Assigning Numbers to the Arrows Parameterizing a Gene

Assigning Numbers to the Arrows Parameterizing a Gene Regulation Network by using Accurate Expression Kinetics

Overview • • • Motivation Gene Regulation Networks Background Our Goal Our Example Parameterizing Algorithm Results

Motivation • Understand regulation factors for different genes • Can help understand a gene’s function • If we can understand how it all works we can use it for medical purposes like fixing and preventing DNA damage!
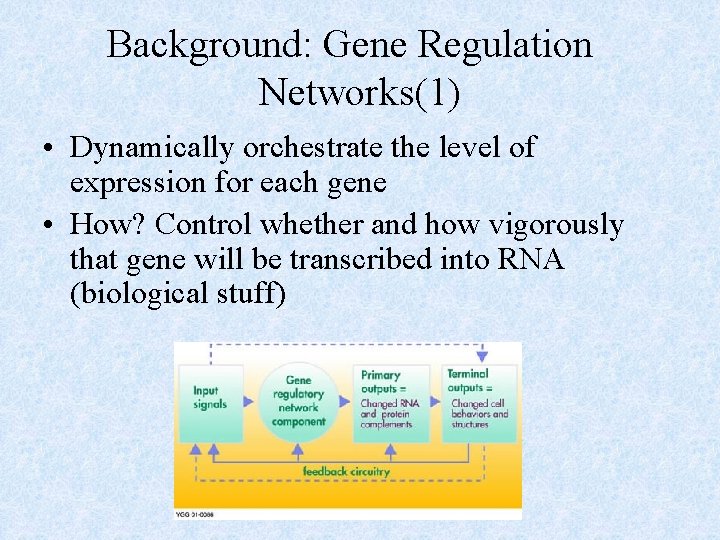
Background: Gene Regulation Networks(1) • Dynamically orchestrate the level of expression for each gene • How? Control whether and how vigorously that gene will be transcribed into RNA (biological stuff)

Background: Gene Regulation Networks(2) • Contains: 1. Input Signals: environmental cues, intracellular signals 2. Regulatory Proteins 3. Target Genes

Our Goal • Assign parameters to a Gene Regulation Network based on experiments: - production of unrepressed promoter. the maximum production - concentration of repressor at half maximal repression. The bigger it is the earlier the gene becomes active and the later it becomes inactive again

Our Example(1) • Escheria coli bacterium • SOS DNA repair system – used to repair damage done by UV light • 8 (out of about 30) gene groups (operons)
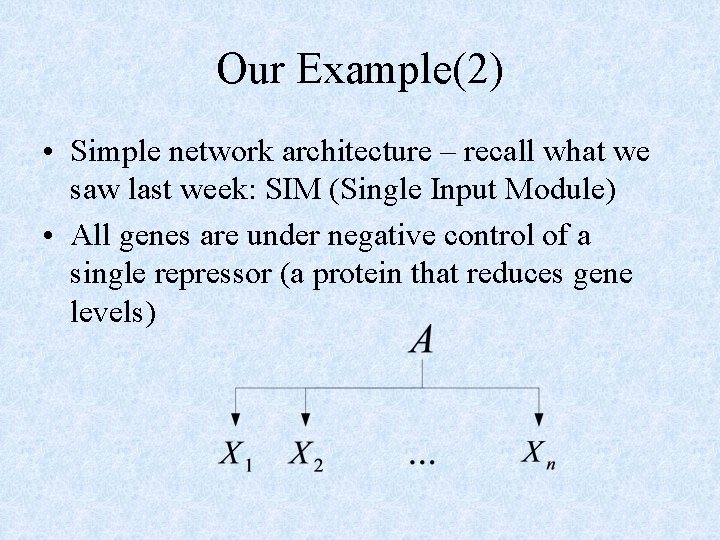
Our Example(2) • Simple network architecture – recall what we saw last week: SIM (Single Input Module) • All genes are under negative control of a single repressor (a protein that reduces gene levels)

Parametrization Algorithm Definitions: - the activity of promoter i in experiment j as function of time - effective repressor concentration in experiment j as function of time - production rate of the unrepressed promoter i - k parameter of promoter i

Parametrization Algorithm 1: Trial Function Why? Michaelis-Menten form: a very useful equation in modeling biological behavior.

Parametrization Algorithm 2: Data Preprocessing(1) • Smoothing the signals using a hybrid Gaussian -median filter with a window size of five measurements: Five time points are taken, sorted and the average of central three points is taken to be the signal.

Parametrization Algorithm 2: Data Preprocessing(2) Some more definitions: - the activity of promoter i as a function of time - GFP fluorescence from the corresponding reporter as a function of time - corresponding Optical Density as a function of time

Parametrization Algorithm 2: Data Preprocessing(3) • The signal is smooth enough to be differentiated • The activity of promoter i is proportional to the number of GFP molecules produced per unit time per cell

Parametrization Algorithm 2: Data Preprocessing(4) • The activity signal is smoothed by a polynomial fit of sixth order to: • The smoothing procedure captures the dynamics well, while removing noise • Data for all experiments is concatenated and normalized by the maximal activity for each operon
![Parametrization Algorithm 3: Parameter Determination(1) • To determine parameters in equation [1] based on Parametrization Algorithm 3: Parameter Determination(1) • To determine parameters in equation [1] based on](http://slidetodoc.com/presentation_image/bf60d0a066b722274337cdf53c704936/image-15.jpg)
Parametrization Algorithm 3: Parameter Determination(1) • To determine parameters in equation [1] based on experimental data we transform it into a bilinear form: where:

Parametrization Algorithm 3: Parameter Determination(2) • Now, the matrix where N is for genes and M for time points, is modeled by two vectors of size N: and one vector of size M: • 2 N*M variables

Parametrization Algorithm 3: Parameter Determination(3) – some algebra • The standard method of least mean squares solution for such a problem uses SVD (Singular Value Decomposition) • The mean over i of is removed:

Parametrization Algorithm 3: Parameter Determination(4) – some algebra • A(t) is the SVD eigenvector with the largest eigenvalue of the matrix: This is the covariance matrix • Results for A(t) are normalized to fit the constraints: • Alternative normalization: add points with A=0 and

Parametrization Algorithm 3: Parameter Determination(5) – some algebra • Perform a second round of optimization for by using a nonlinear least mean squares solver to minimize

Parametrization Algorithm 4: Error Evaluation(1) • The mean error for promoter i is given by: where T is the total time of the experiment • This is considered the quality of the data model in describing the data

Parametrization Algorithm 4: Error Evaluation(2) • The error estimate for the parameters is determined by using a graphic method: is plotted vs. A(t)

Parametrization Algorithm 4: Error Evaluation(3) • From maximal and minimal slopes of the graphs the error for is determined • From maximal and minimal intersections with the y axis the error for is determined

Parametrization Algorithm 5: Additional Trial Function(1) • An extension of the model to the case of cooperative binding – a regulator can be a repressor for some genes and an activator for others, and with different measures:

Parametrization Algorithm 5: Additional Trial Function(2) -Hill coefficient for operon i Hill coefficient? A coefficient that describes binding - repression - activation - no cooperation

Parametrization Algorithm 5: Additional Trial Function(3) Our example: good comparison between measured results and those calculated with trial function suggest there may be no significant cooperativity in the repressor action

Results: Promoter Activity Profiles(1) • After about half a cell cycle the promoter activities begin to decrease • Corresponds to the repair of damaged DNA
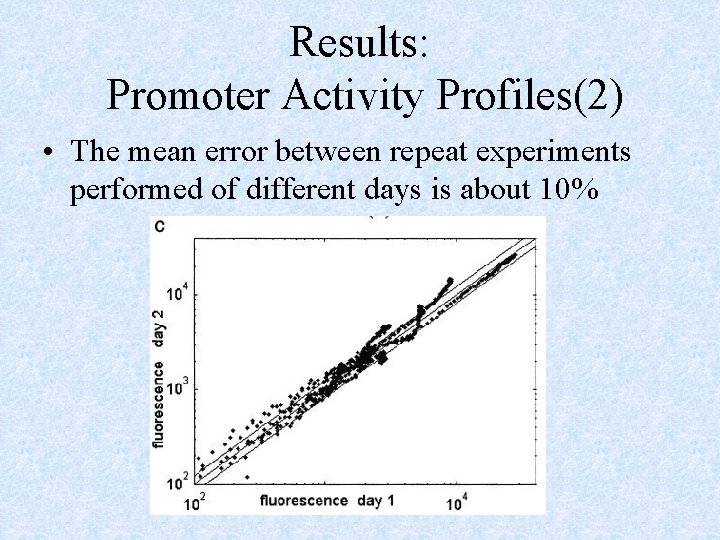
Results: Promoter Activity Profiles(2) • The mean error between repeat experiments performed of different days is about 10%

Results: Assigning Effective Kinetic Parameters • The error is under 25% for most promoters

Results: Detection of Promoters with Additional Regulation • Relatively large error may help to detect operons that have additional regulation. • Examples: 1. lac. Z – very large error (150%) 2. uvr. Y – recently found to participate in another system and to be regulated by other transcription factors (45% error)

Results: Determining Dynamics of an Entire System Based on a Single Representative(1) • Once the parameters are determined for each operon, we need to measure only the dynamics of one promoter in a new experiment to estimate all other SOS promoter kinetics

Results: Determining Dynamics of an Entire System Based on a Single Representative(2) • The estimated kinetics using data from only one of the operons agree quite well with the measured kinetics for all operons • Same level of agreement found by using different operons as the base operon
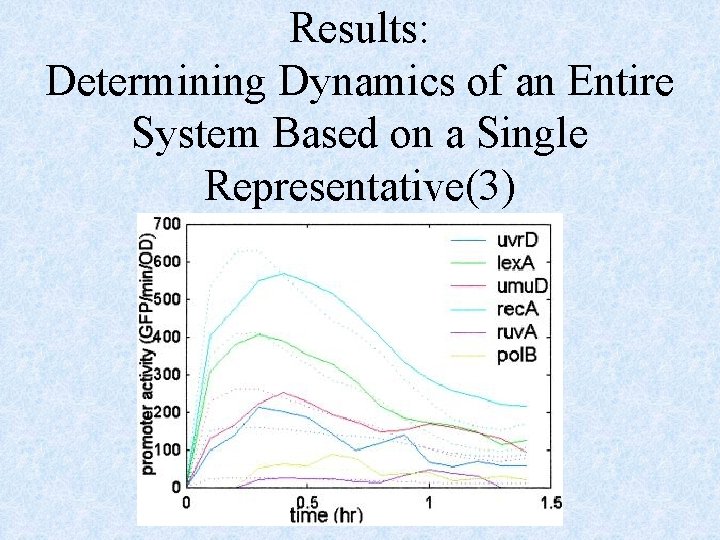
Results: Determining Dynamics of an Entire System Based on a Single Representative(3)

Results: Repressor Protein Concentration Profile • Current measurements don’t directly measure the concentration of the proteins produced by these operons, only the rate at which the corresponding m. RNA’s are produced • The parameterization algorithm allows calculation of the transcriptional repressor A(t), directly.

Summary • We can apply the current method to any SIM motif, in gene regulation networks • The method won’t work with multiple regulatory factors

Questions? Thank You For Listening!
- Slides: 35