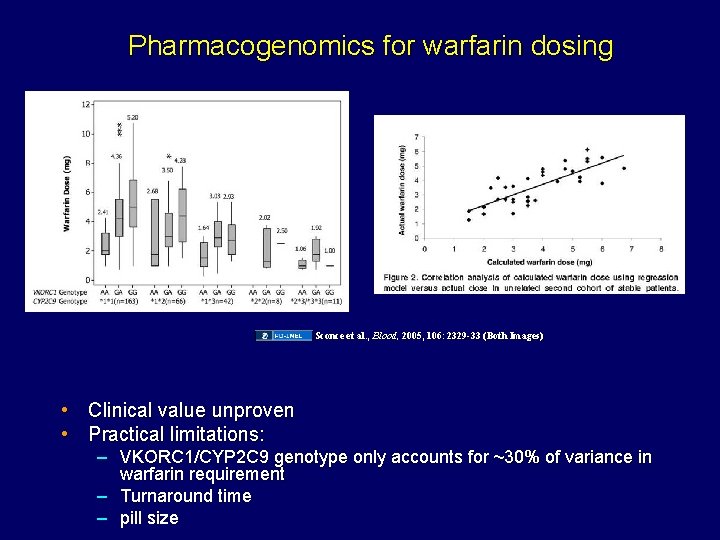
Pharmacogenomics for warfarin dosing Sconce et al. , Blood, 2005, 106: 2329 -33 (Both Images) • Clinical value unproven • Practical limitations: – VKORC 1/CYP 2 C 9 genotype only accounts for ~30% of variance in warfarin requirement – Turnaround time – pill size

The Future… • Will be different – New/improved anticoagulants – Cheaper/faster DNA testing/sequencing – Complex computational analysiscombinatorial risk factors – Much larger data sets– health system wide – Data to support genotype-specific therapy/prophylaxis
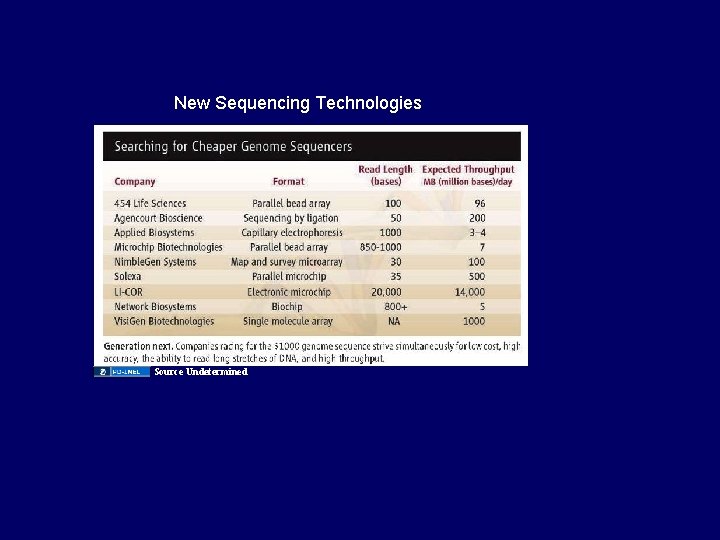
New Sequencing Technologies Source Undetermined

Additional Source Information for more information see: http: //open. umich. edu/wiki/Citation. Policy Slide 6: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 1 Slide 7: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 4 Slide 8: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 2 Slide 9: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 3 Slide 10: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 5 Slide 11: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 6 Slide 12: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 15 Slide 13: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 26 Slide 14: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 9. 31 Slide 16: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E Slide 18: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 5. 22 Slide 19: Adapted from ASCO teaching slides Slide 20: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 10. 3 Slide 21: Source Undetermined Slide 22: Source Undetermined; Andre Karwath (wikipedia); U. S. Federal Government (wikimedia) Slide 24: Source Undetermined Slide 26: National Center for Biotechnology, http: //www. ncbi. nlm. nih. gov/ Slide 27: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E Slide 28: University Of California Santa Cruz, http: //genome. ucsc. edu Slide 29: Levy, et al. Mutations in a member of the ADAMTS gene family cause thrombotic thrombocytopenic purpura. Nature 413: 488 -494, 2001. Slide 30: University Of California Santa Cruz, http: //genome. ucsc. edu Slide 31: National Center for Biotechnology, http: //www. ncbi. nlm. nih. gov/Omim/mimstats. html Slide 41: Regents of The University of Michigan Slide 42: Regents of The University of Michigan Slide 46: Gelehrter, Collins and Ginsburg: Principles of Medical Genetics 2 E, Figure 10. 3 Slide 47: Ricardipus, flickr, http: //creativecommons. org/licenses/by-sa/2. 0/deed. en

Slide 64: Sconce et al. , Blood, 2005, 106: 2329 -33 (Both Images) Slide 66: Source Undetermined Slide 50: Nature Genetics, 2006, 38: 320 -323. Slide 52: Source Undetermined Slide 61: Source Undetermined (All Images)
- Slides: 68