Applied Multivariate Quantitative Methods Discriminant Analysis By Jenpei

Applied Multivariate Quantitative Methods Discriminant Analysis By Jen-pei Liu, Ph. D Division of Biometry, Department of Agronomy, National Taiwan University and Wei-Chie, MD, Ph. D Department of Public Health National Taiwan University 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 1

Introduction n n Introduction Methods for Two Populations n n Methods for Several Populations n n n Fisher Linear Discriminant Function Optimal Classification Rules Estimation of Misclassification Rates Fisher Linear Discriminant Function Minimum Distances Nearest Neighbor Method 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 2

Introduction n Examples n n Identifying variables that significantly differentiate between audited tax returns that resulted in underpayment of taxes and those that did not Identifying genes that have differentially expressed between the groups of breast patients that responds differently to treatment of tamoxifen 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 3

Introduction n Tamoxifen for breast cancer n n A competitive inhibitor of estrogen binding to estrogen receptor (ER) Reduction of 40%-50% in annual risk of recurrence 5. 6% improvement in 10 -year survival ER and progesterone receptor (PR, an indicator of a functional ER pathway) currently the best predicator of tamoxifen response 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 4

Introduction n Tamoxifen for breast cancer n n 25% of ER+/PR+, 66% of ER+/PR-, and 55% of ER-/PR- fail to respond Identify the differentially expressed genes in recurrence To predict tamoxifen treatment outcome in early-stage breast cancer Two-gene expression ratio predicts clinical outcome (Ma, et al, 2004) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 5

Introduction n n Determining differences between onparole prisoners who have and have not violated their parole Identifying salient attributes of differentiating between purchasers and non-purchasers of brands and predicting purchase intention of potential customers 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 6

Introduction n Objectives n n n Identify the variables that discriminate best between groups Develop a index function to parsimoniously represent the differences between groups based on the identified variables Develop a decision rule to classify future observations into one of the groups 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 7

Introduction n Not to discuss how to identify variables that best differentiate groups Assume that the number of groups (populations) is known in advance Focus on development of index function, decision rule and classification errors 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 8

Methods for Two Populations n n Fisher Linear Discriminant Function Data Structure: p-variables Group 1 X 12. . X 1 n 1 1/19/2022 2 X 21 X 22. . X 2 n 2. Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 9

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 10

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 11

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 12

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 13

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 14

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 15

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 16
1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 17

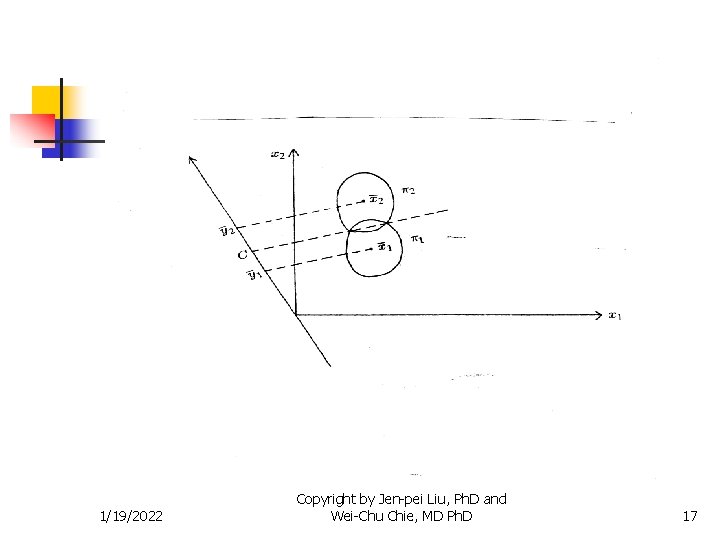
Methods for Two Populations n Example 1: Shen (1998) n Populations n n n Normal female volunteers (n=31) Female haemophilia (n=37) Variables n n 1/19/2022 Coagulant activity of factor VIII % (X 1) Related antigen of factor VIII % (X 2) Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 18

Methods for Two Populations n Example 1: Shen (1998) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 19

Methods for Two Populations n Example 1: Shen (1998) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 20

Methods for Two Populations n Example 1: Shen (1998) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 21

Methods for Two Populations n Example 1: Shen (1998) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 22
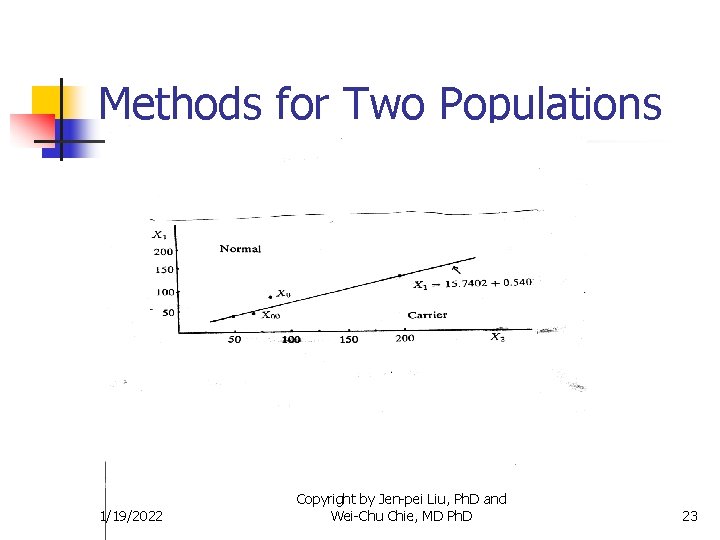
Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 23

Methods for Two Populations n Example 1: Shen (1998) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 24

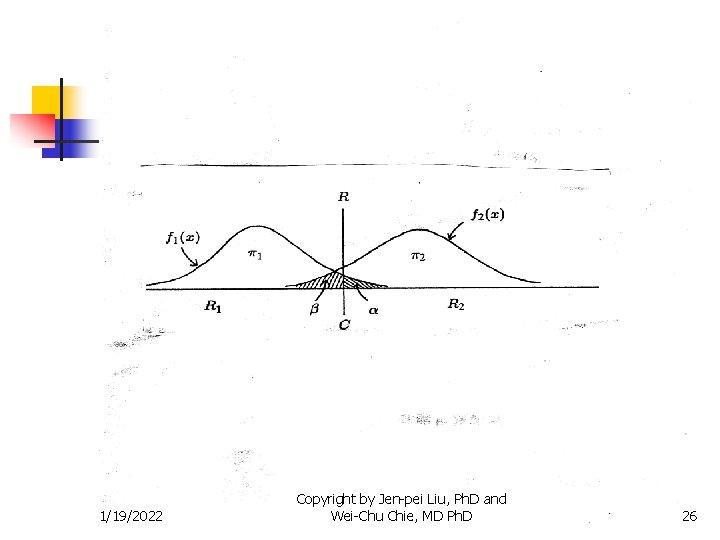
Methods for Two Populations n Classification with known distributions 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 25
1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 26

Methods for Two Populations n Classification with known distributions 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 27

Methods for Two Populations n Classification with known distributions 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 28

Methods for Two Populations n Classification with known distributions 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 29

Methods for Two Populations n Classification with known distributions n n n Example 2: Shen (1998) and Johnson & Wichern (1998) Populations n Normal female volunteers (n=30) n Female Haemophilia (n=22) Variables n AHF activity % (X 1) n AHF antigen % (X 2) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 30

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 31

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 32

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 33

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 34

Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 35

Methods for Two Populations n Optimal Classification True Population I II 1/19/2022 Classification Decision I II 0 C(2|1) C(1|2) 0 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 36

Methods for Two Populations n Optimal Classification 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 37

Methods for Two Populations n Optimal Classification 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 38

Methods for Two Populations n Estimation of Misclassification Rate Assumptions: Normal distribution and C(2|1)=C(1|2) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 39

Methods for Two Populations n Estimation of Misclassification Rate Assumptions: Normal distribution and C(2|1)=C(1|2) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 40

Methods for Two Populations n n Estimation of Misclassification Rate Example 2 (Continued) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 41

Methods for Two Populations n n Estimation of Misclassification Rate Example 2 (Continued) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 42

Methods for Two Populations n n Estimation of Misclassification Rate Example 2 (Continued) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 43

Methods for Two Populations n Estimation of Misclassification Rate If the true form of distribution is unknown True Classification Decision Population I II I a b II c d Overall accuracy = (a+d)/(a+b+c+d) Sensitivity = a/(a+b) False + rate = c/(a+c) Specificity = d/(c+d) False – rate =b/(b+d) 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 44

Methods for Two Populations n n Fisher linear discriminant function required existence inverse of the pooled sample covariance matrix It requires that n 1+n 2 -2 > p (# of genes) In microarray experiments, n 1+n 2 << p and the sample covariance matrix is singular Prediction by discriminant function is even more important in microarray experiment 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 45

Methods for Two Populations n Diagonal Linear Discriminant Function 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 46

Methods for Two Populations The compound covariate discriminant function 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 47

Methods for Two Populations The compound covariate discriminant function 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 48

Methods for Two Populations n Misclassification Rate n Fisher linear discriminant function and methods of calculation of misclassifiction rate assumes n n n 1/19/2022 Variables used in discriminant function is known Distribution form Prevalenece Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 49

Methods for Two Populations n Misclassification Rate n Application to predict diseases or response of treatment based on expression data from microarray experiment n n n 1/19/2022 Differentially expressed genes are not identified Distribution of expression data is unknown and is unlikely to follow normal distribution Prevalence of expressed genes is also unknown Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 50

Methods for Two Populations n Misclassification Rate n Re-substitution method n n 1/19/2022 Use the sample to select expressed genes Use the selected expressed genes in the sample to obtain the discriminant function Use the discriminant function to classify each member in the sample Compare the true class member and misclass member to compute the misclassification rate Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 51

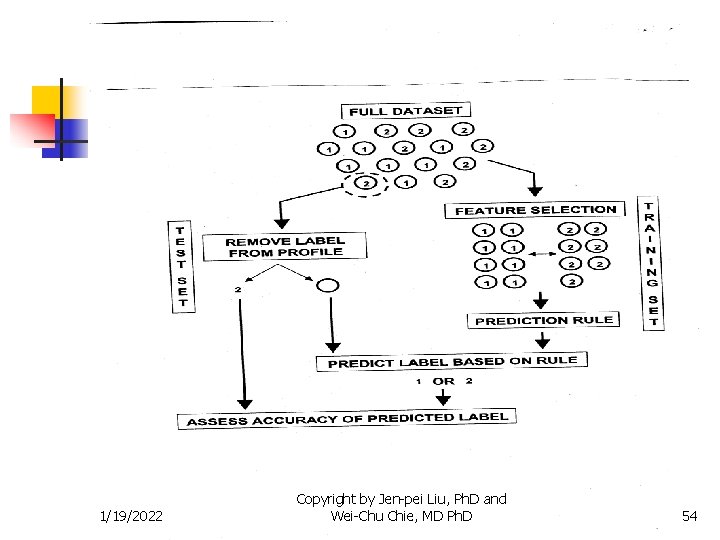
Methods for Two Populations n Misclassification Rate n Cross-validation after gene selection n n Use the sample to find the expressed genes Randomly divided the sample into n n n 1/19/2022 Training set Testing set Use the training set to obtain the discriminant function Use the training set to classify each member of the testing set Compare the true class member and predicted class member to compute the misclassification rate Leave-one-out method when # of case in testing set is 1 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 52

Methods for Two Populations n Misclassification Rate n Cross-validation before gene selection n Randomly divided the sample into n n n Use the training set n n 1/19/2022 Training set Testing set to find the expressed genes and to obtain the discriminant function Use the training set to classify each member of the testing set Compare the true class member and predicted class member to compute the misclassification rate Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 53
1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 54

Methods for Two Populations n Misclassification Rate n Re-substitution method provides a biased estimate of misclassification rate n n Parameters of the model are optimized to fit the data and the model will fit those data better than they will predict for the independent data – issue of overfitting The bias will be large when # of genes >> # of cases 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 55

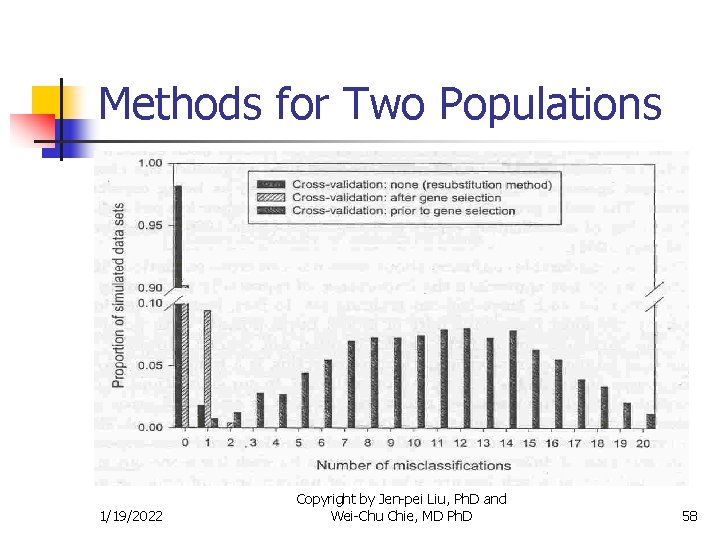
Methods for Two Populations n n Misclassification Rate Simulation by Simon et al. n n n Expression levels of 6000 genes were randomly generated under the same normal distribution for 20 specimens Specimen 1 to 10 were arbitrarily considered to be from population I and 11 -20 from population II Therefore the misclassification rate should be close to 50% 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 56

Methods for Two Populations n n Misclassification Rate Simulation by Simon et al. n n Simulation will repeated 2000 times (datasets) The misclassification rate for re-substitution method is 1. 8% The misclassification rate for cross-validation after gene selection is 9. 8% The misclassification rate for cross-validation before gene selection is around 50% 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 57
Methods for Two Populations 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 58

Methods for Several Populations n Fisher Linear Discriminant Function 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 59

Methods for Several Populations n Fisher Linear Discriminant Function 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 60

Methods for Several Populations n n Minimum Distance Method Assumption: Homogeneous covariance matrix 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 61
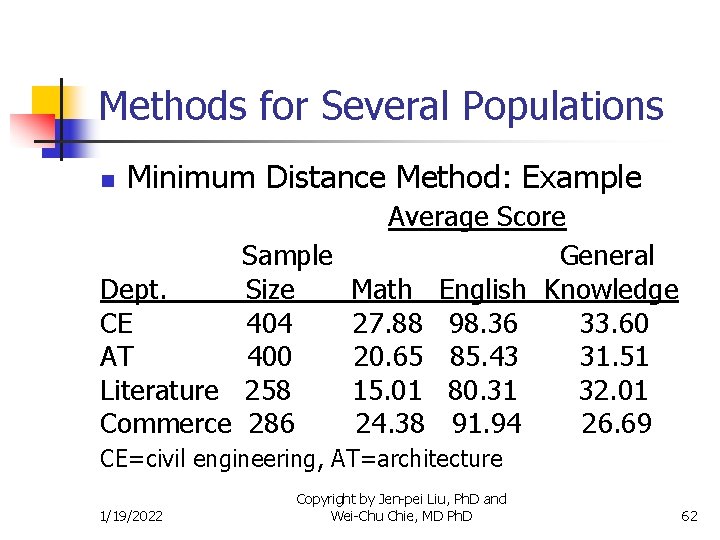
Methods for Several Populations n Minimum Distance Method: Example Average Score Sample General Dept. Size Math English Knowledge CE 404 27. 88 98. 36 33. 60 AT 400 20. 65 85. 43 31. 51 Literature 258 15. 01 80. 31 32. 01 Commerce 286 24. 38 91. 94 26. 69 CE=civil engineering, AT=architecture 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 62

Methods for Several Populations n Minimum Distance Method: Example 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 63

Methods for Several Populations Minimum Distance Method: Example A new student with scores of (20 90 30) n 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 64

Methods for Several Populations Minimum Distance Method: Example A new student with scores of (20 90 30) n 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 65

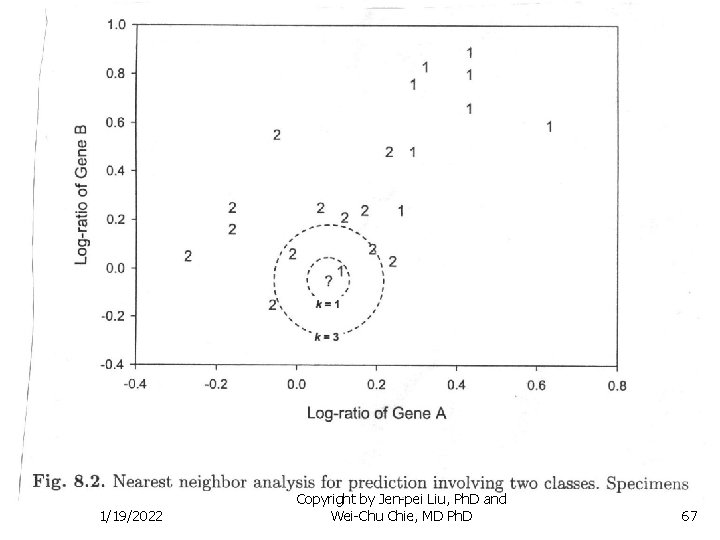
Methods for Several Populations n Nearest Neighbor Classification Method n Select a distance measure between two data points n n n (Xi - Xj)C-1(Xi - Xj) For a data point xo, select the number of data points closest to xo, say K If among K data points, K 1 points belong to population I and K 2 points belong to population II and K 1 > K 2, classify xo to population I; otherwise to population II 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 66
1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 67

Methods for Several Populations n Nearest Neighbor Classification Method n Example: n n n K=1, classify “? ” to population 1 K=3, Two “ 2” and one “ 1”, classify “? ” to population 2 Optimal number of K is less than 7 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 68

Methods for Several Populations n Nearest Neighbor Classification Method n n If sample size are different, classify xo, to population I if K 1/n 1 > K 2/n 2, otherwise to population II If both sample size and prior probability are different, classify xo, to population I if (K 1/n 1)/(K 2/n 2) > p 1/p 2, otherwise to population II 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 69

Summary n n n Objectives of discriminant analysis Fisher linear discriminant function for two populations Misclassification rate Discriminant function for three populations Application of discriminant analysis to to microarray data Nonparametric method: nearest neighbor rule 1/19/2022 Copyright by Jen-pei Liu, Ph. D and Wei-Chu Chie, MD Ph. D 70
- Slides: 70