Architectural TFs MBV 4230 Overview DNAbinding TFs General

Architectural TFs
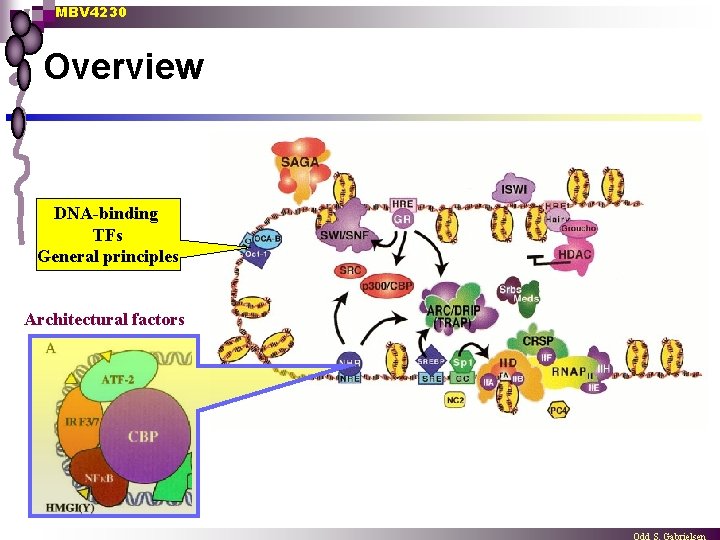
MBV 4230 Overview DNA-binding TFs General principles Architectural factors
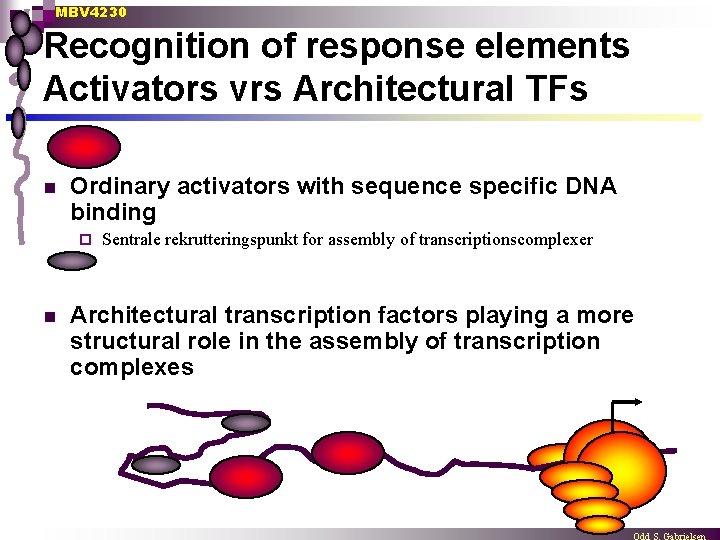
MBV 4230 Recognition of response elements Activators vrs Architectural TFs n Ordinary activators with sequence specific DNA binding ¨ n Sentrale rekrutteringspunkt for assembly of transcriptionscomplexer Architectural transcription factors playing a more structural role in the assembly of transcription complexes
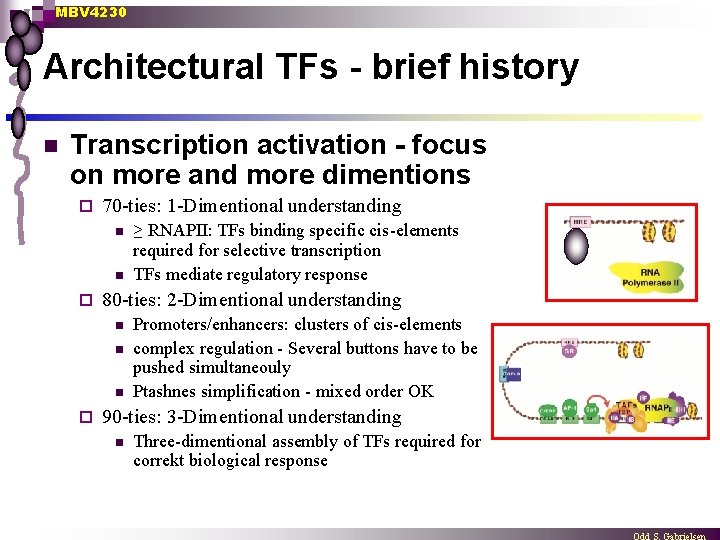
MBV 4230 Architectural TFs - brief history n Transcription activation - focus on more and more dimentions ¨ 70 -ties: 1 -Dimentional understanding n n ¨ 80 -ties: 2 -Dimentional understanding n n n ¨ ≥ RNAPII: TFs binding specific cis-elements required for selective transcription TFs mediate regulatory response Promoters/enhancers: clusters of cis-elements complex regulation - Several buttons have to be pushed simultaneouly Ptashnes simplification - mixed order OK 90 -ties: 3 -Dimentional understanding n Three-dimentional assembly of TFs required for correkt biological response

MBV 4230 3 D protein-promoter complexes - factors dedicated architecture n some factors has a pure architectural function designated architectural transcription factors n They lack a transactivation domain (TAD) n Do not function out of their natural context (in contrast to ordinary acitvators) n Their function is to confer a specific 3 D structure on DNA n
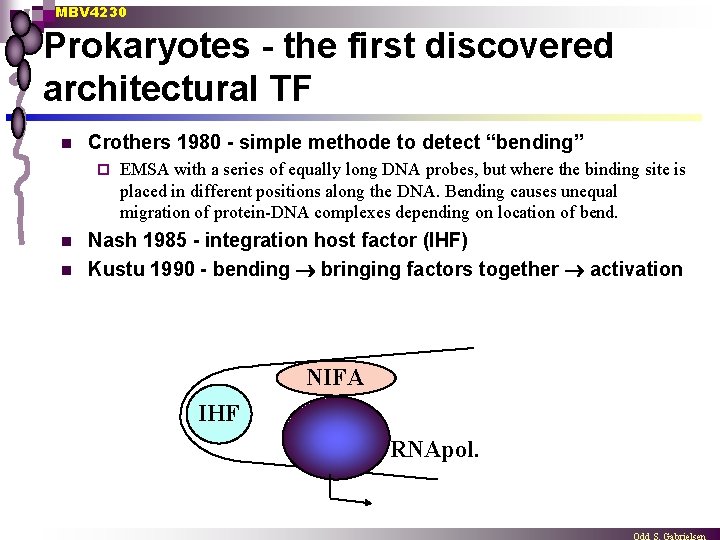
MBV 4230 Prokaryotes - the first discovered architectural TF n Crothers 1980 - simple methode to detect “bending” ¨ n n EMSA with a series of equally long DNA probes, but where the binding site is placed in different positions along the DNA. Bending causes unequal migration of protein-DNA complexes depending on location of bend. Nash 1985 - integration host factor (IHF) Kustu 1990 - bending bringing factors together activation NIFA IHF RNApol.

MBV 4230 Classical HMG-proteins n non-histone chromatin proteins - original defining criteria ¨ ¨ ¨ high mobility in PAGE soluble in 2 -5% TCA small < 30 k. Da High content of charged amino acids abundant: 1 per. 10 -15 nucleosomes

MBV 4230 Classical HMG-proteins n n Three classes of HMG DNA-binding proteins HMGB HMG-box family Eks: HMG 1 and HMG 2 ¨ Bends DNA substantially ¨ Facilitators of nucleoprotein complexes ¨ n HMG-AT-hook family HMGA Eks. : HMGI(Y) ¨ Antagonizing intrinsic distortions in the conformation of AT-rich DNA ¨ n HMG-nucleosome binding family HMGN Eks. : HMG 14 and 17 ¨ Mediates moderate destabilization of chromatin higher-order structure ¨ Not present in yeast or fly ¨

HMGB-proteins
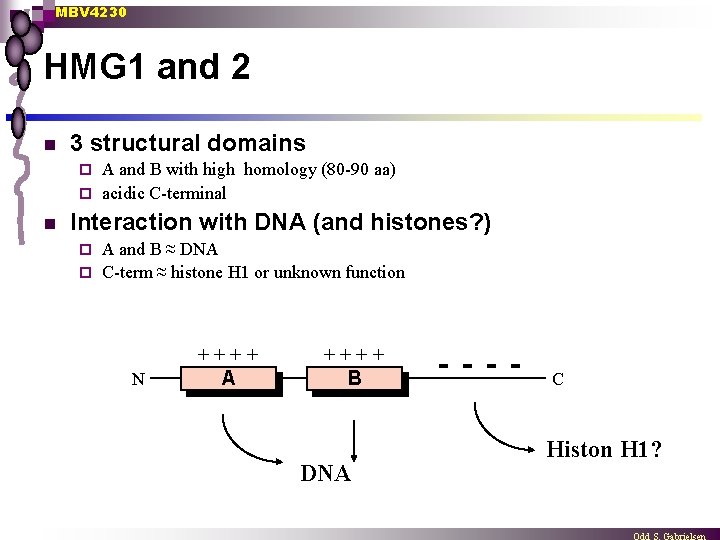
MBV 4230 HMG 1 and 2 n 3 structural domains A and B with high homology (80 -90 aa) ¨ acidic C-terminal ¨ n Interaction with DNA (and histones? ) A and B ≈ DNA ¨ C-term ≈ histone H 1 or unknown function ¨ N ++++ A ++++ B DNA - - C Histon H 1?
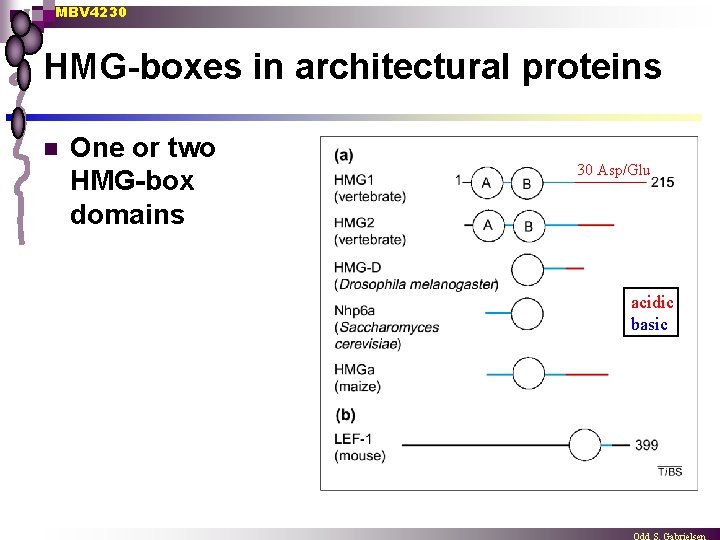
MBV 4230 HMG-boxes in architectural proteins n One or two HMG-box domains 30 Asp/Glu acidic basic

MBV 4230 First eukaryotic architectural TF: LEF 1 (Grosschedl 1992) n n n LEF 1: a cell type-specific TF LEF 1 contains an HMG-related domain LEF 1: a sequence-specific TF that binds CCTTTGAAG ¨ n n found in enhancer of TCR LEF 1 induces strong bending of DNA - about 130 o Induced bending brings nearly TFs in contact
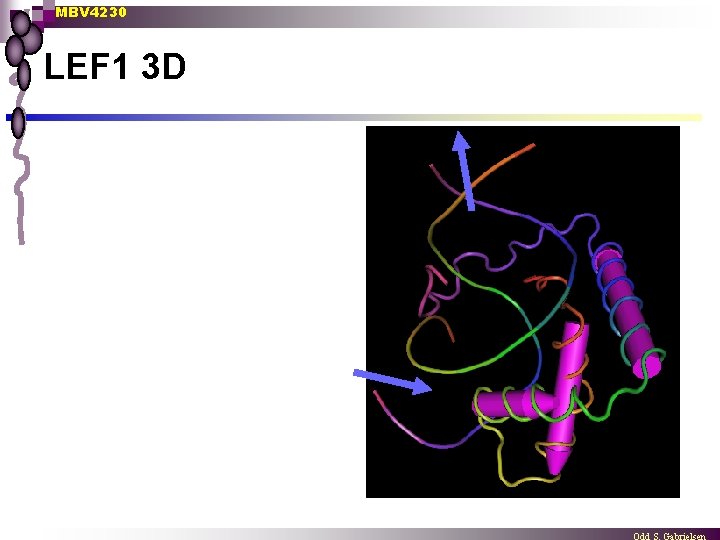
MBV 4230 LEF 1 3 D
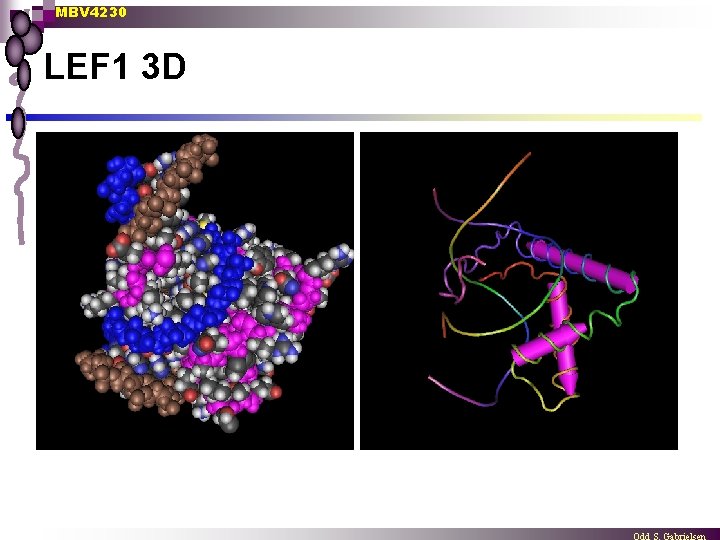
MBV 4230 LEF 1 3 D

MBV 4230 A whole family of architectural TFs with HMG-domains n UBF has repeated HMG-homologous repeats 4 -6 ex dimer ≈ 10 HMG-like domains ¨ activator of r. RNA gener ¨ UBF-DNA complex scaffold for SL-1 recruitment n Interaction with 180 bp that is packed into a distinct structure ¨ n DNA-motif in a series of TFs: “HMG-box” designate the DNA-sequence-motif ¨ “HMG-domain” designate the protein motif ¨

MBV 4230 Two subclasses of HMG-domain proteins n Proteins with multiple HMG-domains low sequence-specificity ¨ Ubiquitous - found in all cell types ¨ eks. : HMG 1, HMG 2, ABF-2, UBF ¨ n Proteins with enkelt HMG-domain (moderate) sequence-specificity ¨ Cell type-specific ¨ eks. : LEF-1, SRY, TCF-1, Sox, Mat- 1, Ste 11, Rox 1 ¨
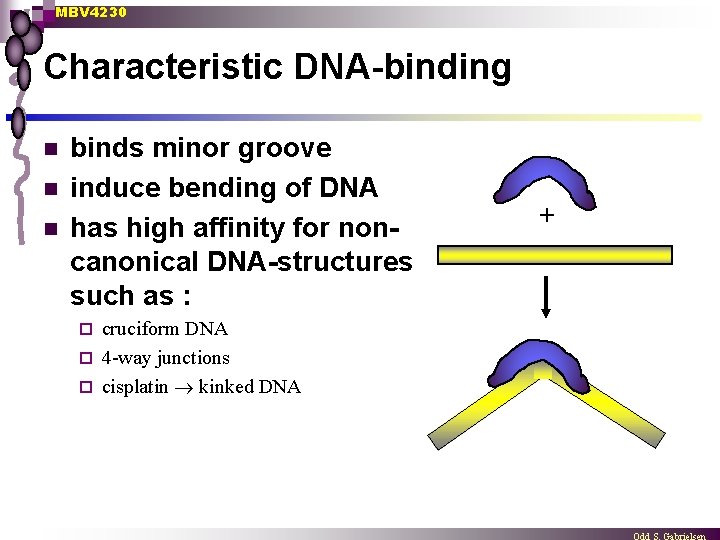
MBV 4230 Characteristic DNA-binding n n n binds minor groove induce bending of DNA has high affinity for noncanonical DNA-structures such as : cruciform DNA ¨ 4 -way junctions ¨ cisplatin kinked DNA ¨ +

MBV 4230 NMR-structures n n n HMG 1 B-domain LEF-1 SRY Yeast Nhp 6 p Drosophila HMG-D Common: 3 helix L-form heliks II and III form an angle of about 80 o ¨ Conserved aromatic aa in kink ¨ Basic concave side interact with DNA ¨
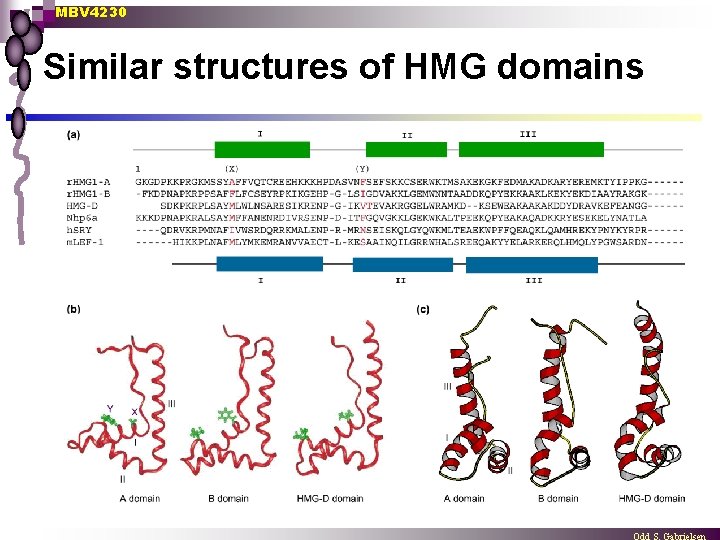
MBV 4230 Similar structures of HMG domains
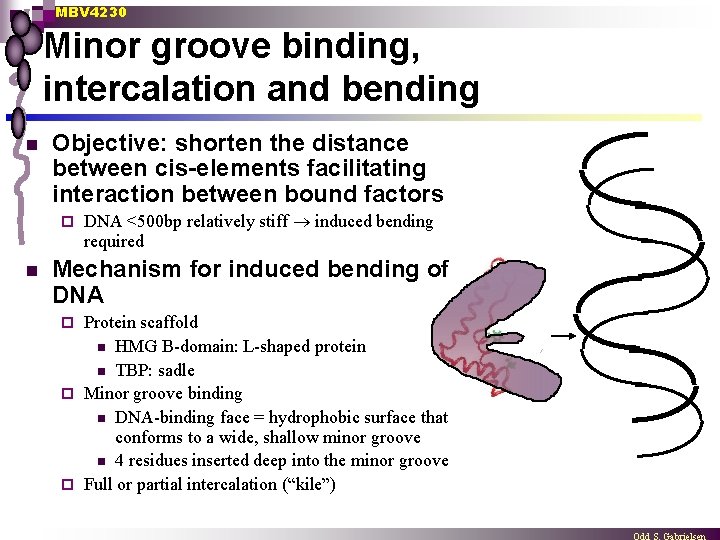
MBV 4230 Minor groove binding, intercalation and bending n Objective: shorten the distance between cis-elements facilitating interaction between bound factors ¨ n DNA <500 bp relatively stiff induced bending required Mechanism for induced bending of DNA Protein scaffold n HMG B-domain: L-shaped protein n TBP: sadle ¨ Minor groove binding n DNA-binding face = hydrophobic surface that conforms to a wide, shallow minor groove n 4 residues inserted deep into the minor groove ¨ Full or partial intercalation (“kile”) ¨

MBV 4230 Intercalation in protein-induced DNA-bending n Partial intercalation in the DNA helix of a protein side chain introduces a kink in the DNA enhancing the bend Large hydrophobic residues (N-term helix I) partially intercalates between two base pairs ¨ The A-box HMG domain has only an Ala in the X position not large enough to intercalate, ¨ n Intercalation linked to bending also seen in other factors ¨ Partial (TBP) n n ¨ Full (ETS 1) n n n Inserted side chain unstacks two basepairs side chain as stacking-partner side chain penetrates into the helix side chain (Trp) as new stacking-partner Result: helix axis direction altered
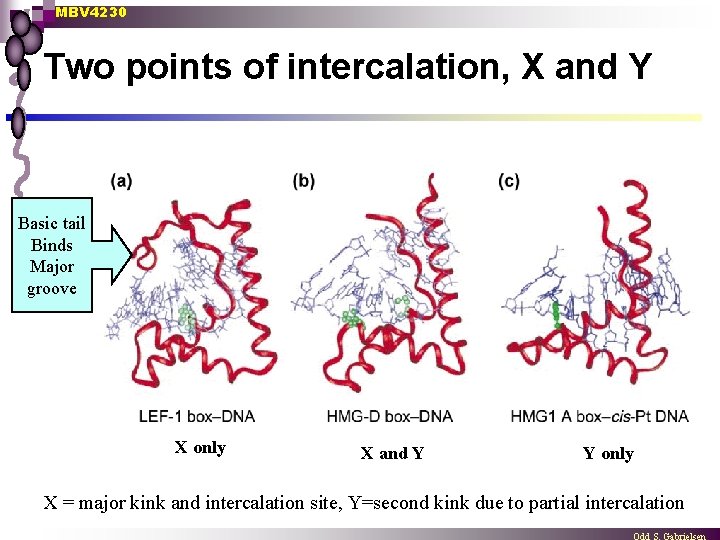
MBV 4230 Two points of intercalation, X and Y Basic tail Binds Major groove X only X and Y Y only X = major kink and intercalation site, Y=second kink due to partial intercalation
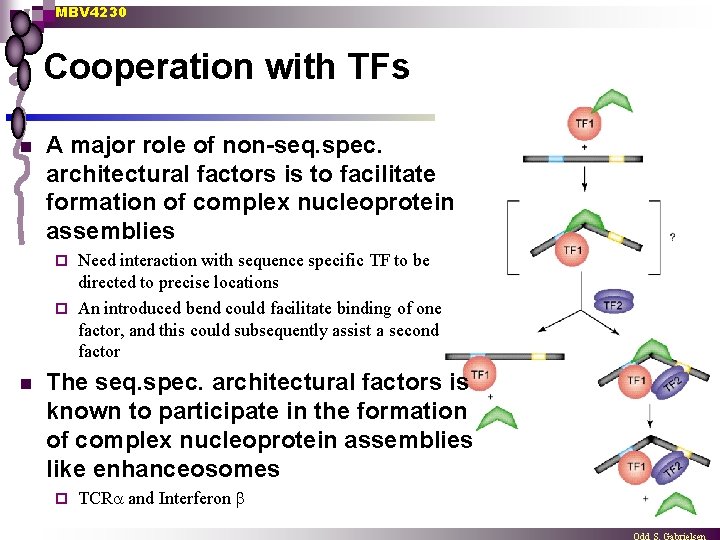
MBV 4230 Cooperation with TFs n A major role of non-seq. spec. architectural factors is to facilitate formation of complex nucleoprotein assemblies Need interaction with sequence specific TF to be directed to precise locations ¨ An introduced bend could facilitate binding of one factor, and this could subsequently assist a second factor ¨ n The seq. spec. architectural factors is known to participate in the formation of complex nucleoprotein assemblies like enhanceosomes ¨ TCR and Interferon b

MBV 4230 Are all TFs architectural? n A large number of publications “TFx bends DNA” positive reports “TFx bends DNA” ¨ negative reports “TFx does not bend DNA” ¨ n n All TFs that bind on one side of DNA will induce bending due to one-sided neutralization of charge Degree of bending will depend on ionic condition Uncertain if biologically relevant The term “Architectural TFs“ should be reserved for factors with a particularly developed bending mechanism

MBV 4230 The charge neutralization model

2. subgruppe: HMGA. . First described by Søren Laland, an almost forgotten discovery

MBV 4230 HMGA - proteins with AT-hook n The mammalian HMGI/Y (HMGA) proteins participate in a wide variety of cellular processes ¨ n All members have multiple copies of a DNA-binding motif called the `AT hook' ¨ n including regulation of gene trx and induction of neoplastic transformation and promotion of metastatic progression. that binds to the narrow minor groove of stretches of AT-rich sequence. The proteins have little secondary structure in solution but assume distinct conformations when bound to DNA or other proteins ¨ Their flexibility allows the HMGI/Y proteins to induce both structural changes in chromatin substrates and the formation of stereospecific complexes called `enhanceosomes'. Reciprocal conformational changes occur in both the HMGI/Y proteins themselves and in their interacting substrates.

MBV 4230 Members n 4 known members Alternatively splicing gives rise to two isoform proteins, HMGA 1 a (HMGI) and HMGA 1 b (HMGY). These two are identical in sequence except for a deletion of 11 residues between the first and second AT hook in the latter. Alternative splicing also produces HMGA 1 c. ¨ The related HMGA 2 (HMGI-C) protein is coded for by a separate gene. ¨ n Conserved ¨ Homologues of the mammalian HMGA proteins have been found in yeast, insects, plants and birds, as well as in all mammalian species examined.
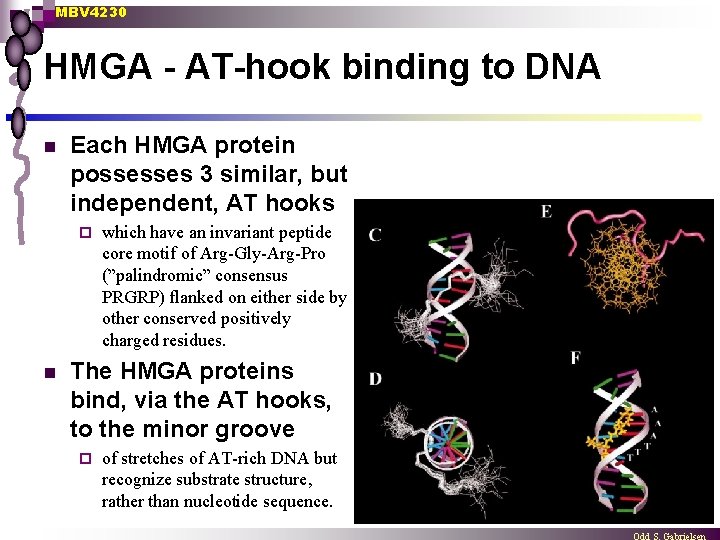
MBV 4230 HMGA - AT-hook binding to DNA n Each HMGA protein possesses 3 similar, but independent, AT hooks ¨ n which have an invariant peptide core motif of Arg-Gly-Arg-Pro (”palindromic” consensus PRGRP) flanked on either side by other conserved positively charged residues. The HMGA proteins bind, via the AT hooks, to the minor groove ¨ of stretches of AT-rich DNA but recognize substrate structure, rather than nucleotide sequence.

MBV 4230 HMGA proteins heavily modified n The HMGA proteins are among the most highly phosphorylated proteins in the mammalian nucleus. Cell cycle-dependent phosphorylation pga cdc 2 activity in the G 2/M phase of the cycle. ¨ Sites: T 53 and T 78 situated at the N-terminal ends of the 2. and 3. AT-hook. Phosphorylation significantly reduces (>20 -fold) DNA binding. ¨ n n HMGA proteins are the downstream targets of a number of signal transduction pathways that lead to phosphorylation. HMGA proteins are also acetylated ¨ n n at Lys 65 by CBP and at Lys 71 by PCAF …as well as methylated and poly-ADP ribosylated Hypothesis: Modifications may alter DNA-binding specificity?

MBV 4230 Architectural effects n Architectural effects ¨ n Binding of full-length HMGA proteins can bend, straighten, unwind and induce loop formation in linear DNA molecules in vitro. Multiple contact points with DNA may alter conformation of DNA A single AT-hook preferentially binds to stretches of 4 -6 bp of AT-rich sequence, and partially neutralizes the negatively charged backbone phosphates on only one face of the DNA helix. ¨ The number and spacing of AT-rich binding sites in DNA influences the conformation of bound DNA and the biological effects elicited. ¨ n HMGA may also induce conformational change in proteins ¨ HMGA forms protein-protein interactions with other transcription factors, which alters the 3 D structure of the factors resulting in enhanced DNA binding and transcriptional activation.
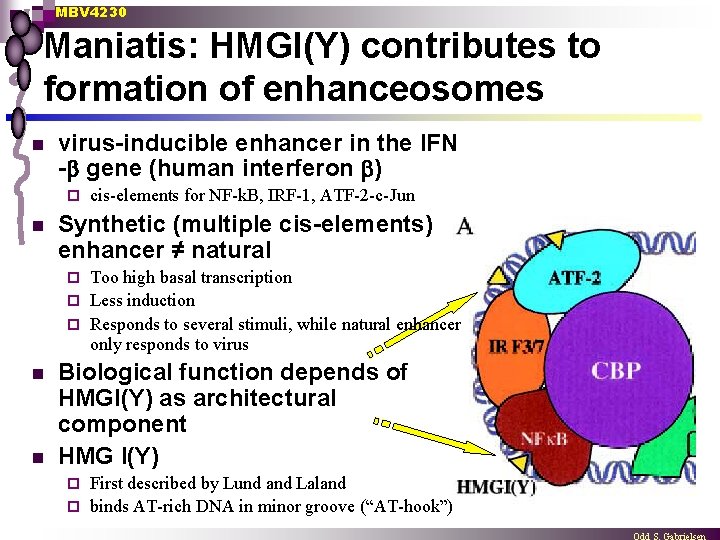
MBV 4230 Maniatis: HMGI(Y) contributes to formation of enhanceosomes n virus-inducible enhancer in the IFN - gene (human interferon ) ¨ n cis-elements for NF-k. B, IRF-1, ATF-2 -c-Jun Synthetic (multiple cis-elements) enhancer ≠ natural Too high basal transcription ¨ Less induction ¨ Responds to several stimuli, while natural enhancer only responds to virus ¨ n n Biological function depends of HMGI(Y) as architectural component HMG I(Y) First described by Lund and Laland ¨ binds AT-rich DNA in minor groove (“AT-hook”) ¨
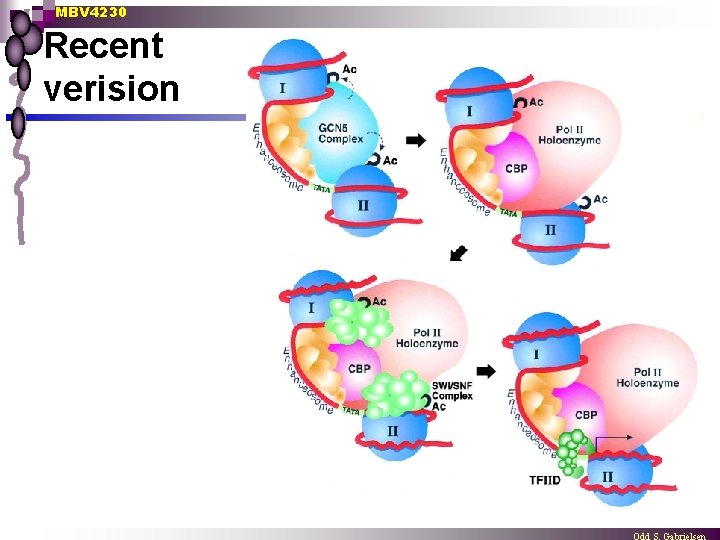
MBV 4230 Recent verision

MBV 4230 Other functions of HMGA proteins n HMGA and cancer ¨ n HMGI/Y proteins are also involved in a diverse range of other cellular processes including pathologic processes such as neoplastic transformation and metastatic progression. Chromosomal translocations in a long 3. intron Intron 3 of the HMGA 2 genes is extremely long (>25 kb in human and >60 kb in mouse) and separates the three exons that contain the AT hook motifs from the remainds of the 3´untranslated tail region of the gene. ¨ Translocation within the exceptionally long third intron are commonly observed in benign mesenchymal tumors. ¨

3. subgruppe: HMGN

MBV 4230 HMGN proteins n Three functional domains of the HMGN proteins: a bipartite nuclear localization signal (NLS), ¨ a nucleosomal binding domain (NBD) ¨ and a chromatin-unfolding domain (CHUD). The CHUD domain has a net negative charge. ¨ n Binding of HMGN proteins to nucleosomes decreases the compactness of chromatin, and facilitates trx
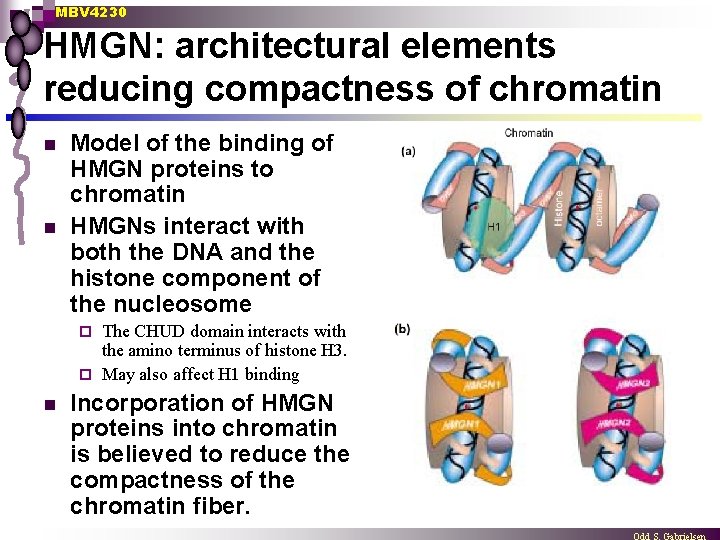
MBV 4230 HMGN: architectural elements reducing compactness of chromatin n n Model of the binding of HMGN proteins to chromatin HMGNs interact with both the DNA and the histone component of the nucleosome The CHUD domain interacts with the amino terminus of histone H 3. ¨ May also affect H 1 binding ¨ n Incorporation of HMGN proteins into chromatin is believed to reduce the compactness of the chromatin fiber.

MBV 4230 Alan P. Wolffe on HMG proteins n `There must be money in proteins that control your fat, your teeth, your sex and your health (besides minor things such as transcription, cell division and DNA recombination and repair). '
- Slides: 38