Applications Using standard Bioinformatics applications Introduction The overall

Applications Using standard Bioinformatics applications

Introduction
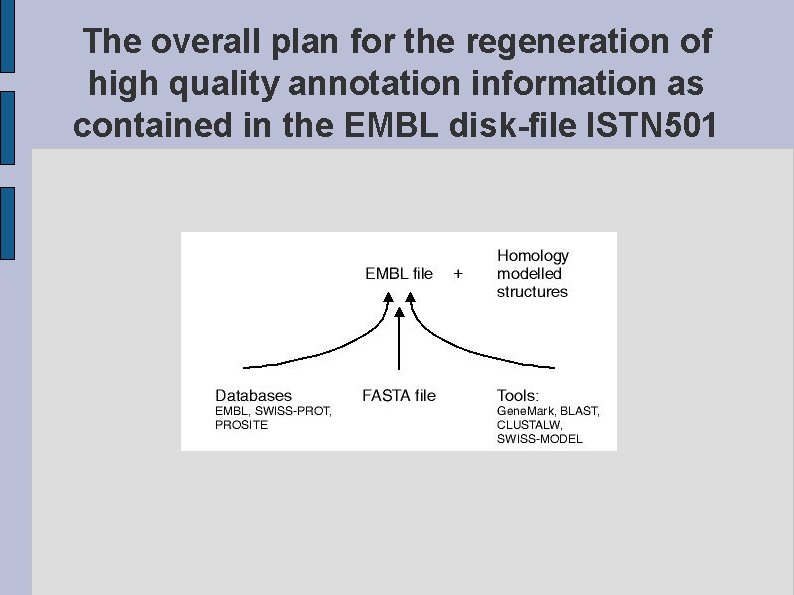
The overall plan for the regeneration of high quality annotation information as contained in the EMBL disk-file ISTN 501 fig. WHAT. eps

Scientific Background To Mer Operon ● Function ● Genetic Structure and Regulation ● Mobility Of The Mer Operon
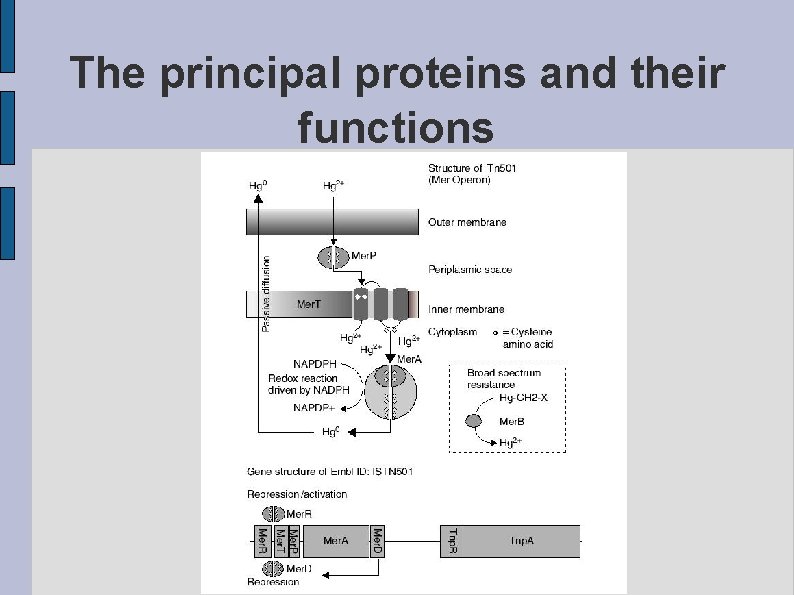
The principal proteins and their functions fig. PRINCIPLE. eps

Downloading The Raw DNA Sequence

Initial BLAST Sequence Similarity Search

Maxim 18. 1 With BLAST scores, up is down and lower is better

Gene. Mark http: //opal. biology. gatech. edu/Gene. Mark/

The web-based interface to Gene. Mark as running at EBI fig. EBIGENEBANK. eps

Using BLAST to identify specific sequences

Dealing with false negatives and missing proteins

Over predicted genes and false positives

Structural Prediction With SWISS-MODEL http: //www. expasy. org/swissmod/

Maxim 18. 2 The major limitation of ``homology modelling'' is that homology to a known structure is needed

Alternatives to homology modelling

Modelling with SWISS-MODEL
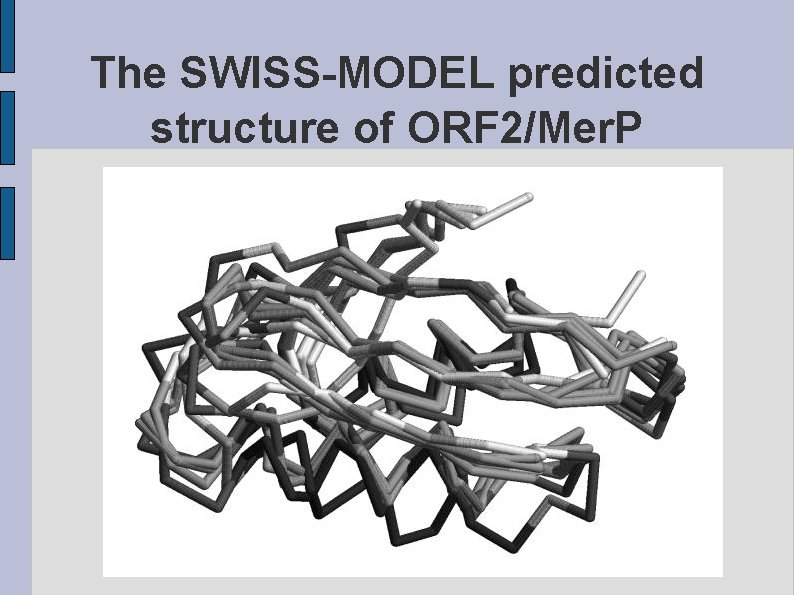
The SWISS-MODEL predicted structure of ORF 2/Mer. P fig. ORF 2 MERP. eps
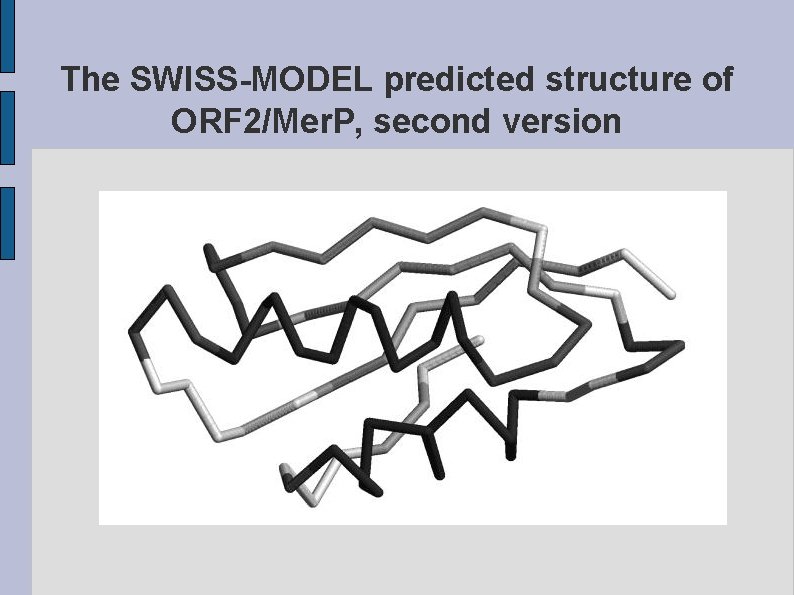
The SWISS-MODEL predicted structure of ORF 2/Mer. P, second version fig. ORF 2 MERP 2. eps
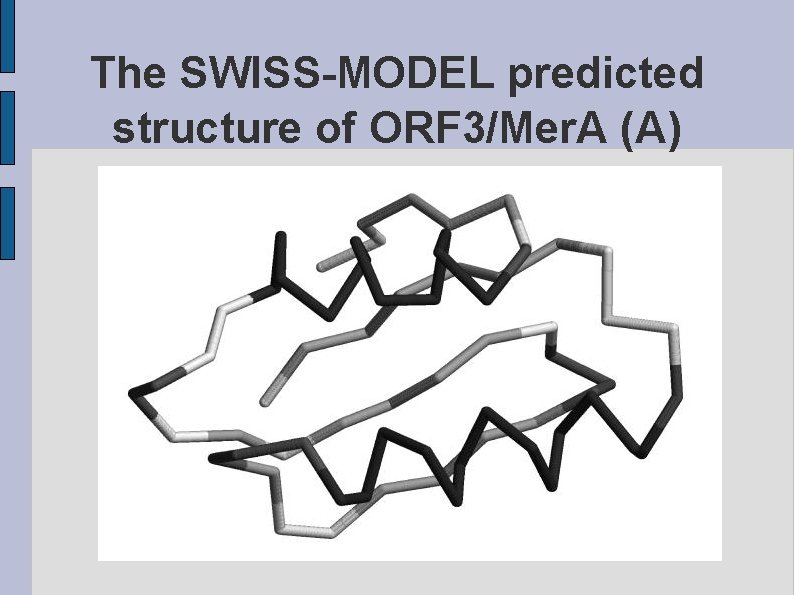
The SWISS-MODEL predicted structure of ORF 3/Mer. A (A) fig. ORF 3 MERAA. eps
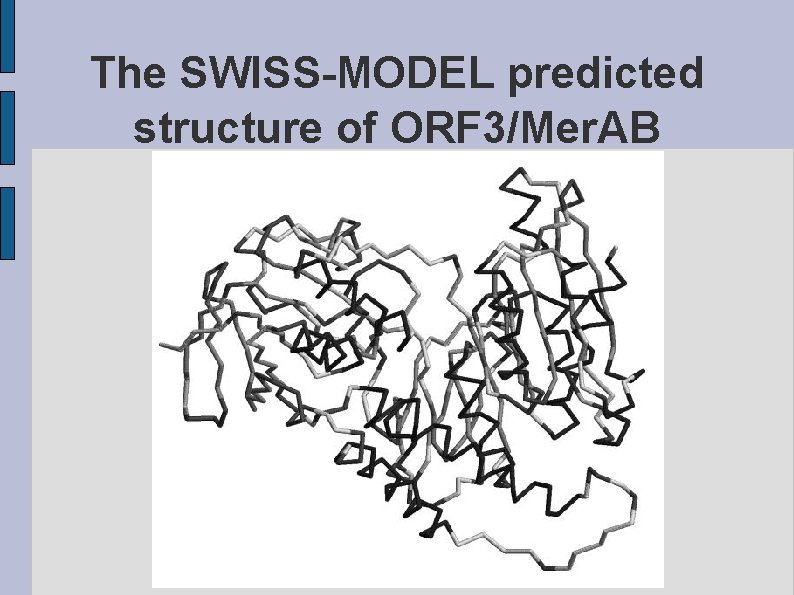
The SWISS-MODEL predicted structure of ORF 3/Mer. AB fig. ORF 3 MERAB. eps
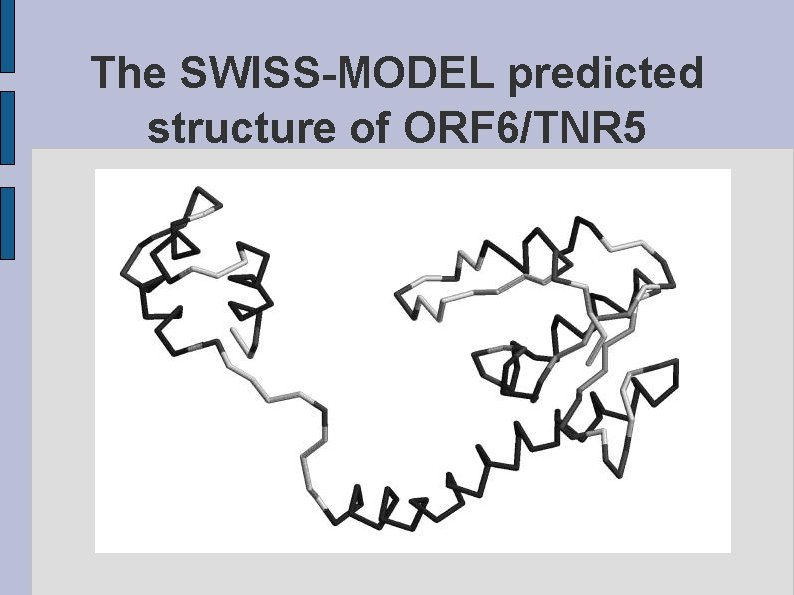
The SWISS-MODEL predicted structure of ORF 6/TNR 5 fig. ORF 6 TNR 5. eps

Deep. View as a Structural Alignment Tool

The ORF 2 and ORF 3_A structures loaded into Deep. View prior to structural alignment fig. DEEPVIEW. eps
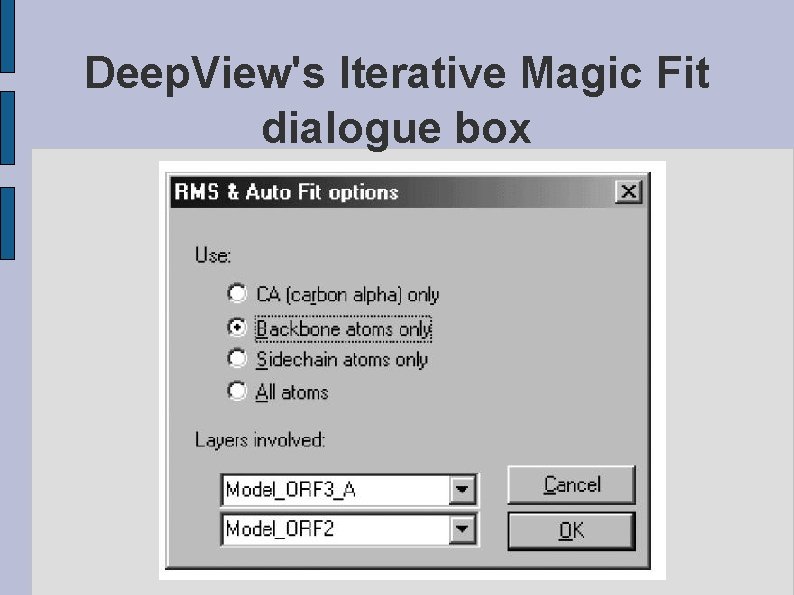
Deep. View's Iterative Magic Fit dialogue box fig. DEEPVIEWDIALOG. eps

Structural Alignment created using the Deep. View's Iterative Magic Fit facility fig. DEEPVIEWEXAMPLE. eps

Selecting the current ``layer'' in Deep. View fig. DEEPLAYER. eps
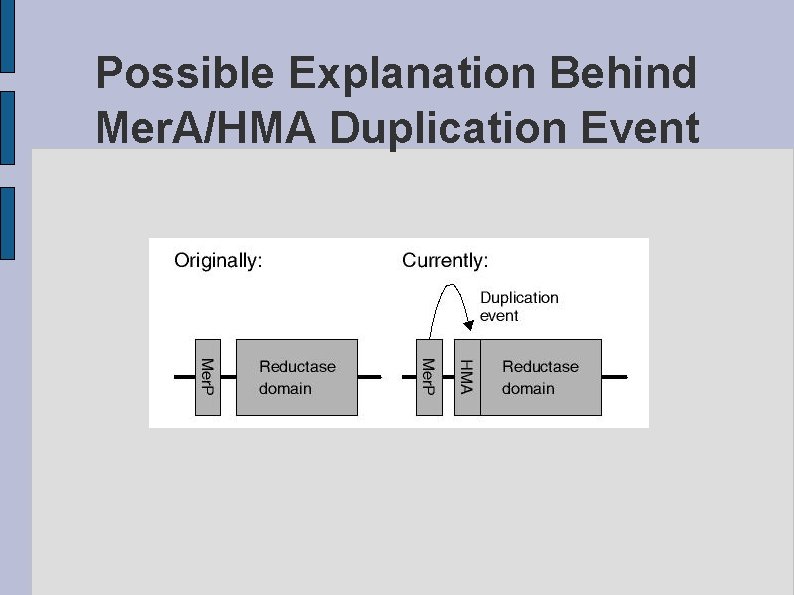
Possible Explanation Behind Mer. A/HMA Duplication Event fig. POSSIBLE. eps
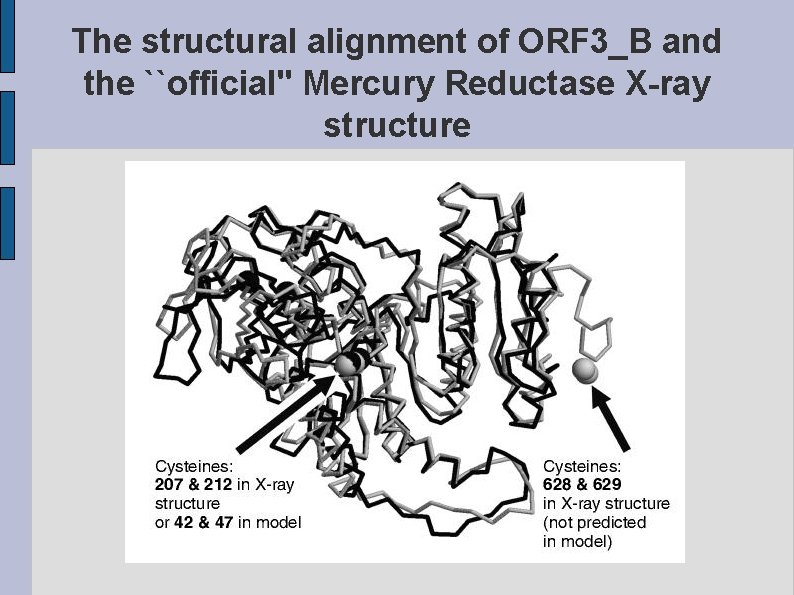
The structural alignment of ORF 3_B and the ``official'' Mercury Reductase X-ray structure fig. CYSTEINES. eps

Maxim 18. 3 Homology modelling can only model protein sequences similar to those which are already known

PROSITE and Sequence Motifs

Maxim 18. 4 Searching large datasets with non-specific, short sequence fragments results in many false positives

Using PROSITE patterns and matrices http: //www. expasy. org/prosite/ http: //www. ebi. ac. uk/interpro/ http: //www. geneontology. org ● http: //www. kegg. org

Phylogenetics

A look at the HMA domain of Mer. A and Mer. P ----------------SWISS-PROT IDs of Mer. P Proteins SWISS-PROT IDs of Mer. A Proteins ----------------MERP_ACICA MERA_ACICA MERP_ALCSP MERA_ALCSP MERP_PSEAE MERA_BACSR MERP_PSEFL MERA_ENTAG MERP_SALTI MERA_PSEAE MERP_SERMA MERA_PSEFL MERP_SHEPU MERA_SERMA MERP_SHIFL MERA_SHEPU MERA_SHIFL MERA_STAEP MERA_STRLI MERA_THIFE -------------------------------

The multiple sequence alignment of the example proteins fig. LISTMERAMERP. eps

The EBI's tree graphical display fig. TREE. eps

Maxim 18. 5 Whenever you make a statement, call for more research (money)!

Maxim 18. 6 Database annotation is hard to do well, so be prepared to update it on a regular basis

Maxim 18. 7 Automation can be very helpful when creating annotation, but to achieve the highest quality, humans are needed to make some value judgments

Maxim 18. 8 Conclusions are based on the available data which, in this case, is the database annotation (which may or may not be current)

Where To From Here?
- Slides: 42