Authored by Peter J Russell DNA Replication Semiconservative

Authored by Peter J. Russell DNA Replication

Semiconservative DNA Replication n The Watson and Crick DNA model implies a mechanism for replication (very important!): ¨ DNA can be denatured ¨ A complementary copy n and separated can be produced for each strand 3 models were proposed for DNA replication: ¨ The conservative model proposed that both strands of one copy would be entirely old DNA, while the other copy would have both strands of new DNA ¨ The dispersive model proposed that ds. DNA would fragment (dangerous idea!), replicate and reassemble. ¨ The semiconservative model proposed that DNA strands separate and a complementary strand is synthesized for each in other words (or pictures)…
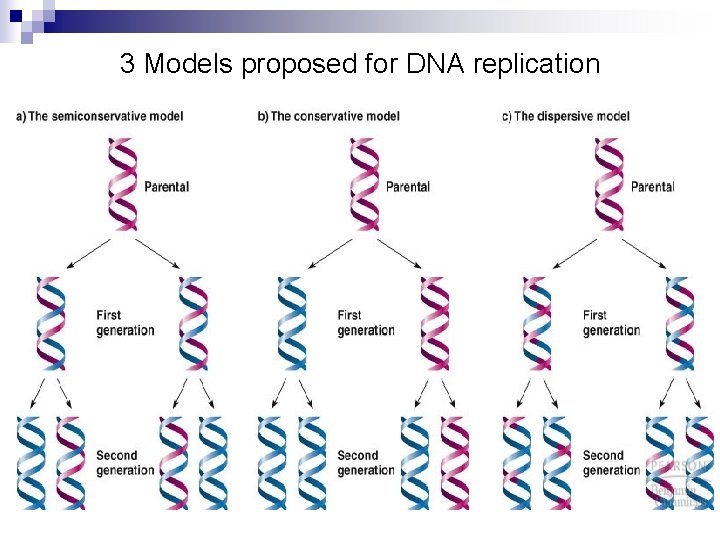
3 Models proposed for DNA replication

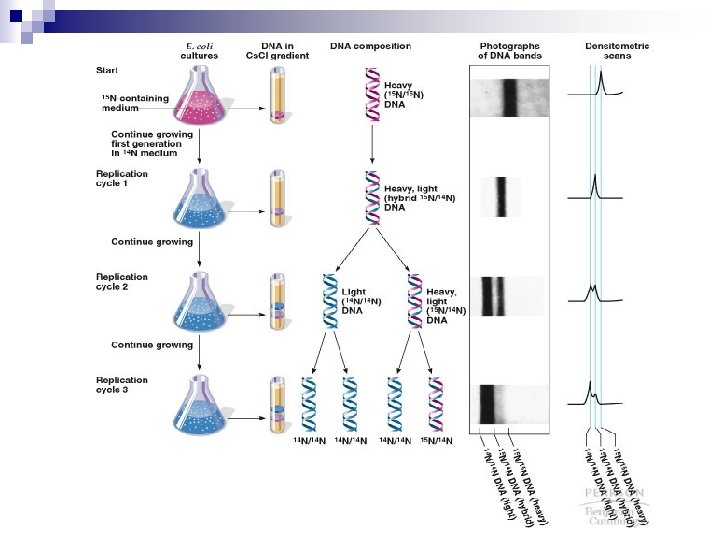
Meselson-Stahl Experiment n Meselson and Stahl grew E. coli in a heavy, nonradioactive isotope of nitrogen. (15 N in the form of 15 NH Cl salt). Because it is heavier, DNA 4 containing 15 N is more dense than DNA with normal 14 N and can be separated by cesium chloride (Cs. Cl) density gradient centrifugation


n n After the E. coli were incubated with heavy 15 N, the investigators shifted the cells to a medium containing normal 14 N, and took cell samples at different time points DNA was then extracted from the treated cells and each sample was subjected to cesium chloride density centrifugation for content analysis Animation: Meselson-Stahl Experiment
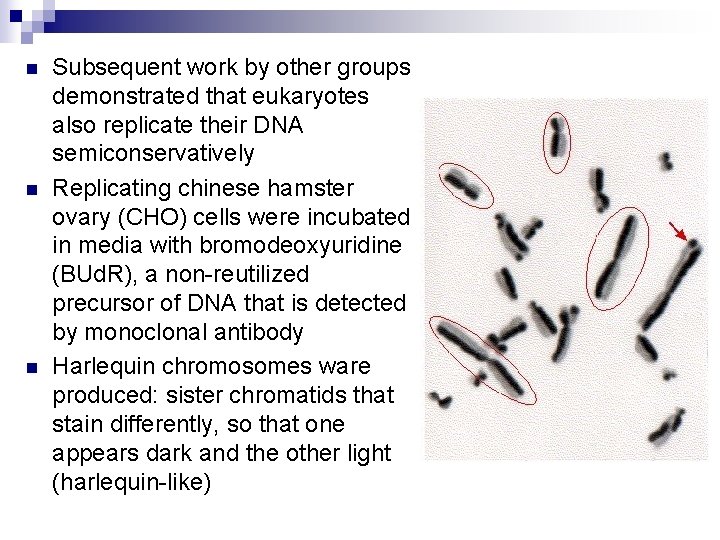
n n n Subsequent work by other groups demonstrated that eukaryotes also replicate their DNA semiconservatively Replicating chinese hamster ovary (CHO) cells were incubated in media with bromodeoxyuridine (BUd. R), a non-reutilized precursor of DNA that is detected by monoclonal antibody Harlequin chromosomes ware produced: sister chromatids that stain differently, so that one appears dark and the other light (harlequin-like)

DNA Polymerases n n Arthur Kornberg isolated the Kornberg enzyme; it was renamed DNA polymerase (thankfully!). His son proved to be a successful biologist as well… In vitro DNA synthesis required 5 components: ¨ All four deoxynucleotide tri-phosphates; d. ATP, d. TTP, d. CTP and d. GTP (substrates) ¨ DNA polymerase I (enzyme) ¨ DNA (template matter) ¨ Primer DNA (is actually RNA in vivo) ¨ Magnesium ions (divalent cations either alter the conformation of the enzyme or substrate, or take part in the chemistry of the catalytic reaction at the active site by stabilizing the intermediate species)
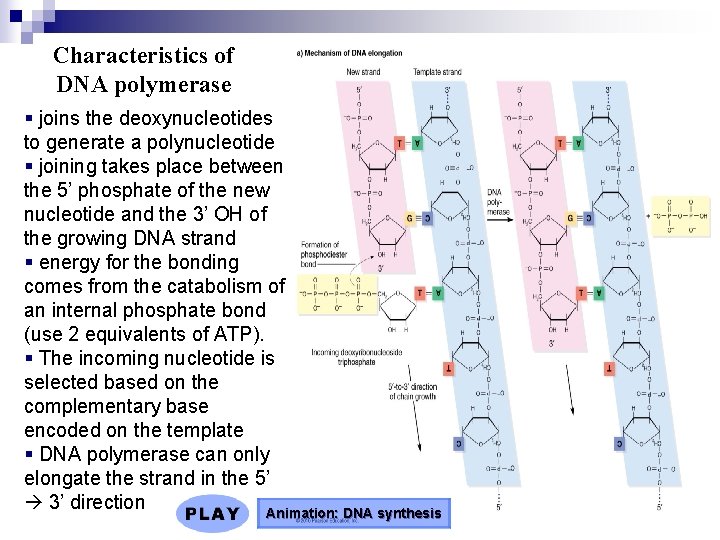
Characteristics of DNA polymerase § joins the deoxynucleotides to generate a polynucleotide § joining takes place between the 5’ phosphate of the new nucleotide and the 3’ OH of the growing DNA strand § energy for the bonding comes from the catabolism of an internal phosphate bond (use 2 equivalents of ATP). § The incoming nucleotide is selected based on the complementary base encoded on the template § DNA polymerase can only elongate the strand in the 5’ 3’ direction Animation: DNA synthesis

Prokaryotic DNA Polymerase Family n DNA pol I ¨ ¨ n DNA Pol II, IV, V ¨ ¨ n Replicates DNA in the 5’ 3’ direction Composed of a single peptide 3’ to 5’ exonuclease activity (proofreading) 5’ to 3’ exonuclease activity (primer removal) Single peptide enzyme DNA repair functions DNA Pol III Replicates DNA in the 5’ 3’ direction Catalytic core is composed of 3 peptide subunits (a, e, q) Complete holoenzyme is composed of an additional 6 peptide subunits (making the total 9 peptides) ¨ 5’ to 3’ exonuclease activity ¨ 3’ to 5’ exonuclease activity ¨ No repair function is observed ¨ ¨ ¨

Molecular Model of DNA Replication n n The copying of DNA is remarkable in its speed and accuracy, using a dozen enzymes and other proteins Replication begins at special DNA sequences called replicators within origins of replication, where the two DNA strands are separated by origin of replication enzymes, opening a replication “bubble” with 2 “forks” A eukaryotic chromosome has thousands of origins of replication; prokaryotes have one origin (called Ori. C) Replication proceeds in both directions from each origin, until the entire molecule is copied Animation: Origins of Replication
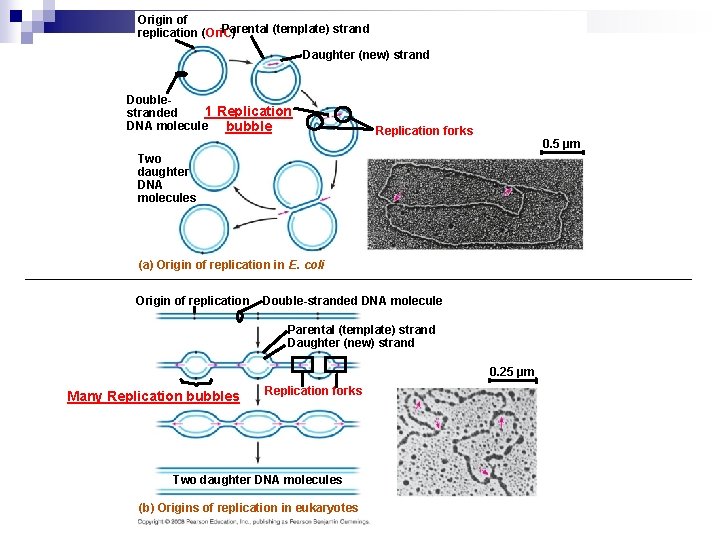
Origin of Parental (template) strand replication (Ori. C) Daughter (new) strand Double 1 Replication stranded DNA molecule bubble Replication forks 0. 5 µm Two daughter DNA molecules (a) Origin of replication in E. coli Origin of replication Double-stranded DNA molecule Parental (template) strand Daughter (new) strand 0. 25 µm Many Replication bubbles Replication forks Two daughter DNA molecules (b) Origins of replication in eukaryotes

Players involved in DNA Replication n Origin of Replication Enzymes (Initiator Proteins) recognize the start site on template DNA and initially open up the DNA (single strands) n Helicases untwist the double helix at the replication forks n Primase begins DNA replication by laying an an RNA primer (10 nt long) for DNA polymerase to start at n Single-strand binding proteins bind to and stabilize the single-stranded DNA until it can be used as a template n DNA polymerase polymerizes new DNA in a 5’ to 3’ direction, proofreads the newly laden DNA, excises error ridden DNA and replaces that DNA with the correct sequence. Speed/fidelity: mutates 1 X 10 -10 n Topoisomerase corrects “overwinding” ahead of replication forks by breaking, swiveling, and rejoining DNA strands n Terminal replication proteins stall replication fork progress for polymeriation to catch-up n DNA ligase anneals nucleotides together, creating a DNA strand
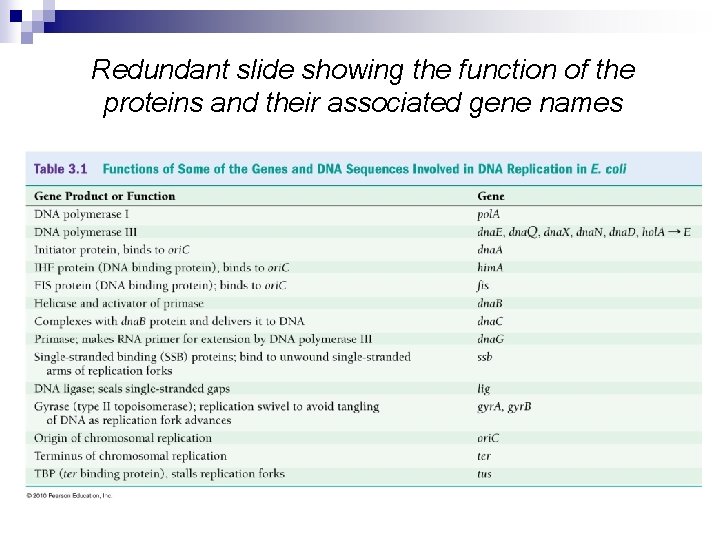
Redundant slide showing the function of the proteins and their associated gene names
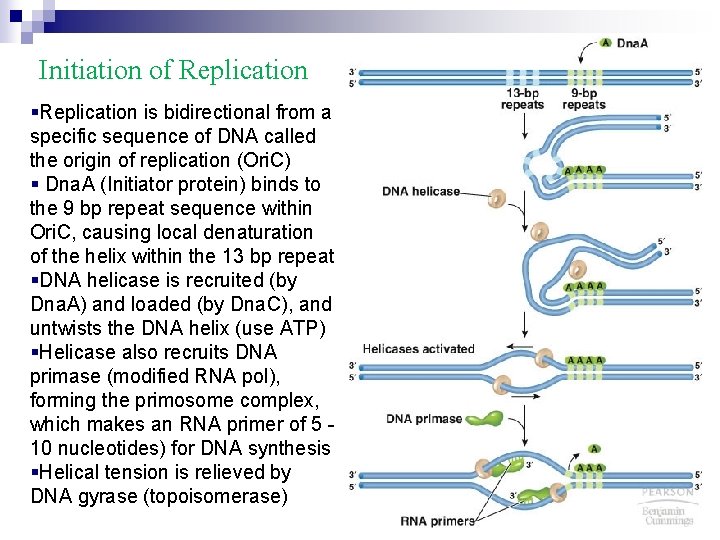
Initiation of Replication §Replication is bidirectional from a specific sequence of DNA called the origin of replication (Ori. C) § Dna. A (Initiator protein) binds to the 9 bp repeat sequence within Ori. C, causing local denaturation of the helix within the 13 bp repeat §DNA helicase is recruited (by Dna. A) and loaded (by Dna. C), and untwists the DNA helix (use ATP) §Helicase also recruits DNA primase (modified RNA pol), forming the primosome complex, which makes an RNA primer of 5 10 nucleotides) for DNA synthesis §Helical tension is relieved by DNA gyrase (topoisomerase)

Semidiscontinuous DNA Replication n n The antiparallel structure of the double helix, with two strands oriented in opposite directions, introduces an interesting paradox to replication DNA polymerases add nucleotides only to the free 3 end of a growing strand; therefore, a new DNA strand can elongate only in the 5 to 3 direction Along one template strand of DNA, DNA polymerase synthesizes a leading strand continuously, moving toward the replication fork To elongate the other new strand, called the lagging strand, DNA polymerase must work in the direction away from the replication fork The lagging strand is synthesized as a series of segments called Okazaki fragments, which are joined together by DNA ligase (semidiscontinuous process) (cool experiment! – see book)
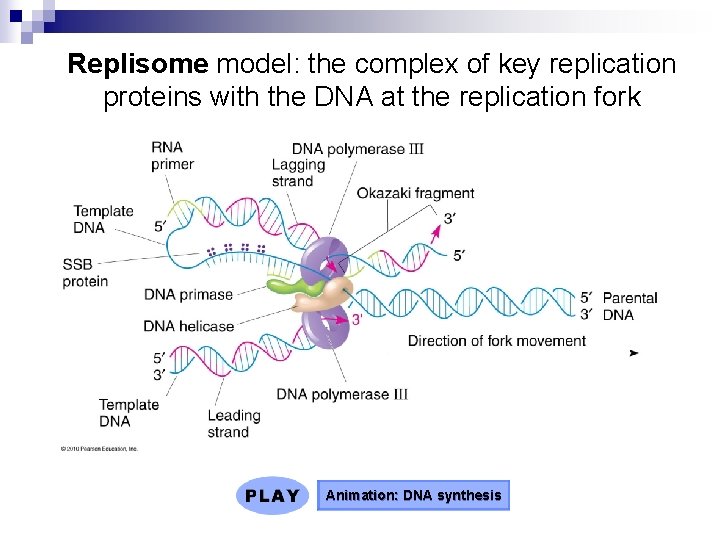
Replisome model: the complex of key replication proteins with the DNA at the replication fork Animation: DNA synthesis
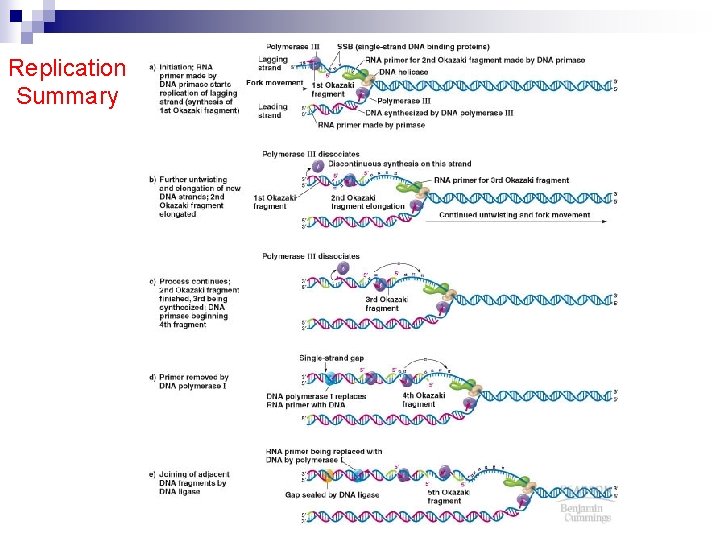
Replication Summary
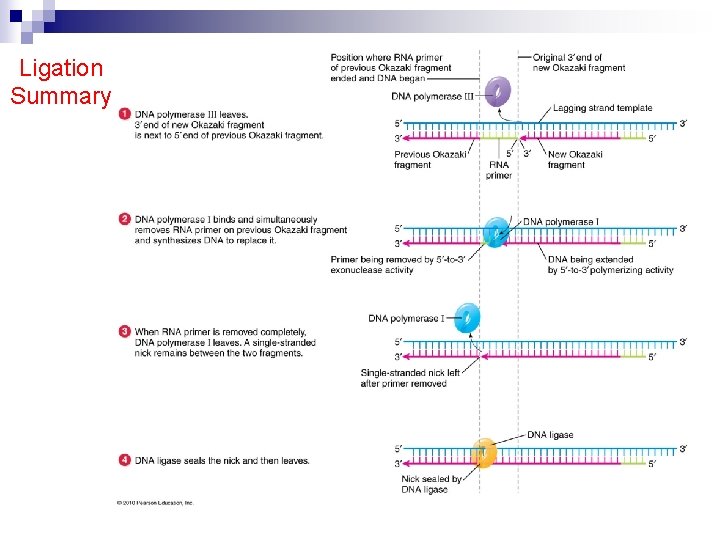
Ligation Summary
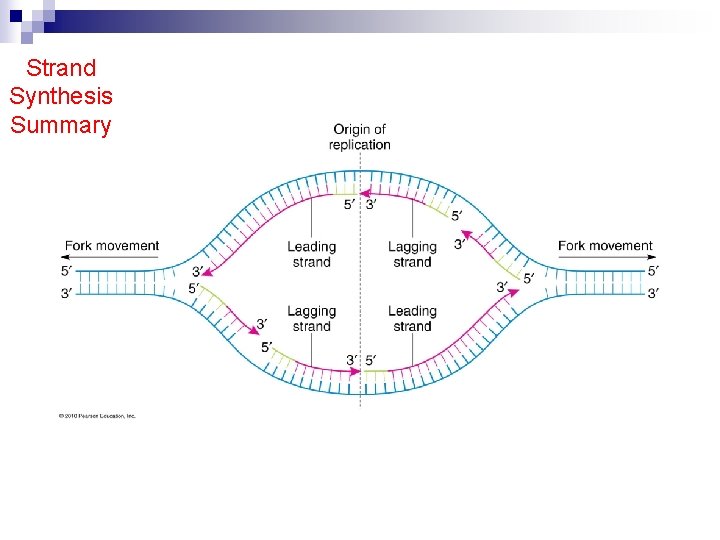
Strand Synthesis Summary
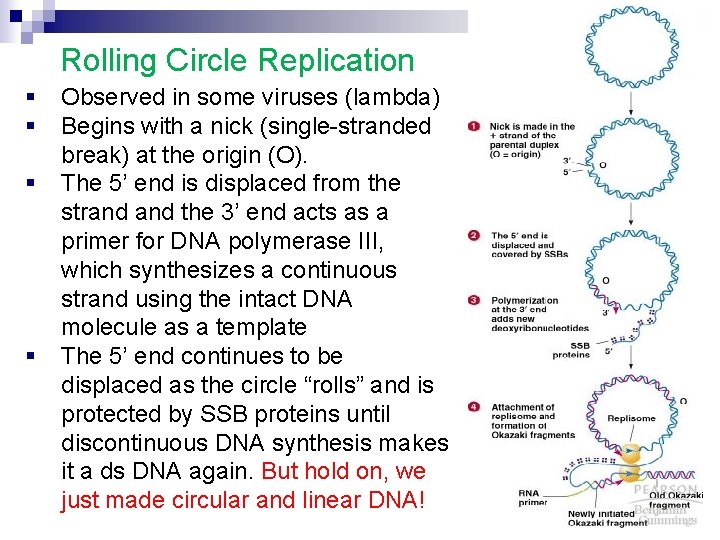
Rolling Circle Replication § § Observed in some viruses (lambda) Begins with a nick (single-stranded break) at the origin (O). The 5’ end is displaced from the strand the 3’ end acts as a primer for DNA polymerase III, which synthesizes a continuous strand using the intact DNA molecule as a template The 5’ end continues to be displaced as the circle “rolls” and is protected by SSB proteins until discontinuous DNA synthesis makes it a ds DNA again. But hold on, we just made circular and linear DNA!
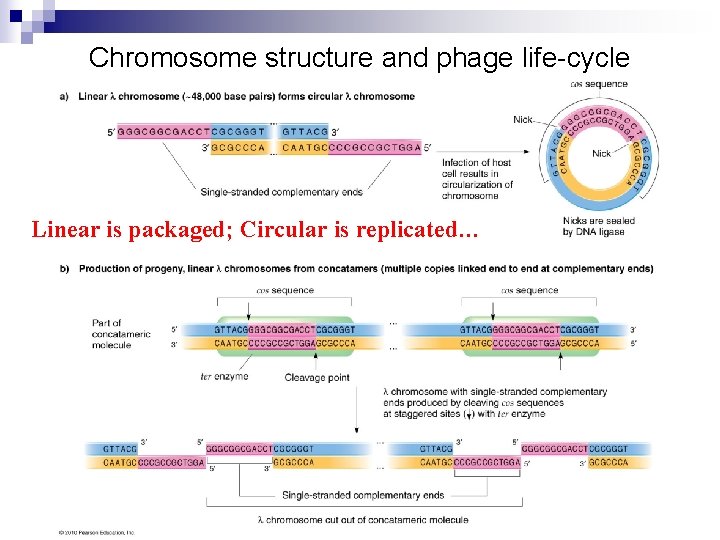
Chromosome structure and phage life-cycle Linear is packaged; Circular is replicated…

DNA Replication in Eukaryotes n n Prokaryotic and eukaryotic DNA replication is similar but for the size, linearity, rate and number of eukaryotic chromosomes Eukaryotes have multiple origins of replication. The DNA replicated from a single origin is called a replicon Replicon size is smaller in eukaryotes. Replication is slower and each chromosome contains many replicons which are activated at distict times Each replicon is used only once when DNA is replicated in the S phase of cell cycle. How you ask? Good question…

Keeping DNA replication in eukaryotes real… n Replicators are less well defined in higher eukaryotes ¨ n Yeast have Autonomously Replicating Sequences (ARSs) In eukaryotes, the key issue is not to use any origin more than once (this would be absolutely disastrous)! To avoid this, these steps are followed: ¨ ¨ First, replicator selection occurs. The multisubunit initiator protein, Origin Recognition Complex (ORC), binds to the replicator during the G 1 phase of the cell cycle Next, other initiator proteins (including licencing factors, which are produced only during G 1) assemble on the replicator to form pre-replicative complexes (pre-RC). When the cell enters the S phase, specific cyclin dependent kinases (Cdks) are present, causing a release and deactivation of licencing factors, allowing the DNA helix to unwind, thus initiating replication Importantly, the presence of these specific Cdks inhibits another round of pre-RC formation until the cell reenters G 1 after a completed round of DNA replication

Eukaryotic DNA polymerase (blah, blah…) n n 15 DNA polymerases are known in mammalian cells Three DNA polymerases are used to replicate nuclear DNA. Pol a (alpha) extends the 10 -nt RNA primer by about 30 nt. Pol d (delta) and Pol e (epsilon) extend the RNA/DNA primers, one on the leading strand the other on the lagging strand (it is not clear which synthesizes which) Other DNA pols replicate mitochondrial or chloroplast DNA or are used in DNA repair Primer removal differs from that in prokaryotes. Pol d continues extension of the newer Okazaki fragment, displacing the RNA and producing a flap that is removed by nucleases, thus allowing the Okazaki fragments to be joined by DNA ligase
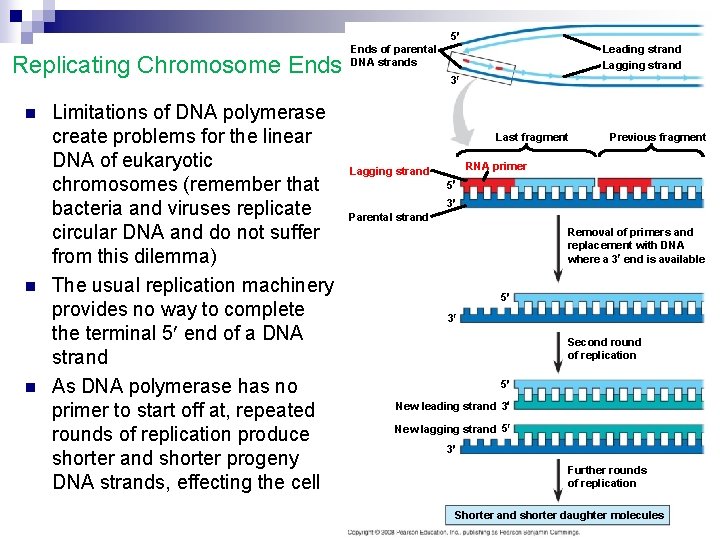
5 Replicating Chromosome Ends n n n Limitations of DNA polymerase create problems for the linear DNA of eukaryotic chromosomes (remember that bacteria and viruses replicate circular DNA and do not suffer from this dilemma) The usual replication machinery provides no way to complete the terminal 5 end of a DNA strand As DNA polymerase has no primer to start off at, repeated rounds of replication produce shorter and shorter progeny DNA strands, effecting the cell Ends of parental DNA strands Leading strand Lagging strand 3 Last fragment Previous fragment RNA primer Lagging strand 5 3 Parental strand Removal of primers and replacement with DNA where a 3 end is available 5 3 Second round of replication 5 New leading strand 3 New lagging strand 5 3 Further rounds of replication Shorter and shorter daughter molecules

the solution… n n n It has been proposed the shortening of telomeres contributes to aging Dilemma: If chromosomes of germ cells became shorter in every cell cycle, the essential genes would eventually be missing from the gametes they produce, destroying the heritability of the species Eukaryotic chromosomal DNA molecules have terminal nucleotide sequences called telomeres (5’ TTAGGG 3’). Telomeres do not prevent shortening of DNA; they postpone genetic erosion near the ends of DNA Solution: An enzyme called telomerase catalyzes the lengthening of telomeres in germ cells (sperm and egg cells), thus protecting the integrity of reproduction Of note: The shortening of telomeres protects somatic cells from cancerous growth by limiting the number of cell divisions (and hence compiled mutations). There is evidence of telomerase activity in cancer cells, allowing cancer cells to persist

§ § Elizabeth Blackburn and Carol Greider have shown that chromosome lengths are maintained by telomerase The enzyme is composed of protein and RNA. The RNA component (451 bases in humans) includes an 11 base template RNA sequence that is used for the synthesis of new telomere repeat DNA. Thus, telomerase acts as a reverse transcriptase (TERT) The 3’CAAUC 5’ sequence on RNA interacts with the 5’GTTAG 3’ sequence on DNA. The remaining 3’CCAAUC 5’ sequence on RNA acts as a template to fill in the 5’GGTTAG 3’ sequence on DNA. The process repeats and telomerase drops off The alternative strand of DNA gets filled in using DNA as a template
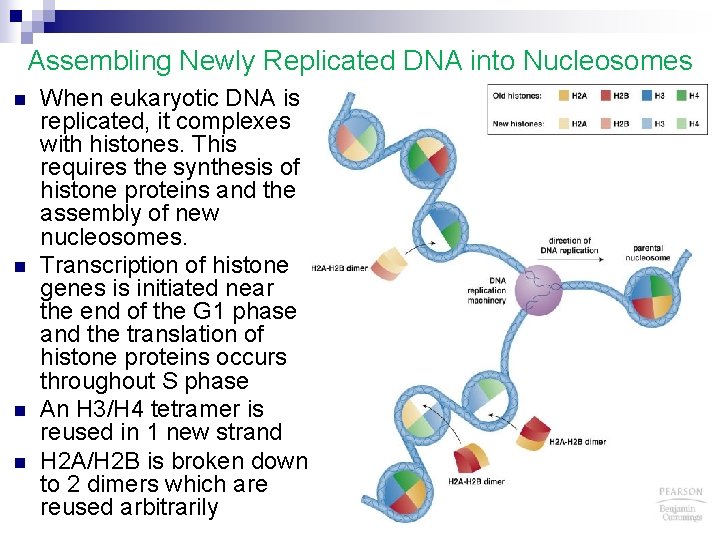
Assembling Newly Replicated DNA into Nucleosomes n n When eukaryotic DNA is replicated, it complexes with histones. This requires the synthesis of histone proteins and the assembly of new nucleosomes. Transcription of histone genes is initiated near the end of the G 1 phase and the translation of histone proteins occurs throughout S phase An H 3/H 4 tetramer is reused in 1 new strand H 2 A/H 2 B is broken down to 2 dimers which are reused arbitrarily

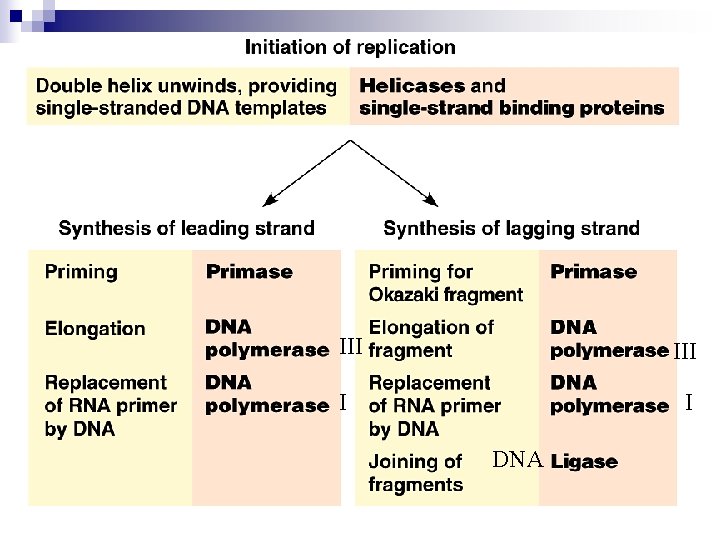
III I I DNA
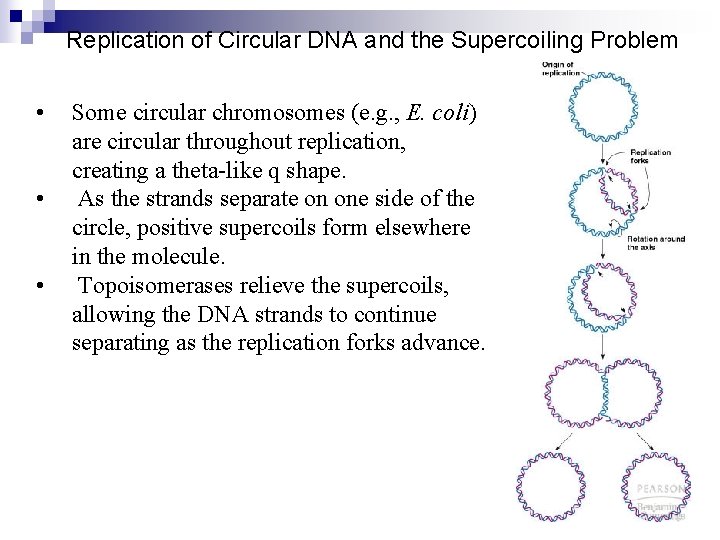
Replication of Circular DNA and the Supercoiling Problem • • • Some circular chromosomes (e. g. , E. coli) are circular throughout replication, creating a theta-like q shape. As the strands separate on one side of the circle, positive supercoils form elsewhere in the molecule. Topoisomerases relieve the supercoils, allowing the DNA strands to continue separating as the replication forks advance.
- Slides: 33