Ara Cyc Metabolic Pathway Annotation 1 Ara Cyc
Ara. Cyc Metabolic Pathway Annotation 1

Ara. Cyc – An overview § Ara. Cyc is a metabolic pathway database for Arabidopsis thaliana; § Computational prediction by Patho. Logic software using Meta. Cyc as the reference database (Peter Karp, SRI); § Predicted pathways were then manually validated; ongoing manual curation. 2
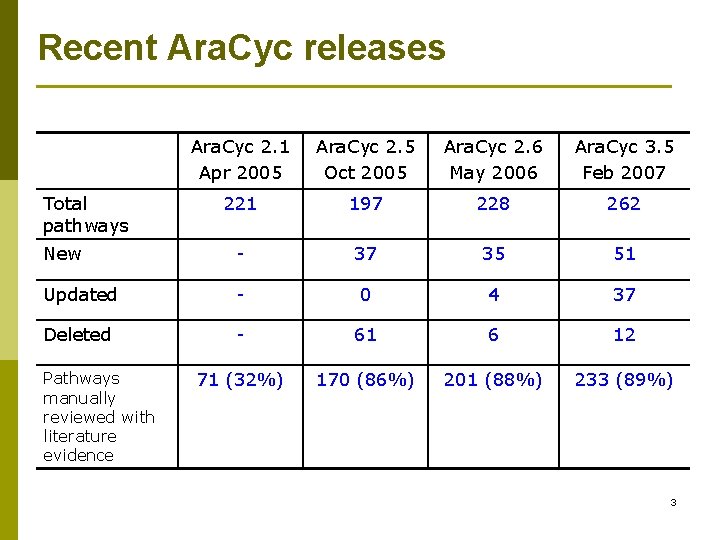
Recent Ara. Cyc releases Ara. Cyc 2. 1 Apr 2005 Ara. Cyc 2. 5 Oct 2005 Ara. Cyc 2. 6 May 2006 Ara. Cyc 3. 5 Feb 2007 221 197 228 262 New - 37 35 51 Updated - 0 4 37 Deleted - 61 6 12 71 (32%) 170 (86%) 201 (88%) 233 (89%) Total pathways Pathways manually reviewed with literature evidence 3

Upcoming Ara. Cyc 4. 0 § New pathways, updated pathways § Gene function annotation updated according to TAIR 7 genome release § Significant changes of the assignment of genes to reactions/pathways 4

Outline § Search and browse Ara. Cyc § Arabidopsis Metabolic map § Omics. Viewer 5
6
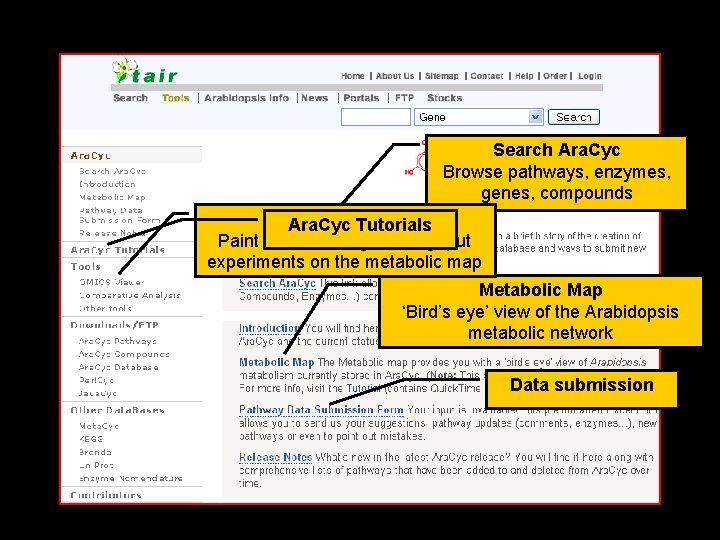
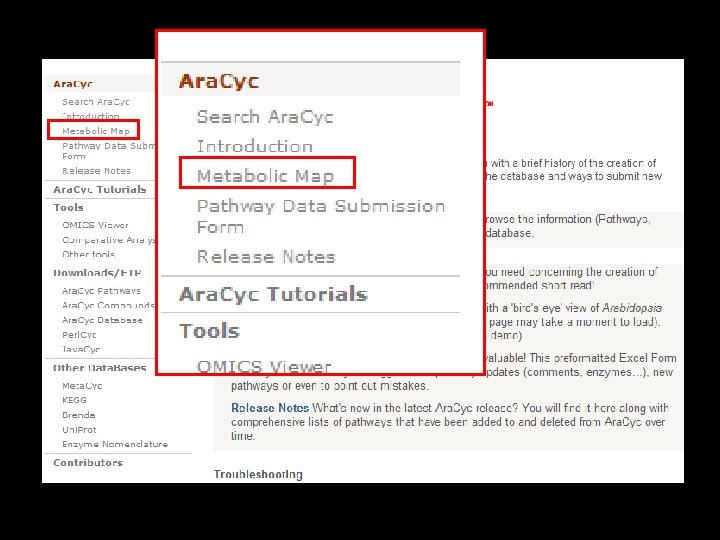
Search Ara. Cyc Browse pathways, enzymes, genes, compounds OMICS Ara. Cyc. Viewer Tutorials Paint data from high-throughput experiments on the metabolic map Metabolic Map ‘Bird’s eye’ view of the Arabidopsis metabolic network Data submission 7
8
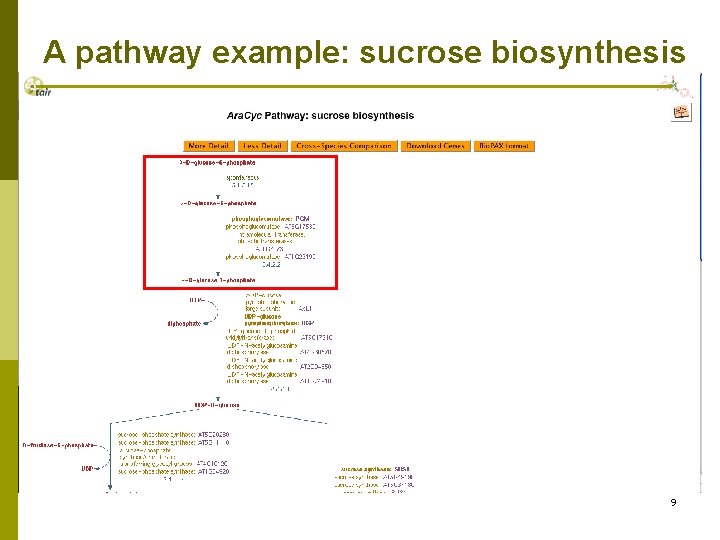
A pathway example: sucrose biosynthesis 9
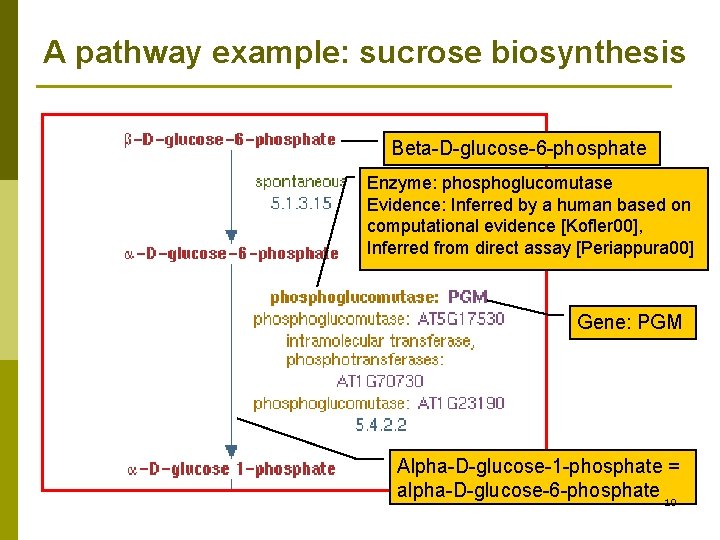
A pathway example: sucrose biosynthesis Beta-D-glucose-6 -phosphate Enzyme: phosphoglucomutase Evidence: Inferred by a human based on computational evidence [Kofler 00], Inferred from direct assay [Periappura 00] Gene: PGM Alpha-D-glucose-1 -phosphate = alpha-D-glucose-6 -phosphate 10
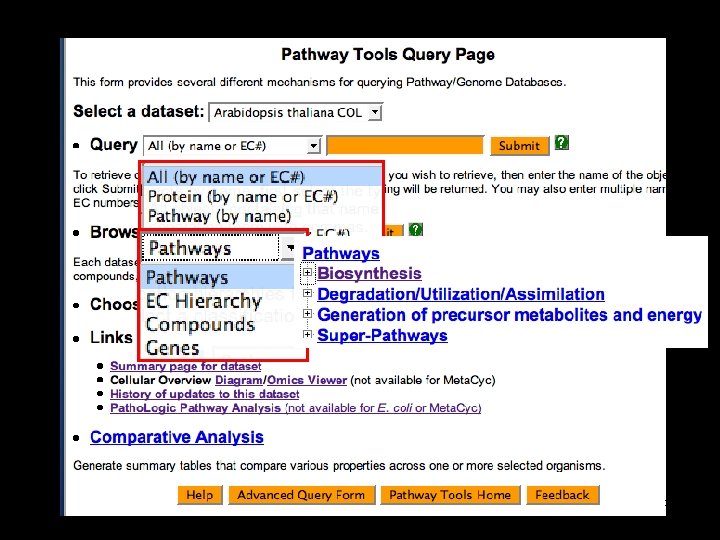
11
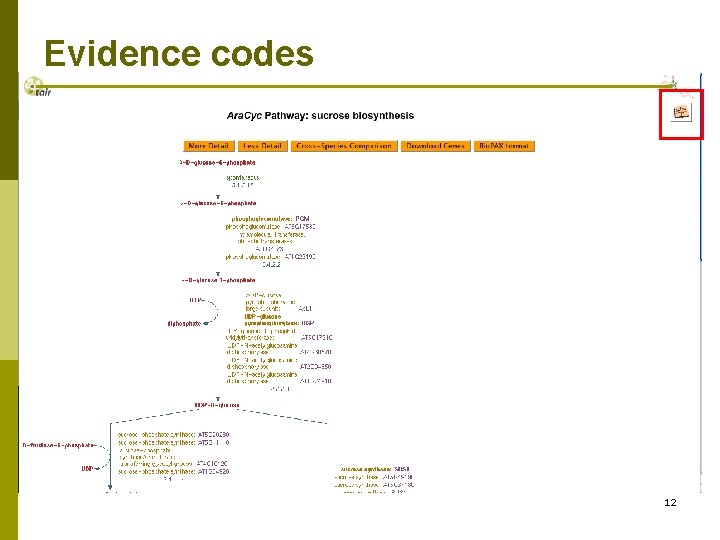
Evidence codes 12

Evidence codes Experimental evidence Computational evidence Evidence based on an Author Statement 13
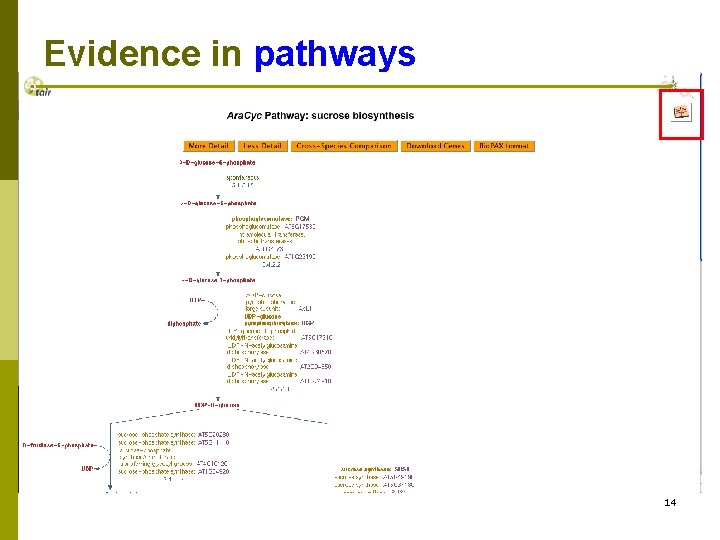
Evidence in pathways 14

Evidence for enzymatic activities 15
16
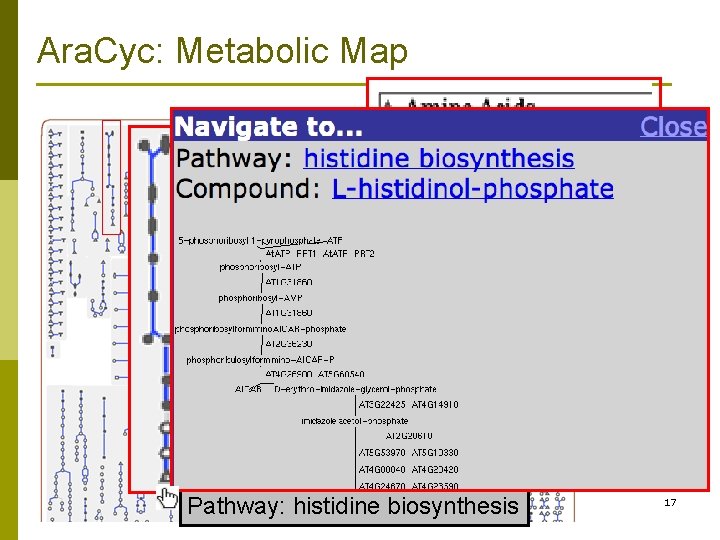
Ara. Cyc: Metabolic Map Compound: L-histidine Pathway: histidine biosynthesis 17
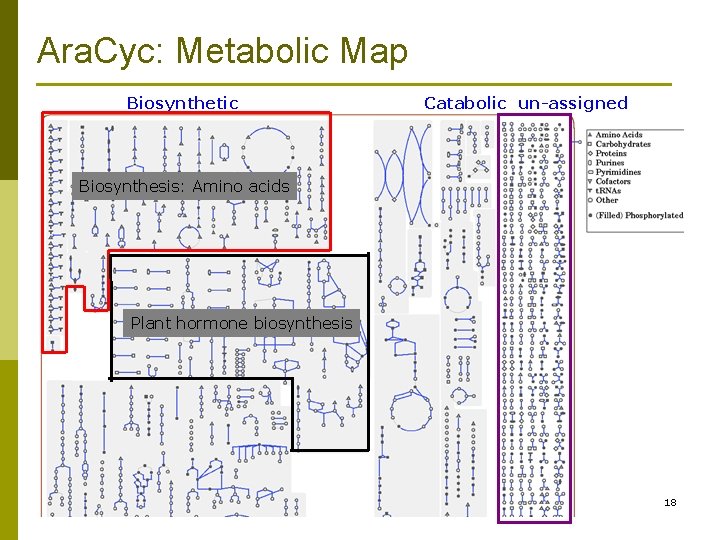
Ara. Cyc: Metabolic Map Biosynthetic Catabolic un-assigned Biosynthesis: Amino acids Plant hormone biosynthesis 18
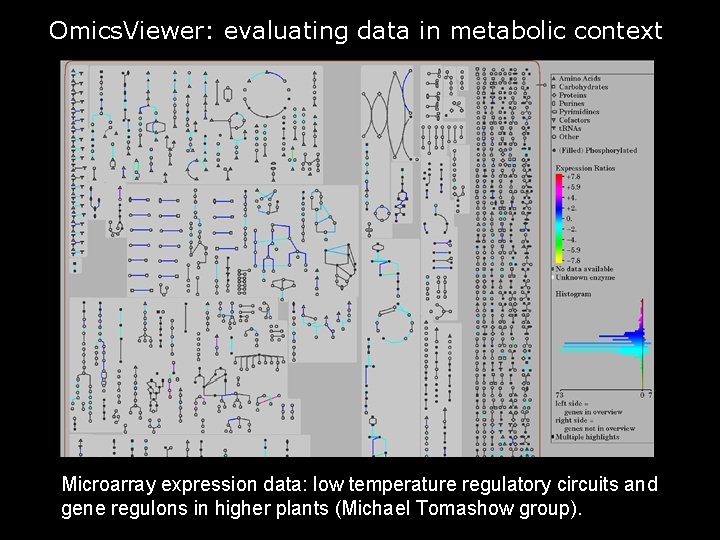
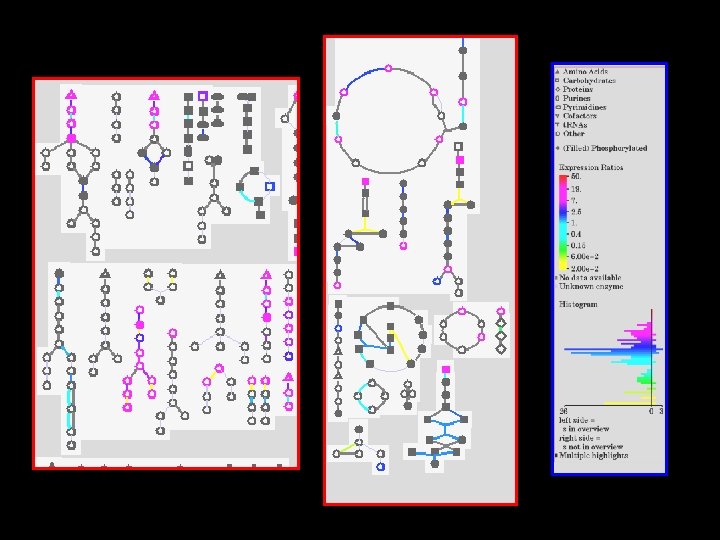
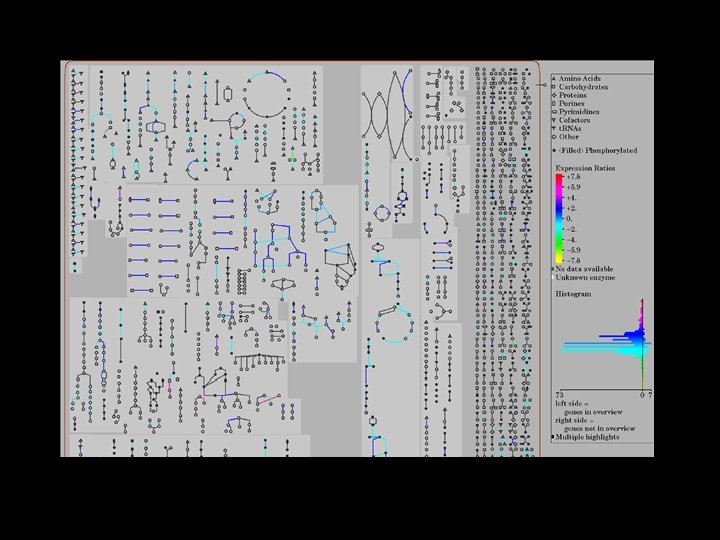
Omics. Viewer: evaluating data in metabolic context Microarray expression data: low temperature regulatory circuits and gene regulons in higher plants (Michael Tomashow group). 19
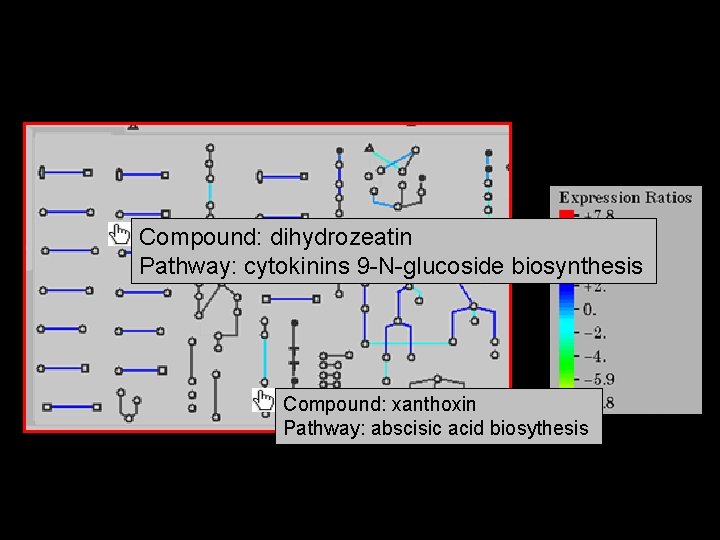
Compound: dihydrozeatin Pathway: cytokinins 9 -N-glucoside biosynthesis Compound: xanthoxin Pathway: abscisic acid biosythesis 20
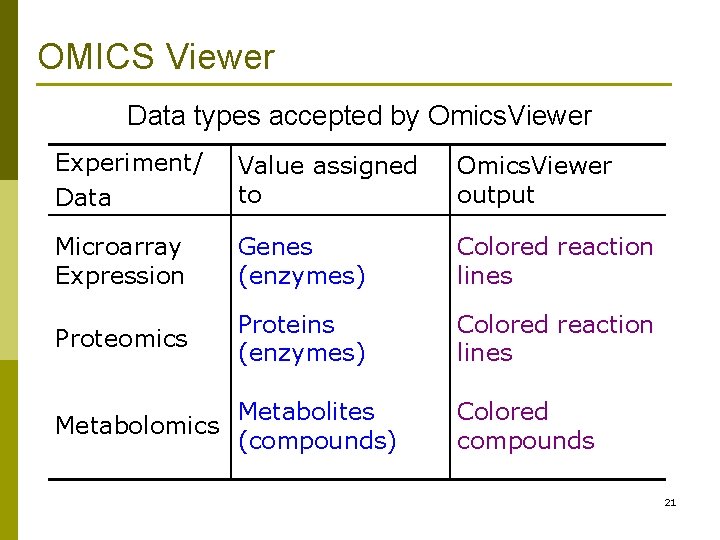
OMICS Viewer Data types accepted by Omics. Viewer Experiment/ Data Value assigned to Omics. Viewer output Microarray Expression Genes (enzymes) Colored reaction lines Proteomics Proteins (enzymes) Colored reaction lines Metabolites Metabolomics (compounds) Colored compounds 21
22

Access Omics Viewer 23
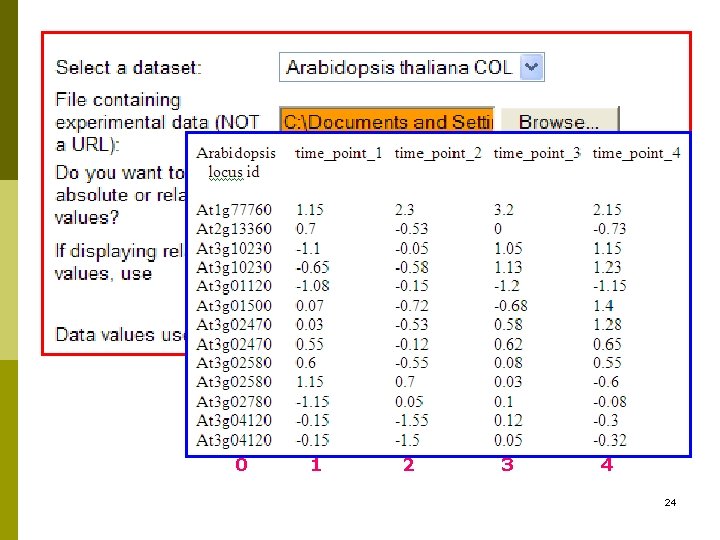
0 1 2 3 4 24
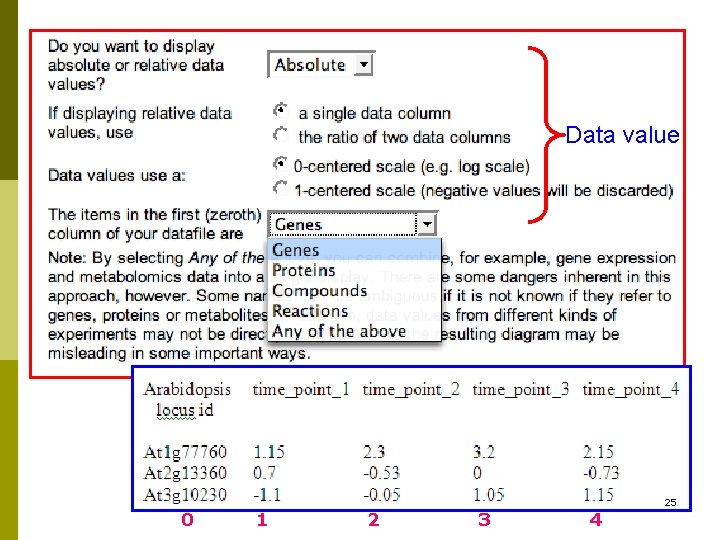
Data value 25 0 1 2 3 4

26

0 1 2 3 4 Result A: single page for a single time point 27
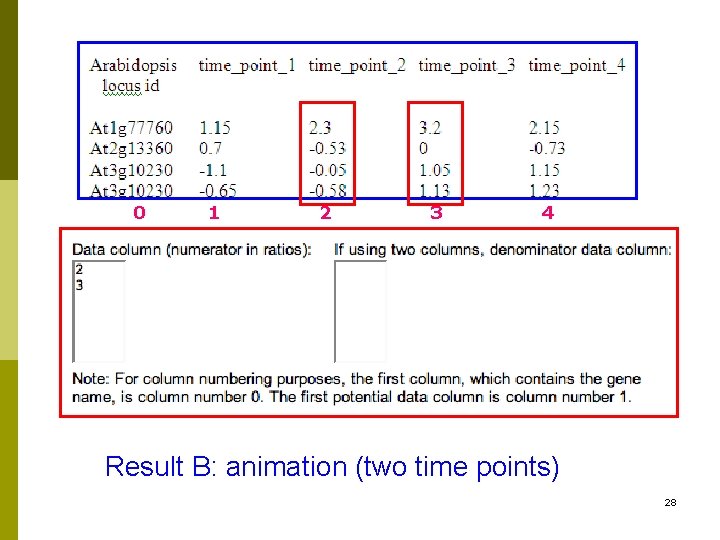
0 1 2 3 4 Result B: animation (two time points) 28

0 1 2 3 4 Result C: single page (ratio of two time points: 2 /3) 29

0 1 2 3 4 Result D: animation (three pages: : 2 /1, 3 /1, 4 /2) 30

31
32
- Slides: 32