Arabidopsis Genome Annotation TAIR 7 Release Arabidopsis Genome

Arabidopsis Genome Annotation TAIR 7 Release

Arabidopsis Genome Annotation Overview of releases p Current release (TAIR 7) p Where to find TAIR 7 release data p Preview of next release (TAIR 8) p
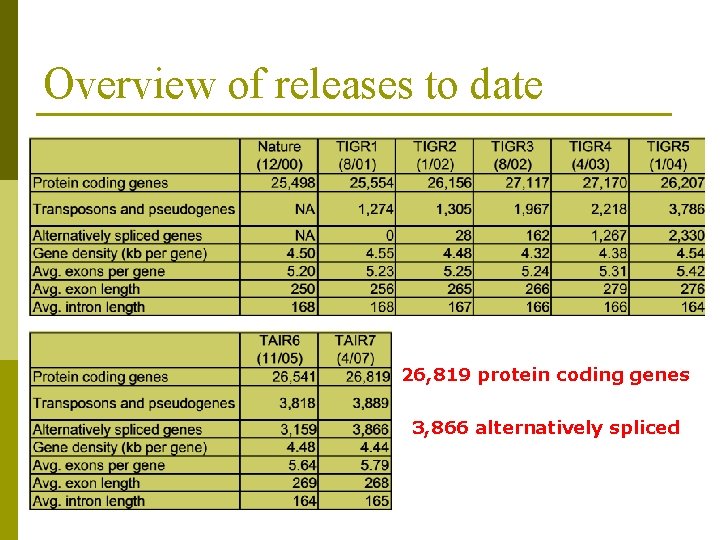
Overview of releases to date 26, 819 protein coding genes 3, 866 alternatively spliced

Average gene in TAIR 7 release 2221 bp long 146 bp Avg 5’ UTR 268 bp Avg Exon 165 bp Avg Intron 1. 16 splice variants per locus 233 bp Avg 3’ UTR

What was done for TAIR 7 p 681 new loci, 1774 new gene models p 211 Cysteine-rich peptides (CRPs) K. Silverstein, Univ. of Minnesota p 71 Micro. RNAs Matt Jones-Rhoades, MIT/mi. RBASE p 34 merges, 41 splits, 47 obsolete loci p 797 models with CDS updates p 10, 792 models with UTR updates p One third of all TAIR 6 loci (10, 098 loci) were updated for TAIR 7
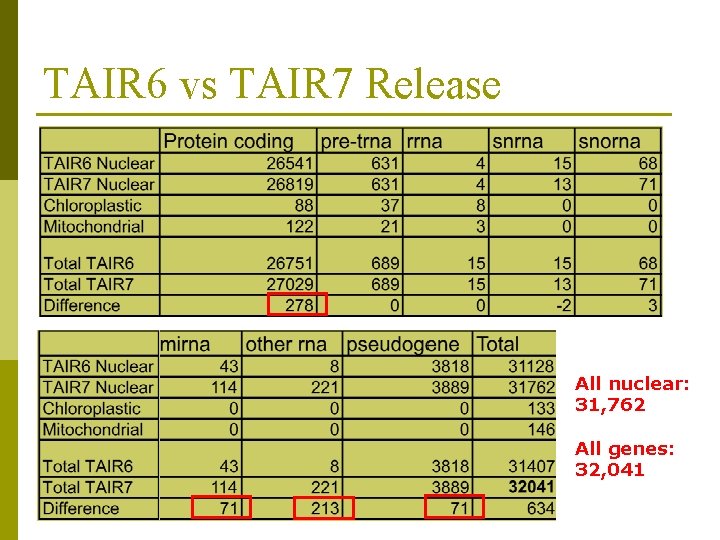
TAIR 6 vs TAIR 7 Release All nuclear: 31, 762 All genes: 32, 041

Annotation pipeline and strategy Gene updates n New Arabidopsis c. DNAs/ESTs incorporated via automated pipeline (PASA) p Result: 1717 non-UTR updates n Community updates (affecting 330 genes) n Manual curation to identify potential errors (targeted approach) n ~10% loci examined manually

Specific problems targeted p Small introns (65), long introns (89) p AT-AC splicing (55) p UTR errors (1098) p nc. RNAs and small proteins (251)

AT-AC splicing genes p 55 Gene models updated TAIR 6 Model AT-AC splice junction
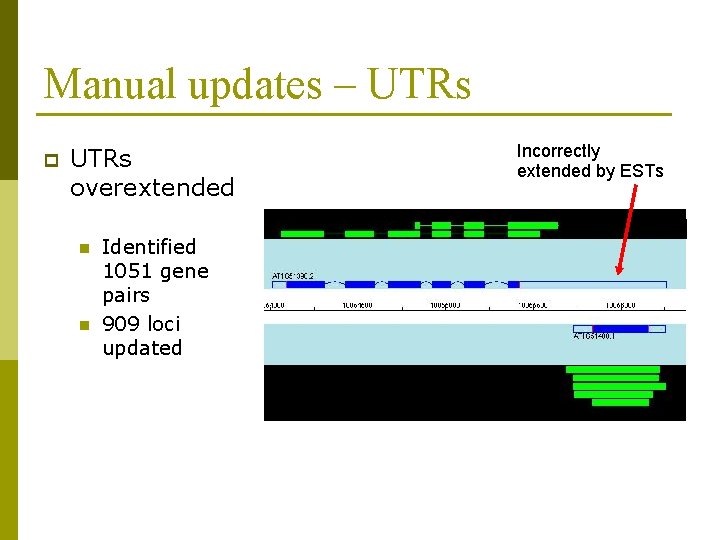
Manual updates – UTRs p UTRs overextended n n Identified 1051 gene pairs 909 loci updated Incorrectly extended by ESTs

nc. RNAs & small proteins p c. DNAa not represented in TAIR 6 gene set p p p 1260 c. DNAs do not map to TAIR 6 annotation (385 splice) 947 separate c. DNA clusters (“Loci”) (291 splice) 251 new loci added TAIR 7 1619 overlapping loci 1459 exon-exon overlaps 127 possible natural antisense genes nc. RNA

nc. RNAs & small proteins p c. DNAa not represented in TAIR 6 gene set p p p 1260 c. DNAs do not map to TAIR 6 annotation (385 splice) 947 separate c. DNA clusters (“Loci”) (291 splice) 251 new loci added TAIR 7 Small protein

Computational descriptions p Updated all computational descriptions n ANAC 001 (Arabidopsis NAC domain containing protein 1); transcription factor; similar to ANAC 069 (Arabidopsis NAC domain containing protein 69), transcription factor [Arabidopsis thaliana] (TAIR: AT 4 G 01550. 1); similar to putative NAC 2 protein [Oryza sativa (japonica cultivar-group)] (GB: BAD 09612. 1); contains Inter. Pro domain No apical meristem (NAM) protein; (Inter. Pro: IPR 003441). p ~4000 loci have similarity only to uncharacterised proteins (i. e. hypothetical, predicted, unknown etc). p 758 have no significant protein similarity to Genbank proteins p 286 also have no supporting EST/c. DNA evidence

TAIR 7 Summary p Chromosome sequence not changed p 681 new loci p 10, 098 loci updated p ~10% loci manually examined

Where to find TAIR 7 data p TAIR: n n p Genome Annotation Portal Bulk Download Tool (Sequences) Seq. Viewer (genome browser) FTP site NCBI n genomes section
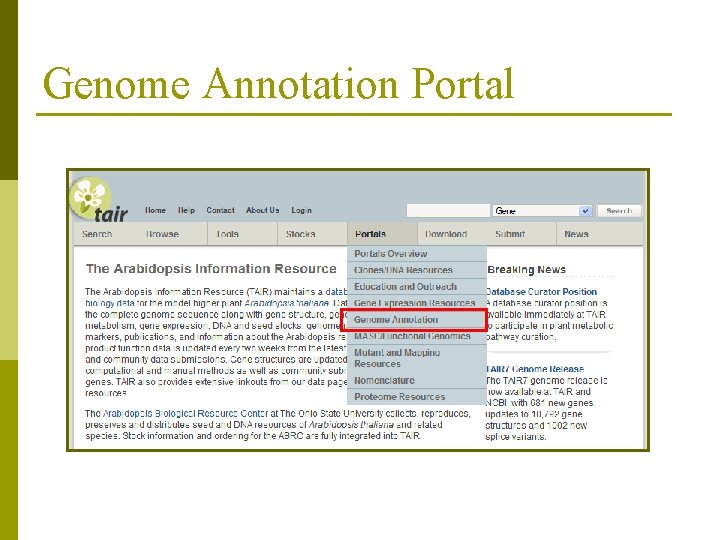
Genome Annotation Portal
Seq. Viewer (Genome Browser)
FTP download whole datasets

Preview of TAIR 8 release p Genome assembly updates p Annotation maintenance n n n Correct structural errors New transcript data Community submissions Missing genes and splice variants p Improved transposon annotation p

Missing genes and splice variants Continued identification of missing genes p Alternative splicing p 8, 264 alternative splicing events affecting 4, 707 genes, (Brendel V et. al. Proc Natl Acad Sci 2006) p 16, 252 events in 11665 models affecting 5, 313 genes, (Buell 2006 Genomics) p TAIR 7 alternative splicing giving 8844 models affecting 3866 genes p p Retained introns ~48% of alternatively spliced genes/loci
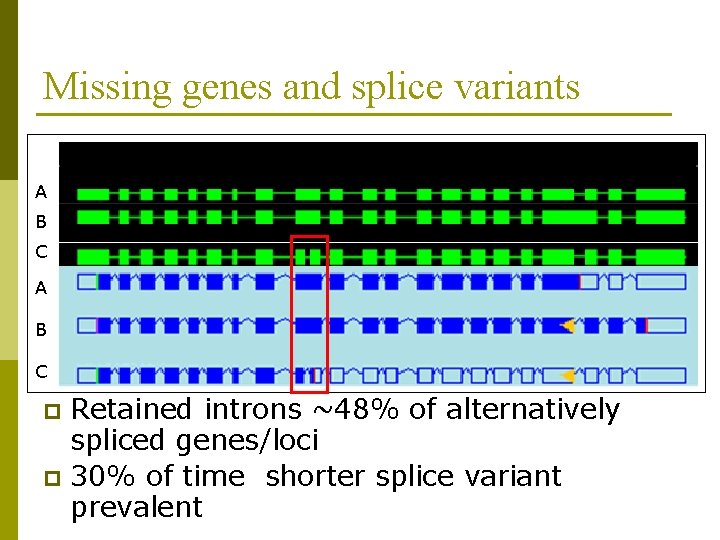
Missing genes and splice variants Continued identification of missing genes B p Alternative splicing Ap C A B C 8, 264 alternative splicing events affecting 4, 707 genes, (Brendel V et. al. Proc Natl Acad Sci 2006) p 16, 252 events in 11665 models affecting 5, 313 genes, (Buell 2006 Genomics) p TAIR 7 alternative splicing giving 8844 models affecting 3866 genes p Retained introns ~48% of alternatively spliced genes/loci p 30% of time shorter splice variant prevalent p

Transposons and pseudogenes 3889 “pseudogenes” p 2490 transposons 1399 pseudogenes p ~100 TEs not currently tagged as pseudo’s p Defined by a single pair of coordinates p At 3 g 26295
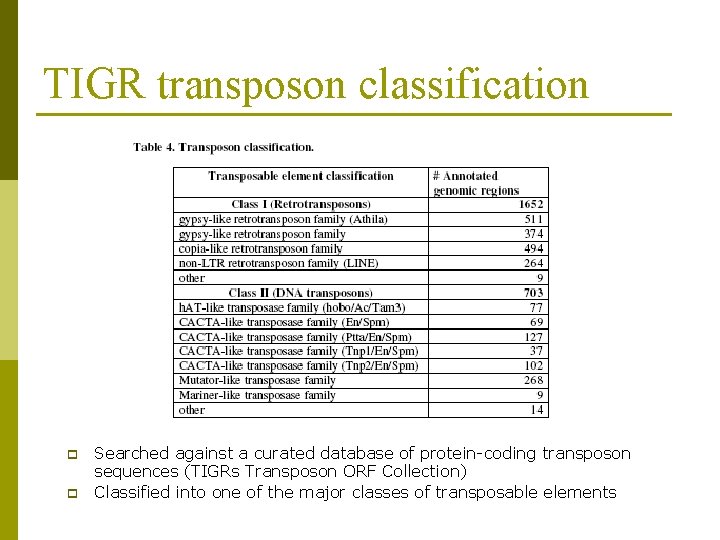
TIGR transposon classification p p Searched against a curated database of protein-coding transposon sequences (TIGRs Transposon ORF Collection) Classified into one of the major classes of transposable elements

Who cares about TEs? p p p Efficient markers in gene tagging and phylogenetic studies. Similarity with virus replication machinery and transcription factors Role in heterochromatin formation Involved in epigenetic gene regulation Genome annotators
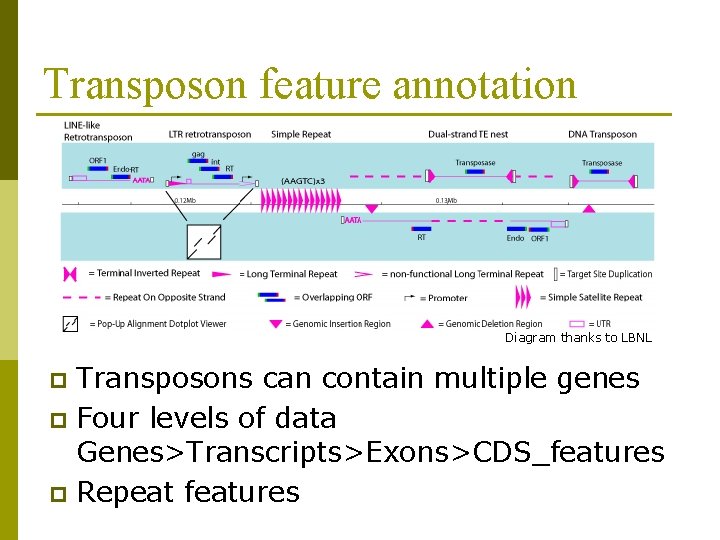
Transposon feature annotation Diagram thanks to LBNL Transposons can contain multiple genes p Four levels of data Genes>Transcripts>Exons>CDS_features p Repeat features p

Beyond TAIR 8 Mitochondrial and chloroplast gene reannotation p Comparative analysis using new genome sequences p Improved pseudogene annotation p Guide to supporting evidence for gene structure p
- Slides: 29