Applying Neural Networks to Predict AntibioticResistant Bacterial Infection

Applying Neural Networks to Predict Antibiotic-Resistant Bacterial Infection from Shotgun Metagenomic Data Innovative Statistics and Machine Learning for Precision Medicine September 16 th, 2017 David Haslam, M. D. Associate Professor Director Antimicrobial Stewardship Program Division of Infectious Diseases
Deaths due to infectious disease are on the rise • largely due to antibiotic-resistant organisms
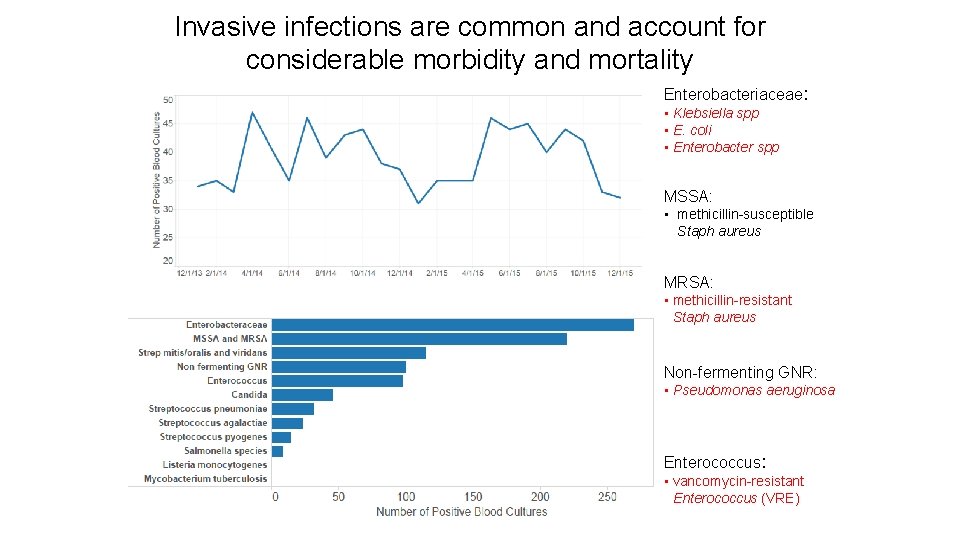
Invasive infections are common and account for considerable morbidity and mortality Enterobacteriaceae: • Klebsiella spp • E. coli • Enterobacter spp MSSA: • methicillin-susceptible Staph aureus MRSA: • methicillin-resistant Staph aureus Non-fermenting GNR: • Pseudomonas aeruginosa Enterococcus: • vancomycin-resistant Enterococcus (VRE)

Where do the infections come from? Can we predict which antibiotic will be most effective before the child becomes ill? Can we prevent these serious infections?

Where do the infections come from? Can we predict which antibiotic will be most effective before the child becomes ill? Can we prevent these serious infections?
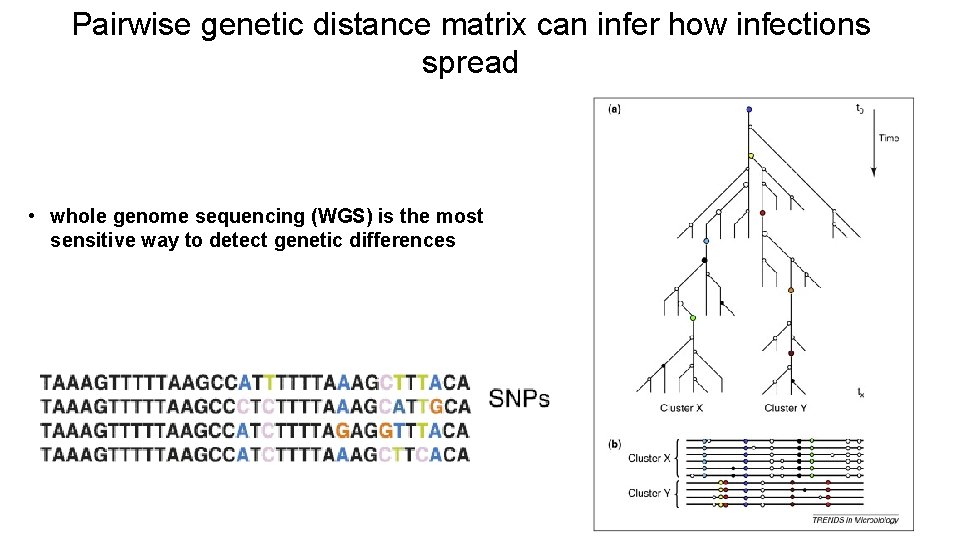
Pairwise genetic distance matrix can infer how infections spread • whole genome sequencing (WGS) is the most sensitive way to detect genetic differences
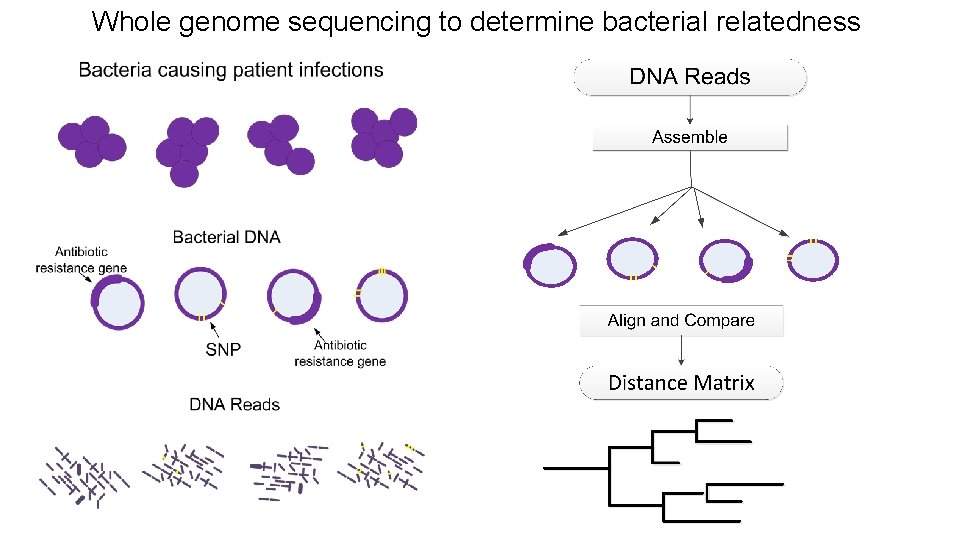
Whole genome sequencing to determine bacterial relatedness
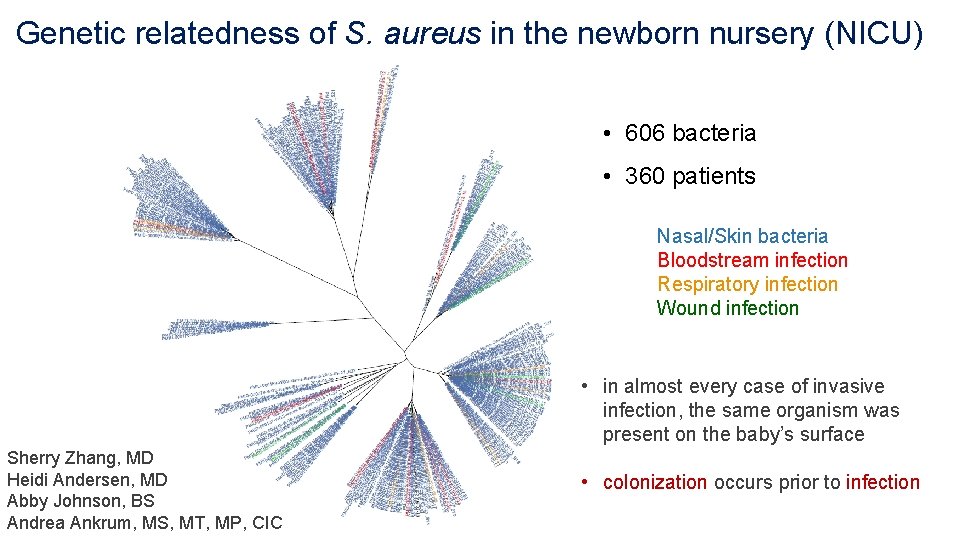
Genetic relatedness of S. aureus in the newborn nursery (NICU) • 606 bacteria • 360 patients Nasal/Skin bacteria Bloodstream infection Respiratory infection Wound infection • in almost every case of invasive infection, the same organism was present on the baby’s surface Sherry Zhang, MD Heidi Andersen, MD Abby Johnson, BS Andrea Ankrum, MS, MT, MP, CIC • colonization occurs prior to infection
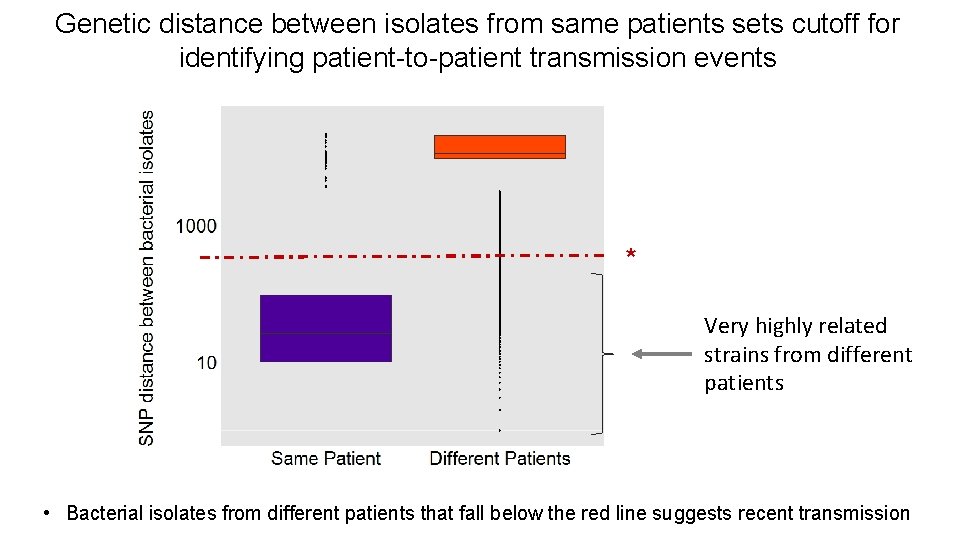
Genetic distance between isolates from same patients sets cutoff for identifying patient-to-patient transmission events * Very highly related strains from different patients • Bacterial isolates from different patients that fall below the red line suggests recent transmission
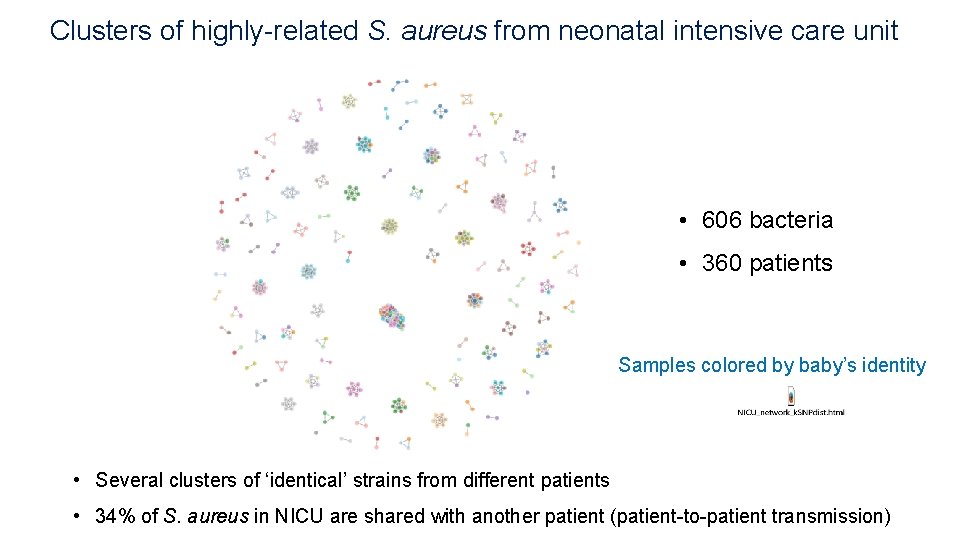
Clusters of highly-related S. aureus from neonatal intensive care unit • 606 bacteria • 360 patients Samples colored by baby’s identity • Several clusters of ‘identical’ strains from different patients • 34% of S. aureus in NICU are shared with another patient (patient-to-patient transmission)

Where do these infections come from? • in many cases the infection comes from another patient (poor handwashing? ) • in almost all cases, the bacteria was on the patient’s body before the infection developed Can we predict which antibiotic will be most effective before the child becomes ill? Can we prevent these infections?

Where do the infections come from? • in many cases the infection comes from another patient (poor handwashing? ) • in almost all cases, the bacteria was on the patient’s body before the infection developed Can we predict which antibiotic will be most effective before the child becomes ill? Can we prevent these infections?
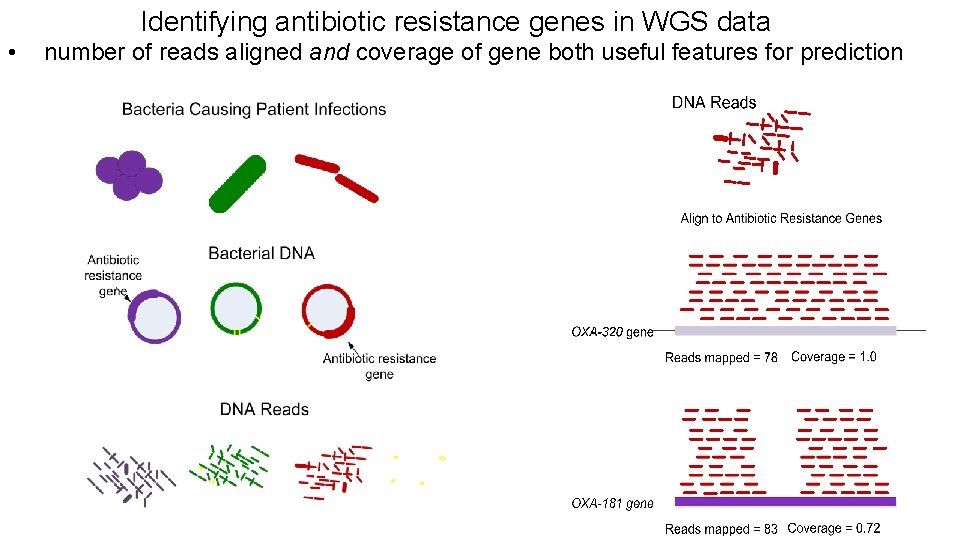
Identifying antibiotic resistance genes in WGS data • number of reads aligned and coverage of gene both useful features for prediction
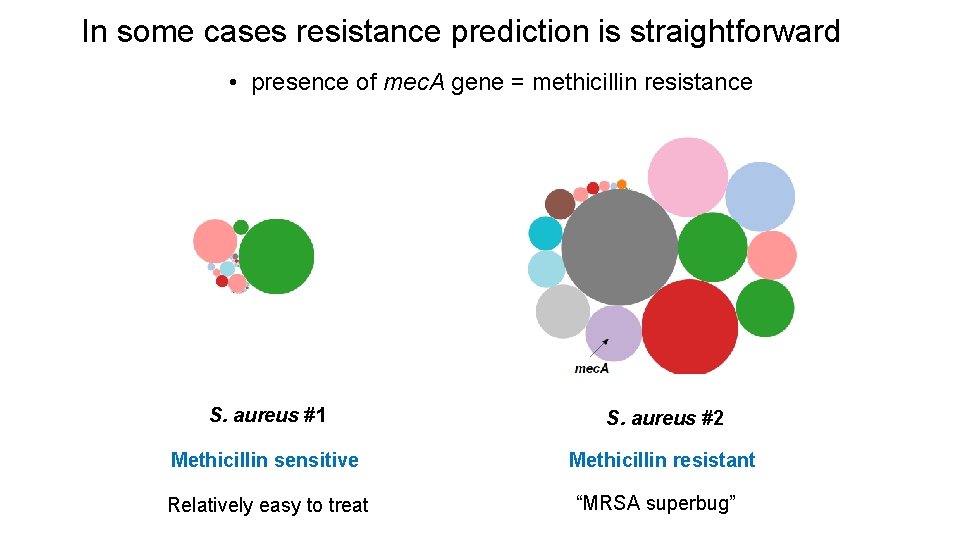
In some cases resistance prediction is straightforward • presence of mec. A gene = methicillin resistance S. aureus #1 S. aureus #2 Methicillin sensitive Methicillin resistant Relatively easy to treat “MRSA superbug”
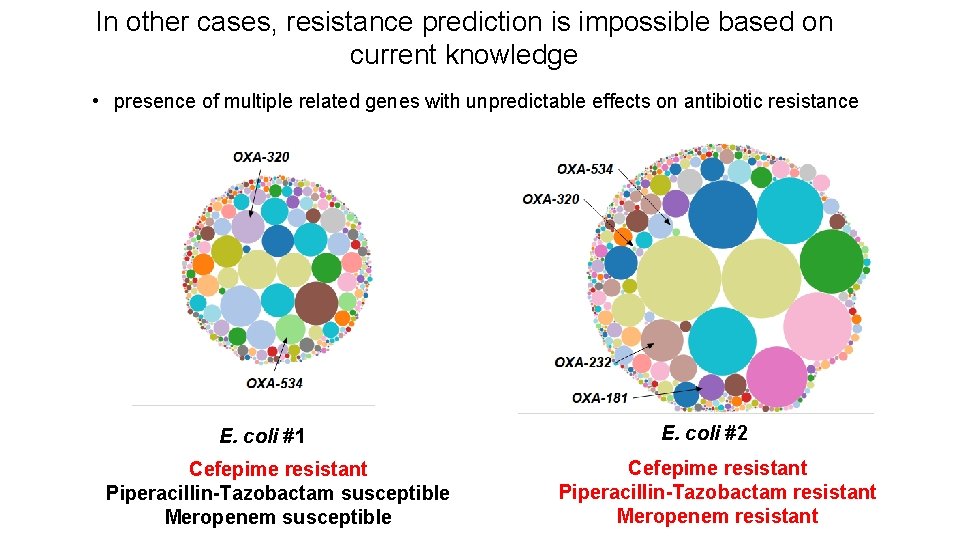
In other cases, resistance prediction is impossible based on current knowledge • presence of multiple related genes with unpredictable effects on antibiotic resistance E. coli #1 Cefepime resistant Piperacillin-Tazobactam susceptible Meropenem susceptible E. coli #2 Cefepime resistant Piperacillin-Tazobactam resistant Meropenem resistant
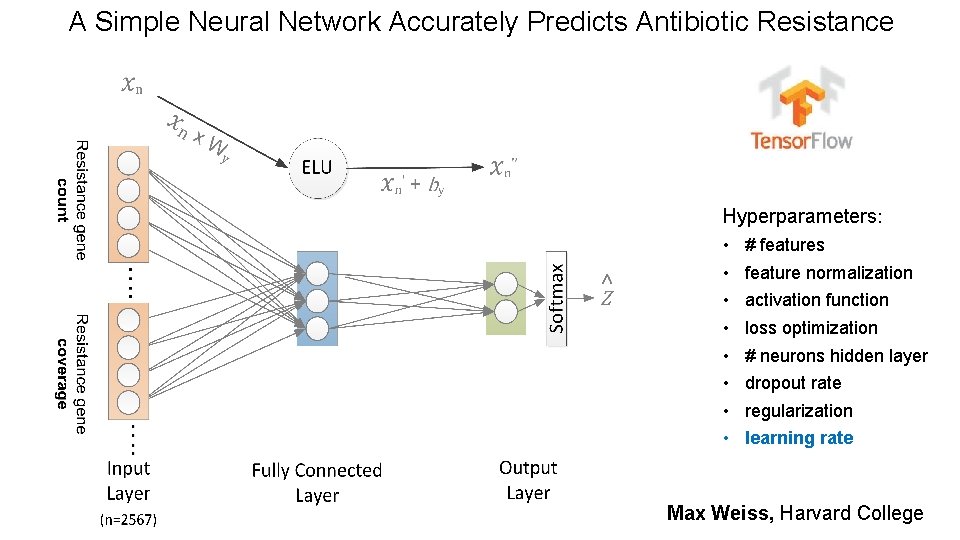
A Simple Neural Network Accurately Predicts Antibiotic Resistance Hyperparameters: • • # features feature normalization activation function loss optimization # neurons hidden layer dropout rate regularization learning rate Max Weiss, Harvard College
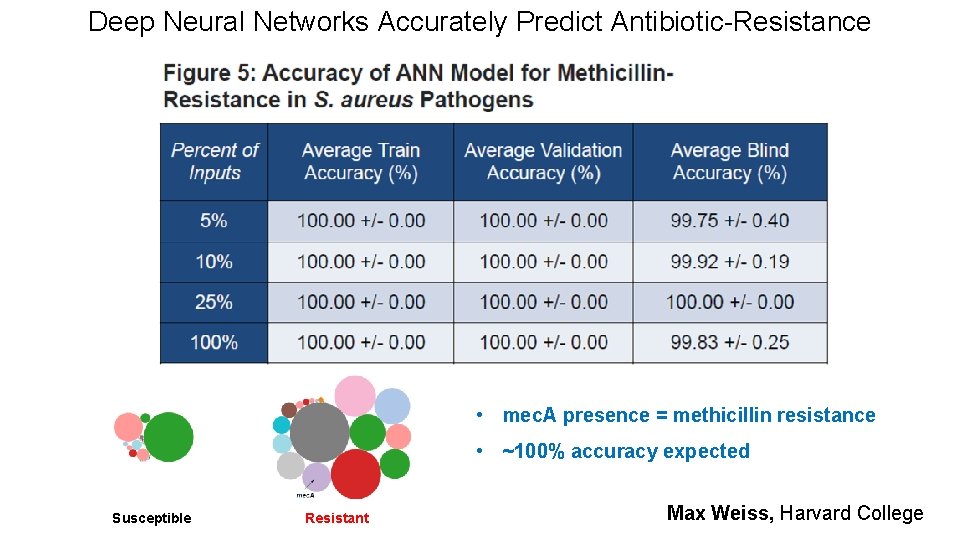
Deep Neural Networks Accurately Predict Antibiotic-Resistance • mec. A presence = methicillin resistance • ~100% accuracy expected Susceptible Resistant Max Weiss, Harvard College
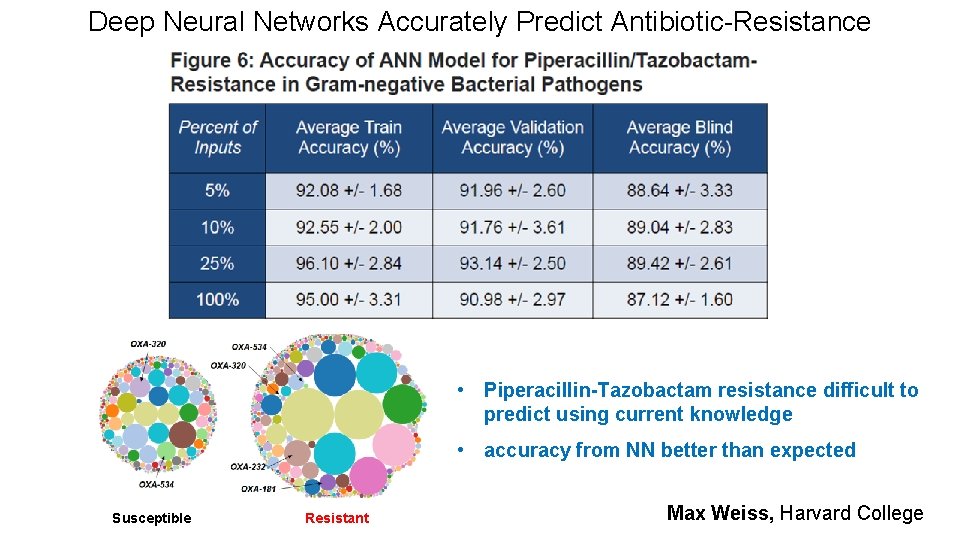
Deep Neural Networks Accurately Predict Antibiotic-Resistance • Piperacillin-Tazobactam resistance difficult to predict using current knowledge • accuracy from NN better than expected Susceptible Resistant Max Weiss, Harvard College
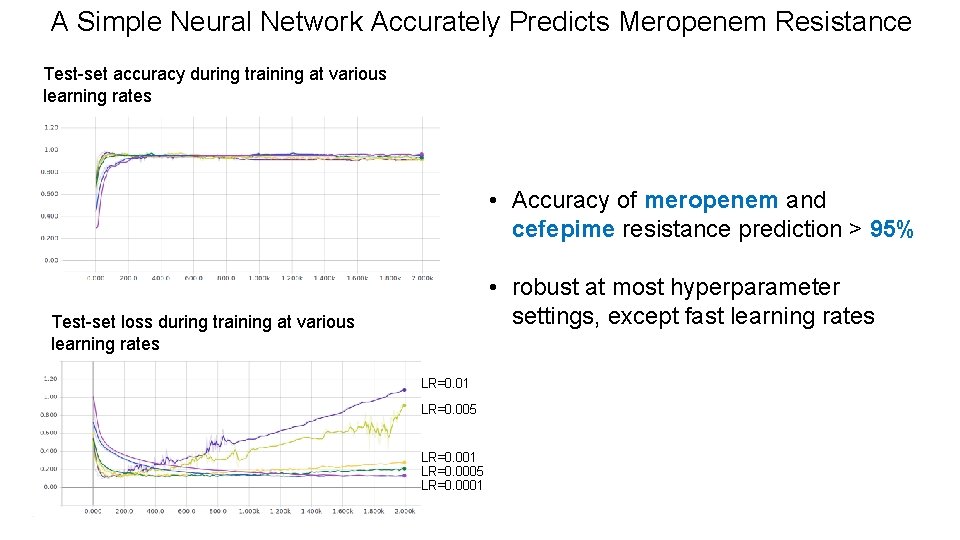
A Simple Neural Network Accurately Predicts Meropenem Resistance Test-set accuracy during training at various learning rates • Accuracy of meropenem and cefepime resistance prediction > 95% • robust at most hyperparameter settings, except fast learning rates Test-set loss during training at various learning rates LR=0. 01 LR=0. 005 LR=0. 001 LR=0. 0005 LR=0. 0001

Summary 1: Whole genome sequencing (WGS) • Reveals patient-to-patient transmission is common • Reveals most children “carry” a bacterium on their body before they become ill • Neural networks accurately predict antibiotic resistance • However, requires growth in laboratory and only a single bacterium can be analyzed at a time (slow and expensive = impractical)

Metagenomic Shotgun Sequencing (MSS) as a screening diagnostic test : • Analyze all possible bacteria in one sample • No requirement to grow bacteria in culture • “Dirty” data : requires deconvolution • No existing analytic tools Heidi Andersen, MD Staff Physician, Division of Infectious Diseases
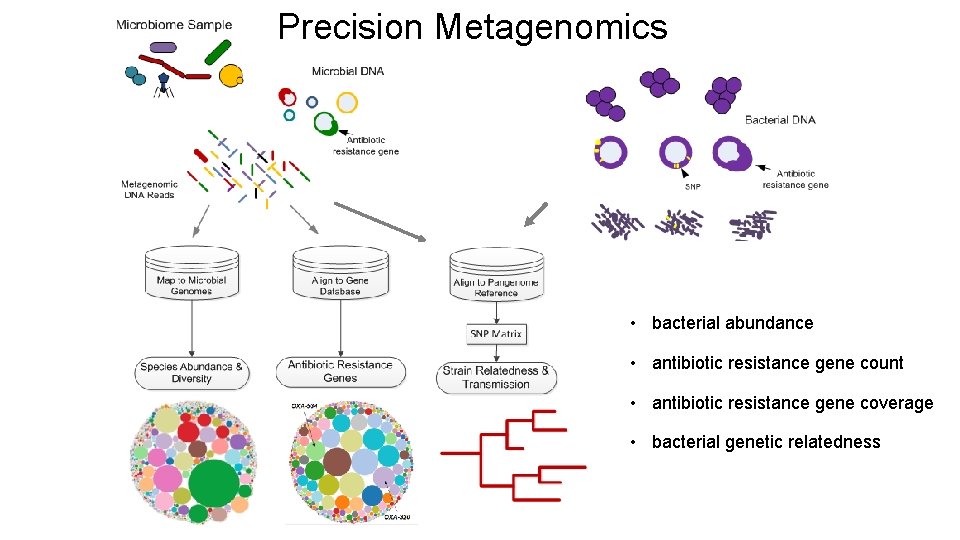
Precision Metagenomics • bacterial abundance • antibiotic resistance gene count • antibiotic resistance gene coverage • bacterial genetic relatedness
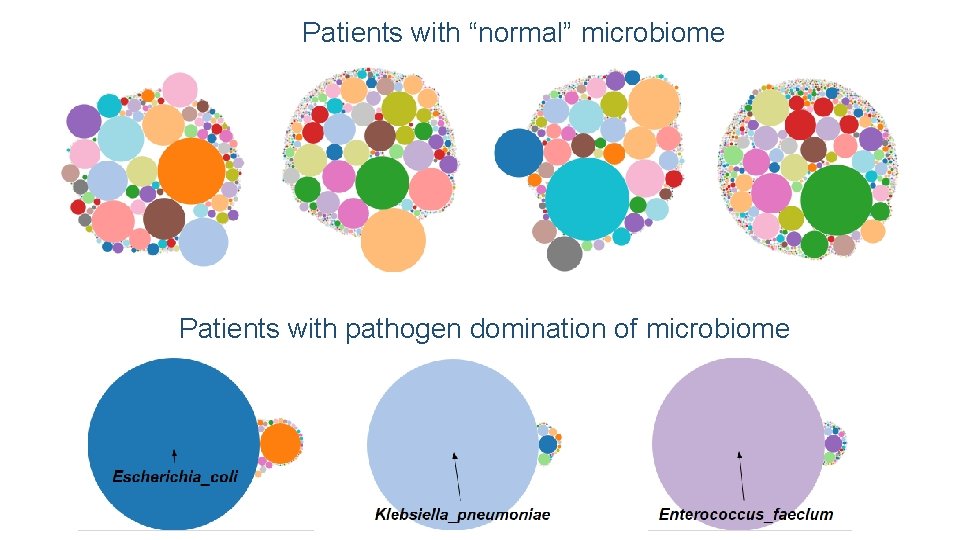
Patients with “normal” microbiome Patients with pathogen domination of microbiome
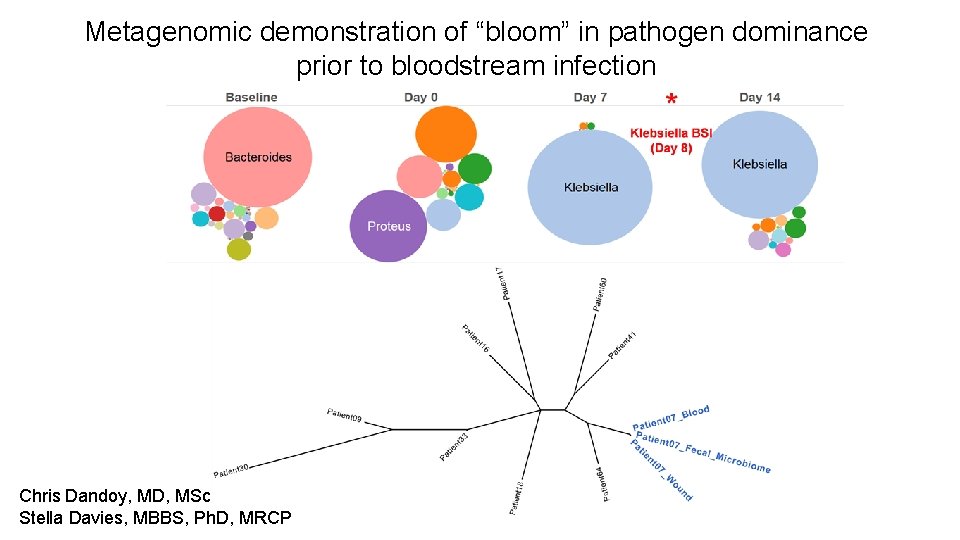
Metagenomic demonstration of “bloom” in pathogen dominance prior to bloodstream infection Chris Dandoy, MD, MSc Stella Davies, MBBS, Ph. D, MRCP

Where do the infections come from? Before the child develops infection can we predict which antibiotic will be most effective? Can we prevent these serious infections?
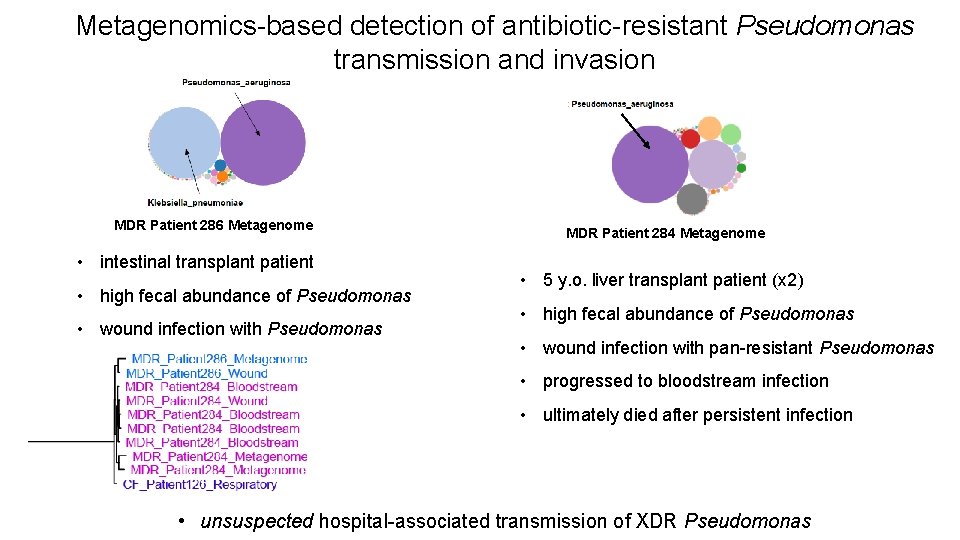
Metagenomics-based detection of antibiotic-resistant Pseudomonas transmission and invasion MDR Patient 286 Metagenome • intestinal transplant patient • high fecal abundance of Pseudomonas • wound infection with Pseudomonas MDR Patient 284 Metagenome • 5 y. o. liver transplant patient (x 2) • high fecal abundance of Pseudomonas • wound infection with pan-resistant Pseudomonas • progressed to bloodstream infection • ultimately died after persistent infection • unsuspected hospital-associated transmission of XDR Pseudomonas
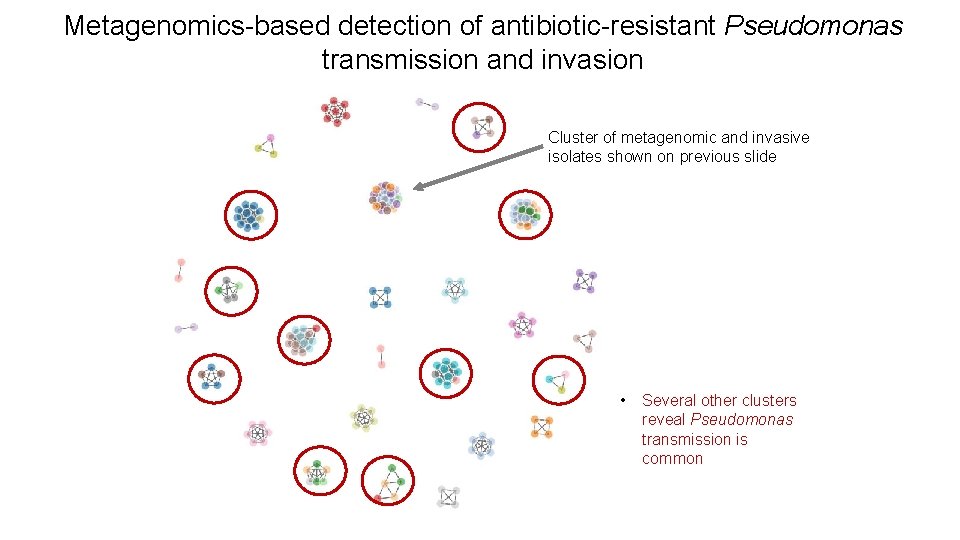
Metagenomics-based detection of antibiotic-resistant Pseudomonas transmission and invasion Cluster of metagenomic and invasive isolates shown on previous slide • Several other clusters reveal Pseudomonas transmission is common

At our hospital we have identified transmission and invasion of: • E. coli • Klebsiella pneumoniae • Klebsiella oxytoca • Enterobacter cloacae • Pseudomonas aeruginosa • Enterococcus faecalis • Enterococcus faecium • transmission events would not have been identified without metagenomic sequencing

Where do these infections come from? • in many cases the infection comes from another patient (poor handwashing? ) • in almost all cases, the bacteria was on the patient’s body before the infection developed Before the child develops infection can we predict which antibiotic will be most effective? Can we prevent these infections?

Where do these infections come from? • in many cases the infection comes from another patient (poor handwashing? ) • in almost all cases, the bacteria was on the patient’s body before the infection developed Can we predict which antibiotic will be most effective before the child develops infection? Can we prevent these infections?
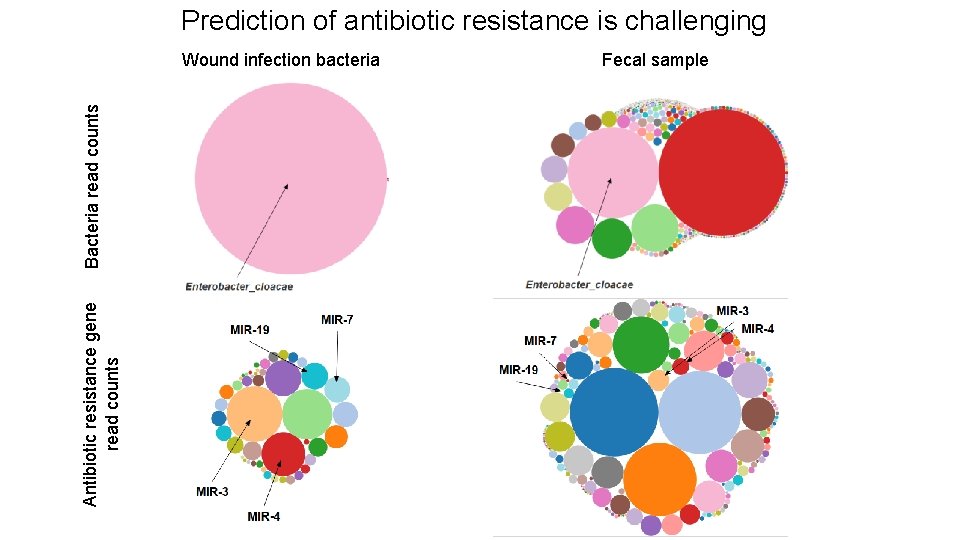
Prediction of antibiotic resistance is challenging Antibiotic resistance gene read counts Bacteria read counts Wound infection bacteria Fecal sample
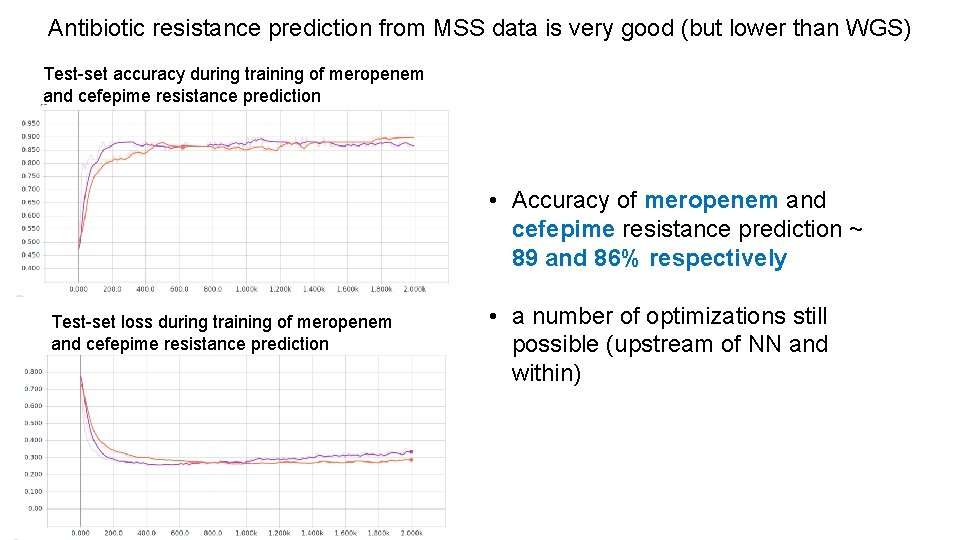
Antibiotic resistance prediction from MSS data is very good (but lower than WGS) Test-set accuracy during training of meropenem and cefepime resistance prediction • Accuracy of meropenem and cefepime resistance prediction ~ 89 and 86% respectively Test-set loss during training of meropenem and cefepime resistance prediction • a number of optimizations still possible (upstream of NN and within)

Summary 2: Metagenomic Shotgun Sequencing • As with WGS of isolated bacterium, metagenomic sequencing: • Reveals patient-to-patient transmission is common • Reveals most children “carry” a bacterium in or on their body before they become ill • Neural networks accurately predict antibiotic resistance from crude fecal samples • Samples all bacteria simultaneously • No need to incubate the samples and grow the bacterium • Relatively fast and inexpensive – suitable for diagnostic test

Where do these infections come from? • in many cases the infection comes from another patient (poor handwashing? ) • in almost all cases, the bacteria was on the patient’s body before the infection developed Can we predict which antibiotic will be most effective before the child develops infection? Can we prevent these infections?
Fecal transplantation (FMT) is highly effective for treatment of C. difficile infection • 90 to 95 % cure rate • better than any other treatment Mir, S. , et al. (2014). Current Pediatr Rep 2: 227 -234.

Fecal microbial transplantation (FMT) displaces antibiotic resistant pathogens Before FMT After FMT

Metagenomic sequencing has the capacity to prevent infection and improve outcomes by: 1. Isolating colonized patients to prevent transmission 2. Preventing inappropriate antibiotic use 3. Choosing correct antibiotic for patient who develops infection after colonization with antibiotic-resistant bacteria 4. Displacing pathogenic organism by probiotics or fecal transplantation

Heidi Andersen, MD T 32 NIH Grant Staff Physician, Division of Infectious Diseases MECEH Fellowship Program at UC Center for Pediatric Genomics at CCHMC Max Weiss Harvard College, SURF Summer Program Xinhua Zhang, MD Conrad Cole, MD, MPH Neonatology Shanxi Provincial Children’s Hospital Jaimie Nathan, MD Abigail Johnson Stella Davies, MBBS, Ph. D, MRCP MA Candidate, UC Alexey Porollo, Ph. D Chris Dandoy, MD, MSc Hansraj Bangar, Ph. D Gurjit Neeru Hershey, MD, Ph. D Research Associate, Infectious Diseases Tammy Hunter, MSTP Student Andrea Ankrum, MS, MT, MP, CIC Ted Densen, MD & Jennifer Hellman, MD Preventionist, Infection Control

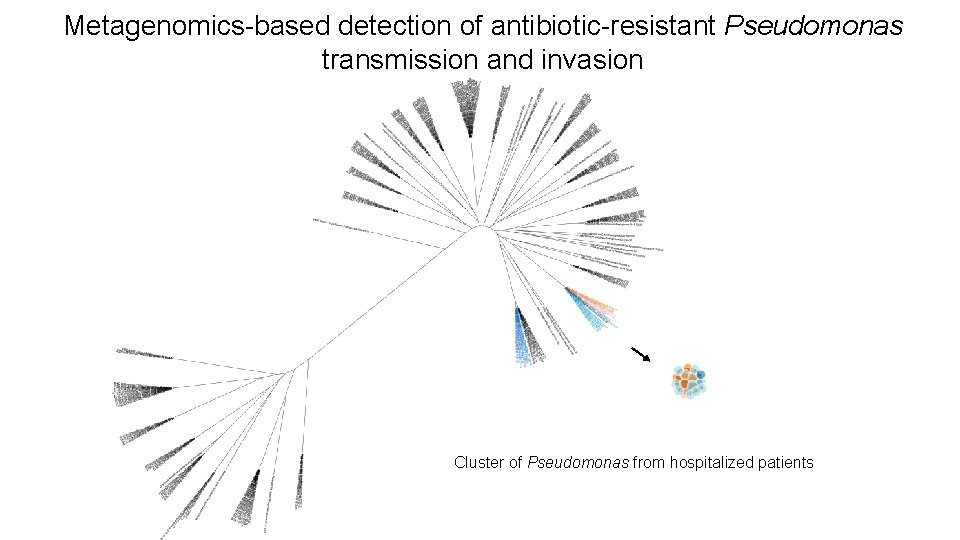
Metagenomics-based detection of antibiotic-resistant Pseudomonas transmission and invasion Cluster of Pseudomonas from hospitalized patients
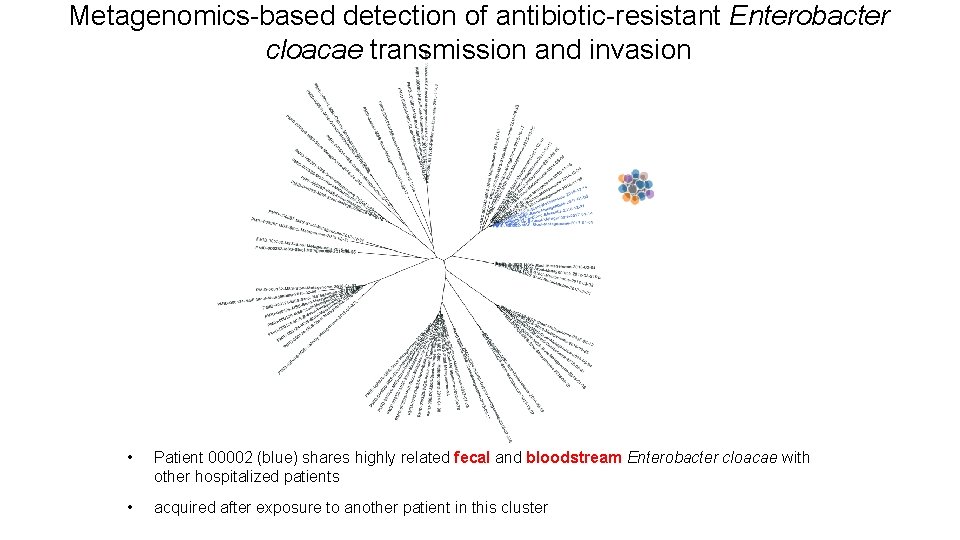
Metagenomics-based detection of antibiotic-resistant Enterobacter cloacae transmission and invasion • Patient 00002 (blue) shares highly related fecal and bloodstream Enterobacter cloacae with other hospitalized patients • acquired after exposure to another patient in this cluster

Metagenomics-based detection of vancomycin-resistant Enterococcus faecium transmission and invasion
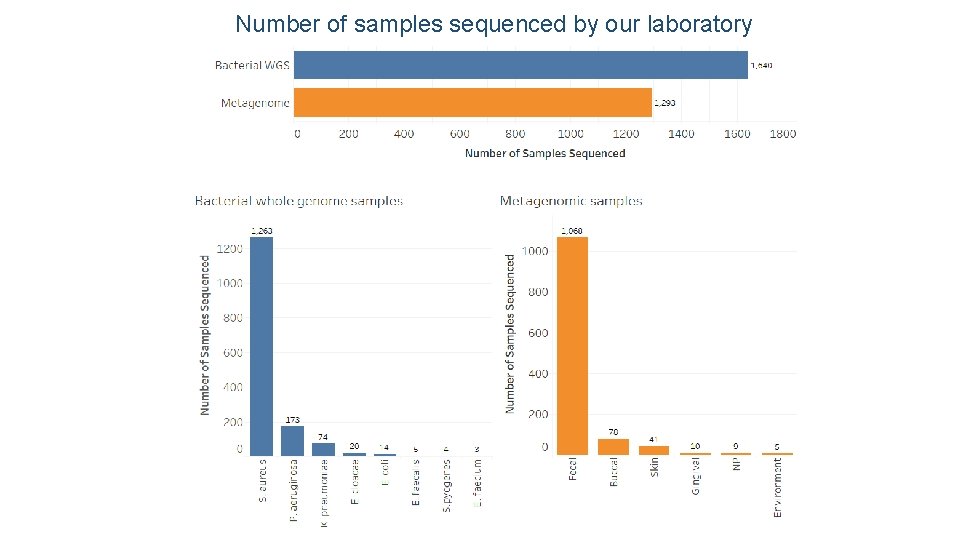
Number of samples sequenced by our laboratory
Pathogen Abundance Associated with Increased Risk of Invasive Infection BSI Pathogen BSI Events (No. ) Relative Abundance Threshold Adjusted Hazard Ratio P value Confidence Interval Enterococcus 21 >12% 3. 30 0. 004 1. 47 to 7. 41 Klebsiella 16 >4% 3. 19 0. 02 1. 20 to 8. 52 Pseudomonas 8 >1% 8. 34 <0. 001 3. 57 to 19. 48 Fungi 10 >2% 5. 62 0. 001 2. 00 to 15. 79 • patients with ‘normal’ microbiome did not develop invasive infection
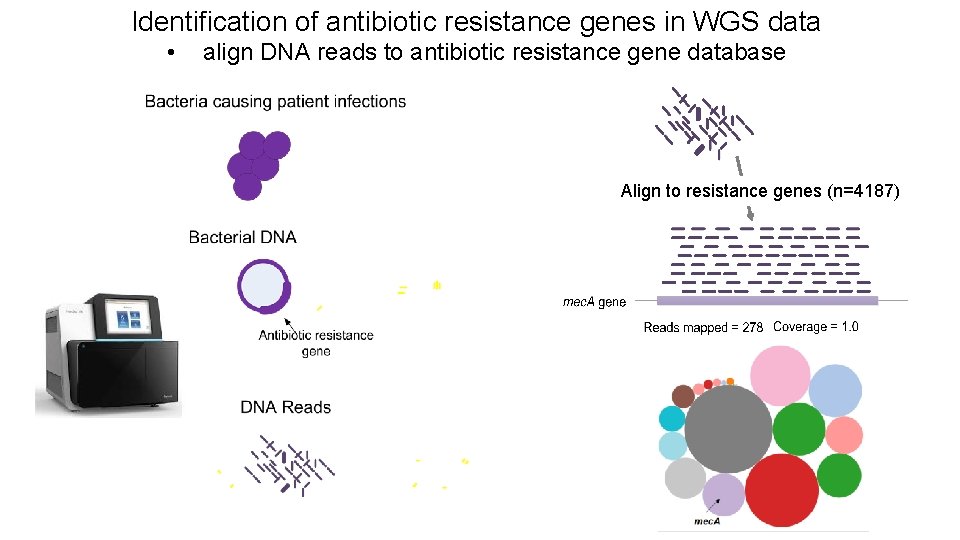
Identification of antibiotic resistance genes in WGS data • align DNA reads to antibiotic resistance gene database Align to resistance genes (n=4187)
- Slides: 45