Applied Bioinformatics Introduction to Linux and R Bing

Applied Bioinformatics Introduction to Linux and R Bing Zhang Department of Biomedical Informatics Vanderbilt University bing. zhang@vanderbilt. edu
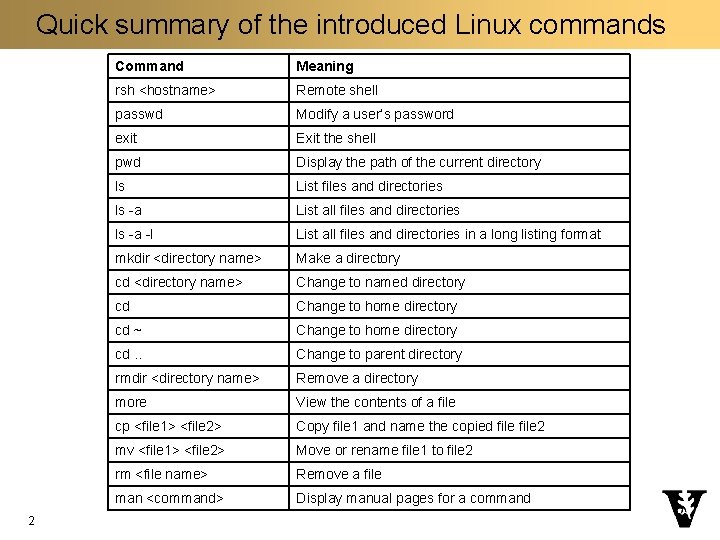
Quick summary of the introduced Linux commands 2 Command Meaning rsh <hostname> Remote shell passwd Modify a user’s password exit Exit the shell pwd Display the path of the current directory ls List files and directories ls -a List all files and directories ls -a -l List all files and directories in a long listing format mkdir <directory name> Make a directory cd <directory name> Change to named directory cd Change to home directory cd ~ Change to home directory cd. . Change to parent directory rmdir <directory name> Remove a directory more View the contents of a file cp <file 1> <file 2> Copy file 1 and name the copied file 2 mv <file 1> <file 2> Move or rename file 1 to file 2 rm <file name> Remove a file man <command> Display manual pages for a command

Getting help n 3 man (display manual pages for a command) q space bar to show next page q up and down arrows to move up and down q q to exist
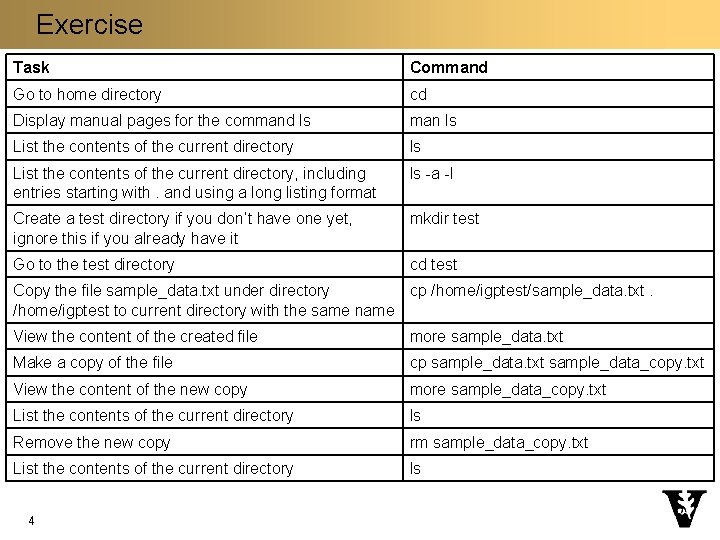
Exercise Task Command Go to home directory cd Display manual pages for the command ls man ls List the contents of the current directory, including entries starting with. and using a long listing format ls -a -l Create a test directory if you don’t have one yet, ignore this if you already have it mkdir test Go to the test directory cd test Copy the file sample_data. txt under directory cp /home/igptest/sample_data. txt. /home/igptest to current directory with the same name View the content of the created file more sample_data. txt Make a copy of the file cp sample_data. txt sample_data_copy. txt View the content of the new copy more sample_data_copy. txt List the contents of the current directory ls Remove the new copy rm sample_data_copy. txt List the contents of the current directory ls 4
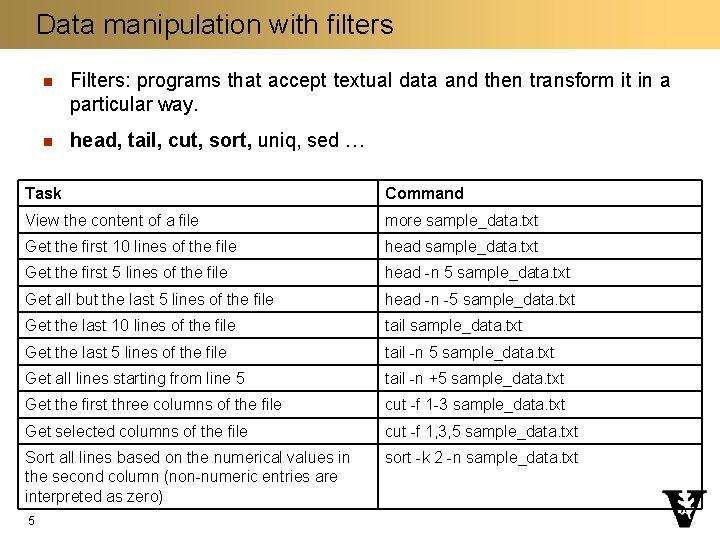
Data manipulation with filters n Filters: programs that accept textual data and then transform it in a particular way. n head, tail, cut, sort, uniq, sed … Task Command View the content of a file more sample_data. txt Get the first 10 lines of the file head sample_data. txt Get the first 5 lines of the file head -n 5 sample_data. txt Get all but the last 5 lines of the file head -n -5 sample_data. txt Get the last 10 lines of the file tail sample_data. txt Get the last 5 lines of the file tail -n 5 sample_data. txt Get all lines starting from line 5 tail -n +5 sample_data. txt Get the first three columns of the file cut -f 1 -3 sample_data. txt Get selected columns of the file cut -f 1, 3, 5 sample_data. txt Sort all lines based on the numerical values in the second column (non-numeric entries are interpreted as zero) sort -k 2 -n sample_data. txt 5

Data manipulation with piping and redirection n Piping (|) : sending data from one program to another program. n Redirection: sending output from one program to a file q >: save output to a file q >>: append output to a file Task Command Get the first 10 lines of the file and then get the first three columns head sample_data. txt | cut -f 1 -3 Get the first 10 lines of the file, then get the first three columns of these lines, and then redirect the content to a new file head sample_data. txt | cut -f 1 -3 >sample_data_subset. txt View the new file more sample_data_subset. txt Append the last 10 lines of the old file to the end of the new file tail sample_data. txt >> sample_data_subset. txt View the new file more sample_data_subset. txt 6
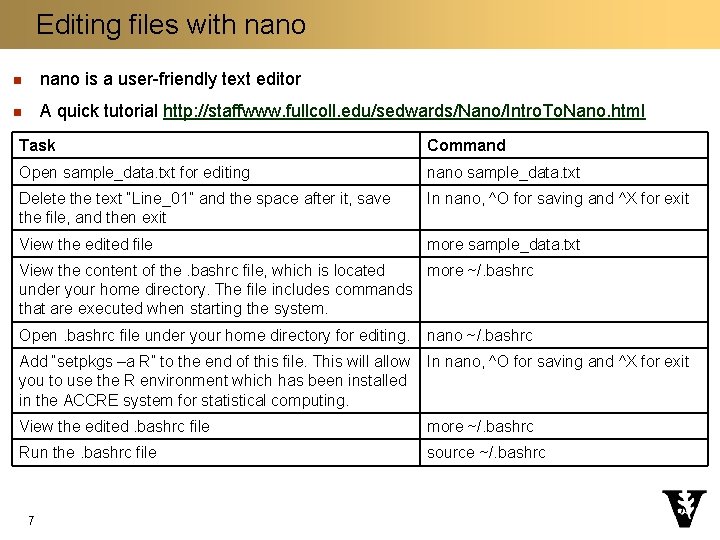
Editing files with nano n nano is a user-friendly text editor n A quick tutorial http: //staffwww. fullcoll. edu/sedwards/Nano/Intro. To. Nano. html Task Command Open sample_data. txt for editing nano sample_data. txt Delete the text “Line_01” and the space after it, save the file, and then exit In nano, ^O for saving and ^X for exit View the edited file more sample_data. txt View the content of the. bashrc file, which is located more ~/. bashrc under your home directory. The file includes commands that are executed when starting the system. Open. bashrc file under your home directory for editing. nano ~/. bashrc Add “setpkgs –a R” to the end of this file. This will allow you to use the R environment which has been installed in the ACCRE system for statistical computing. In nano, ^O for saving and ^X for exit View the edited. bashrc file more ~/. bashrc Run the. bashrc file source ~/. bashrc 7

What is R n R is a free software environment for statistical computing and graphics. It includes: q q q 8 an effective data handling and storage facility a suite of operators for calculations on arrays, in particular matrices a large, coherent, integrated collection of intermediate tools for data analysis graphical facilities for data analysis and display either on-screen or on hardcopy a well-developed, simple and effective programming language which includes conditionals, loops, user-defined recursive functions and input and output facilities

R Installation and tutorial n n 9 Download and install R q http: //www. r-project. org/ q Choose a CRAN (Comprehensive R Archive Network) mirror q Binary distributions of the base system and contributed packages n Windows version n Mac OS X version n Linux version (already installed on the ACCRE cluster, will be used for this module) Tutorials q http: //cran. r-project. org/doc/manuals/r-release/R-intro. html q An introduction to R
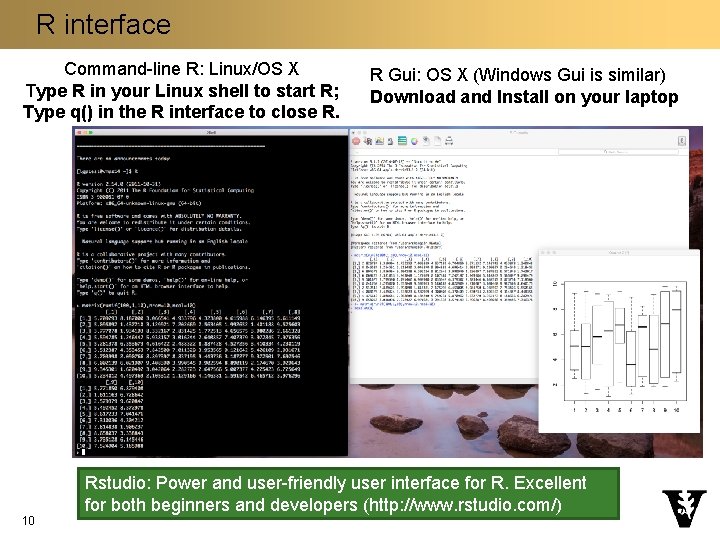
R interface Command-line R: Linux/OS X Type R in your Linux shell to start R; Type q() in the R interface to close R. 10 R Gui: OS X (Windows Gui is similar) Download and Install on your laptop Rstudio: Power and user-friendly user interface for R. Excellent for both beginners and developers (http: //www. rstudio. com/)

Install and load packages n n CRAN packages q http: //cran. r-project. org/web/packages/ q >6000 packages Bio. Conductor packages q http: //www. bioconductor. org/ q ~1000 packages for the analysis of high-throughput genomics data Task R code Install a CRAN package install. packages (“package name”) Install a Bio. Conductor package souce (“http: //www. bioconductor. org/bioc. Lite. R”) bioc. Lite (“package name”) Load a package/library (“package name”) 11

Basic R syntax n Object <- function (arguments) q n <-: assignment operator Object <- object[arguments] Task R code Assign a numeric vector with five numbers to object x using the c() function x <- c(1. 3, 10. 4, 5. 6, 3. 1, 6. 4, 21. 7) Assign a subset of x to a new object y y <- x[1: 3] Show the content of x x Show the content of y y Getting information on function c ? c Display the output of a function without assignment c(1, 2, 5) 12

Data types n Numeric data q n Character data q n 1, 2, 3 “a”, “b”, “c” Logical data q TRUE, FALSE, TRUE Task R code Assign a numeric vector with five numbers to object x using the c() function x <- c(1. 3, 10. 4, 5. 6, 3. 1, 6. 4, 21. 7) Create a character vector from x as. character(x) Create a logical vector from x x>5 13

Data objects n Vectors: an ordered collection of items of the same data type (numeric, character, or logical), 1 -dimensional n Matrices: 2 -dimensional objects, all items must have the same data type n Arrays: similar to matrices but can have more than two dimensions n Data frames: similar to a matrices but can have different data types n Lists: an ordered collection of objects n Functions Task R code Create a numeric vector with numbers ranging from 1 to 9 c(1: 9) Create a 3 x 3 numeric matrix(c(1: 9), nrow=3, ncol=3, byrow=TRUE) Create another 3 x 3 numeric matrix by changing an argument matrix(c(1: 9), nrow=3, ncol=3, byrow=FALSE) 14
Operators and calculations n Comparison operators: ==, !=, <, >, <=, >= n Logical operators: & (AND), | (OR), ! (NOT) n Calculations q Arithmetic operators: +, -, *, /, ^ q Arithmetic functions: log, exp, sqrt, mean, var, sd, sum, etc. Task R code Comparisons 3==5 3!=5 3<5 Logical operators x<-5 y<-(-8) x>0 | y>0 x>0 & y>0 Calculations (4+2^2)/(2*2) x<-c(1, 3, 5, 7, 9) y<-c(2, 4, 6, 8, 10) x+y sum((x-mean(x))^2)/(length(x)-1) var(x) 15
Data import, simple analyses, and export Task R code Import data from a tabular file my. Data<-read. table(“~/test/sample_data. txt”, head=T, sep=“t”) Display the new object my. Data Get class name of the object class(my. Data) Convert data frame to matrix my. Matrix<-as. matrix(my. Data) Get class name of the matrix class(my. Matrix) Display the matrix object my. Matrix Get dimensions of the matrix dim(my. Matrix) Get a high-level summary(my. Matrix) Log transformation of the data my. Matrix_log<-log 2(my. Matrix) Calculate variance for row #1 var(my. Matrix_log[1, ]) Calculate variances for all rows variances<-apply(my. Matrix_log, 1, var) Calculate means for all rows means<-apply(my. Matrix_log, 1, mean) Data subsetting my. Matrix_log[1: 3, 1: 2] my. Matrix_log[c(“Line_02”, ”Line_04”), ] my. Matrix_log[means>median(means), ] Combining data results<-cbind(my. Matrix_log, means, variances) Write data to a tabular file write. table(results, “~/test/sample_data_output. txt”, sep=“t”, quote=FALSE) Quit R q() 16 Go to your test directory, and check the file sample_data_output. txt
Copying files to/from a local computer n Windows q Application: Bitvise SSH (https: //www. bitvise. com/ssh-client-download) n Mac q Application: Cyberduck (https: //cyberduck. io/) 17 q Click on “Open Connection” q Select “SFTP (SSH File Transfer Protocol)” q Server: vmplogin. accre. vanderbilt. edu q Username: your_user_name q Password: your-password q Don’t change other items
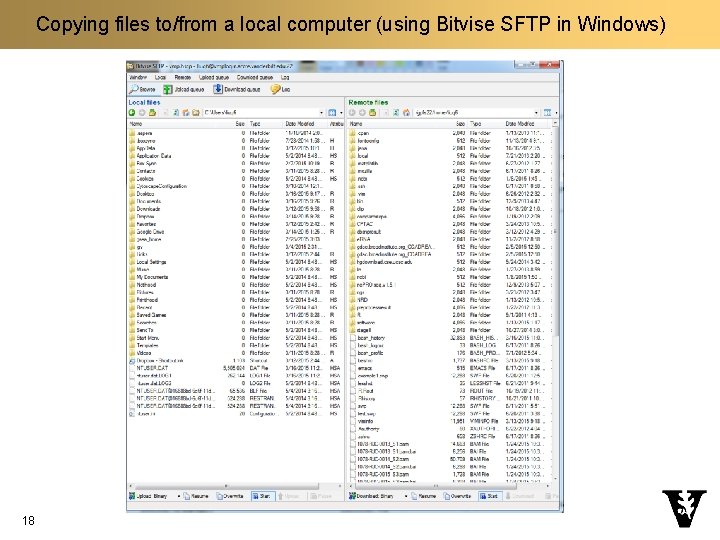
Copying files to/from a local computer (using Bitvise SFTP in Windows) 18
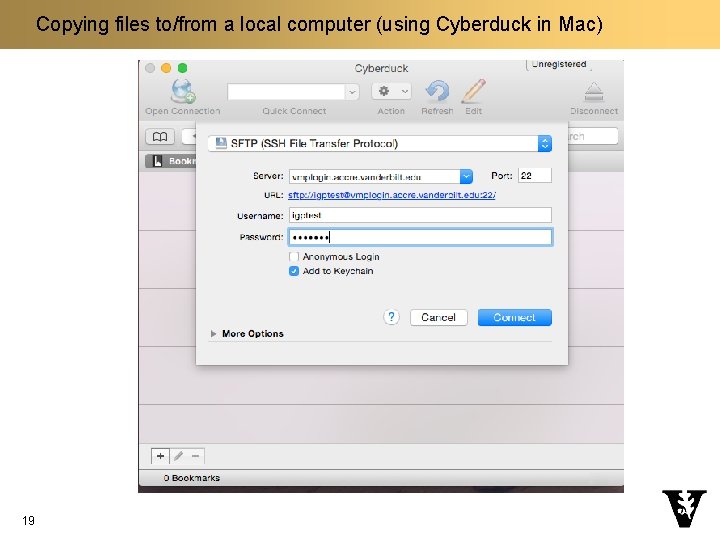
Copying files to/from a local computer (using Cyberduck in Mac) 19
- Slides: 19