Application of Reduced Order Modeling to Time Parallelization








- Slides: 16

Application of Reduced Order Modeling to Time Parallelization Ashok Srinivasan, Yanan Yu, and Namas Chandra Florida State University http: //www. cs. fsu. edu/~asriniva Aim: Simulate for long time scales Solution features: Use data from prior simulations and experiments to parallelize the time domain Acknowledgements NSF, ORNL, NCSA

Outline • Limitations of Conventional Parallelization • Example Application: Carbon Nanotube Tensile Test – A Drawback of Molecular Dynamics Simulations • Small Time Step Size • Data-Driven Time Parallelization – Reduced order modeling is used for prediction • Experimental Results – Scaled efficiently to 400 processors, for a problem where conventional parallelization scales to just 2 -3 processors • Conclusions

Limitations of Conventional Parallelization • Conventional parallelization decomposes the state space across processors – It is effective for large state space – It is not effective when computational effort arises from a large number of time steps • … or when granularity becomes very fine due to a large number of processors

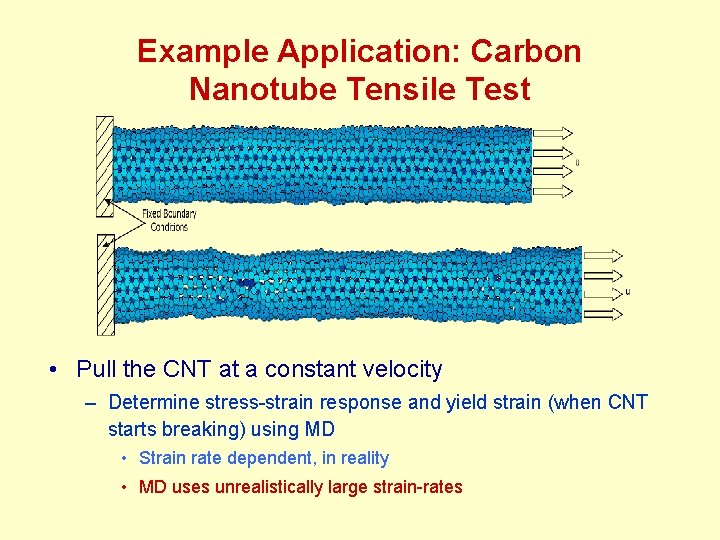
Example Application: Carbon Nanotube Tensile Test • Pull the CNT at a constant velocity – Determine stress-strain response and yield strain (when CNT starts breaking) using MD • Strain rate dependent, in reality • MD uses unrealistically large strain-rates

A Drawback of Molecular Dynamics Simulations • Molecular dynamics – In each time step, forces of atoms on each other modeled using some potential – After force is computed, update positions – Repeat for desired number of time steps • Time steps size ~ 10 – 15 seconds, due to physical and numerical considerations – Desired time range is much larger • A million time steps are required to reach 10 -9 s • Around a day of computing for a 3000 -atom CNT

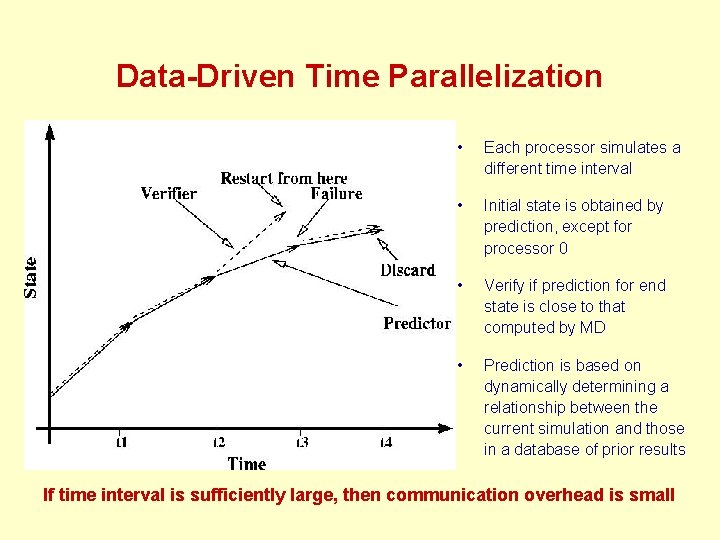
Data-Driven Time Parallelization • Each processor simulates a different time interval • Initial state is obtained by prediction, except for processor 0 • Verify if prediction for end state is close to that computed by MD • Prediction is based on dynamically determining a relationship between the current simulation and those in a database of prior results If time interval is sufficiently large, then communication overhead is small

Dimensionality Reduction • Movement of atoms in a 1000 -atom CNT is the motion of a point in 3000 -dimensional space • Find a lower dimensional subspace close to which the points lie • We use principal orthogonal decomposition – Find a low dimensional affine subspace • Motion may be complex in this subspace – Use results for different strain rates • Velocity = 10 m/s, 5 m/s, and 1 m/s – At five different time points • [U, S, V] = svd(Shifted Data) – Shifted Data = U*S*VT – States of CNT expressed as • m + c 1 u 1 + c 2 u 2 m

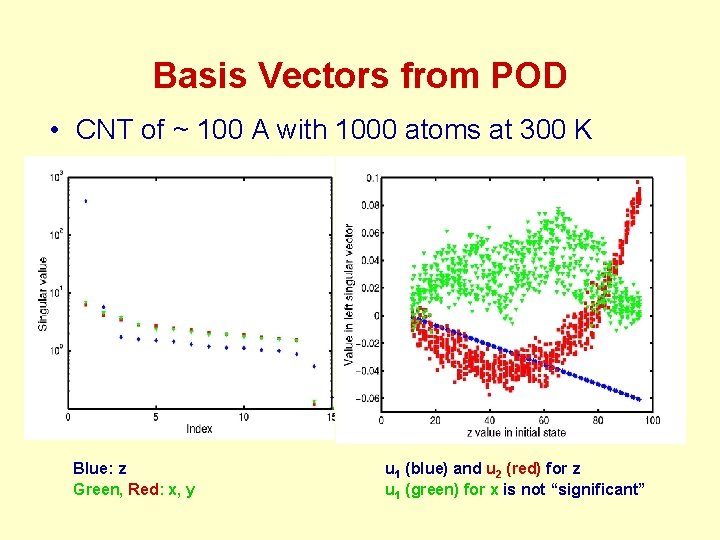
Basis Vectors from POD • CNT of ~ 100 A with 1000 atoms at 300 K Blue: z Green, Red: x, y u 1 (blue) and u 2 (red) for z u 1 (green) for x is not “significant”

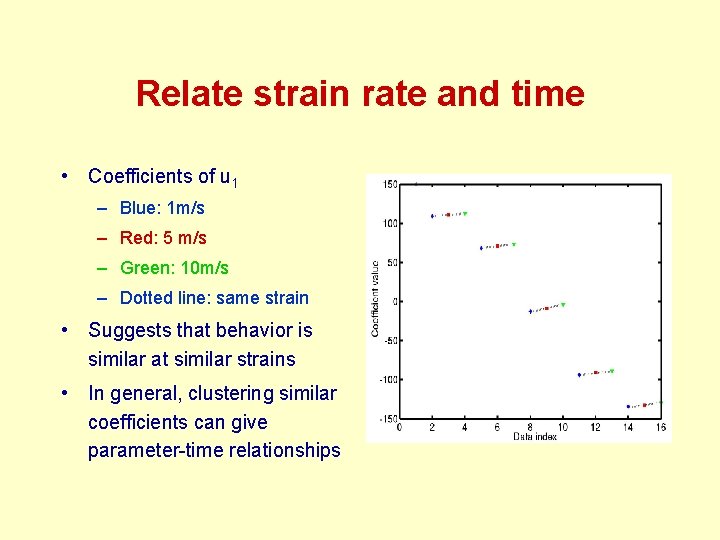
Relate strain rate and time • Coefficients of u 1 – Blue: 1 m/s – Red: 5 m/s – Green: 10 m/s – Dotted line: same strain • Suggests that behavior is similar at similar strains • In general, clustering similar coefficients can give parameter-time relationships

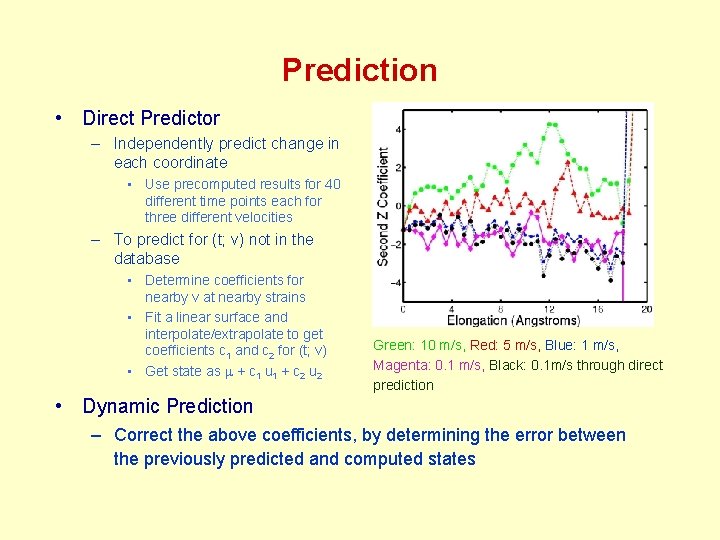
Prediction • Direct Predictor – Independently predict change in each coordinate • Use precomputed results for 40 different time points each for three different velocities – To predict for (t; v) not in the database • Determine coefficients for nearby v at nearby strains • Fit a linear surface and interpolate/extrapolate to get coefficients c 1 and c 2 for (t; v) • Get state as m + c 1 u 1 + c 2 u 2 Green: 10 m/s, Red: 5 m/s, Blue: 1 m/s, Magenta: 0. 1 m/s, Black: 0. 1 m/s through direct prediction • Dynamic Prediction – Correct the above coefficients, by determining the error between the previously predicted and computed states

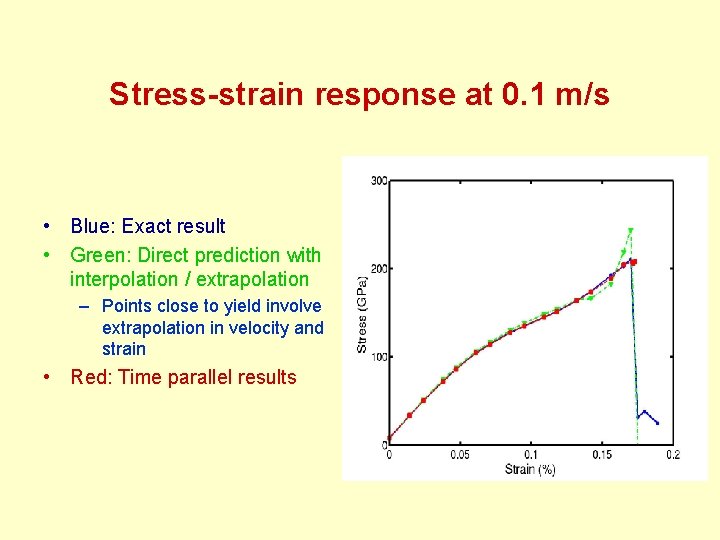
Stress-strain response at 0. 1 m/s • Blue: Exact result • Green: Direct prediction with interpolation / extrapolation – Points close to yield involve extrapolation in velocity and strain • Red: Time parallel results

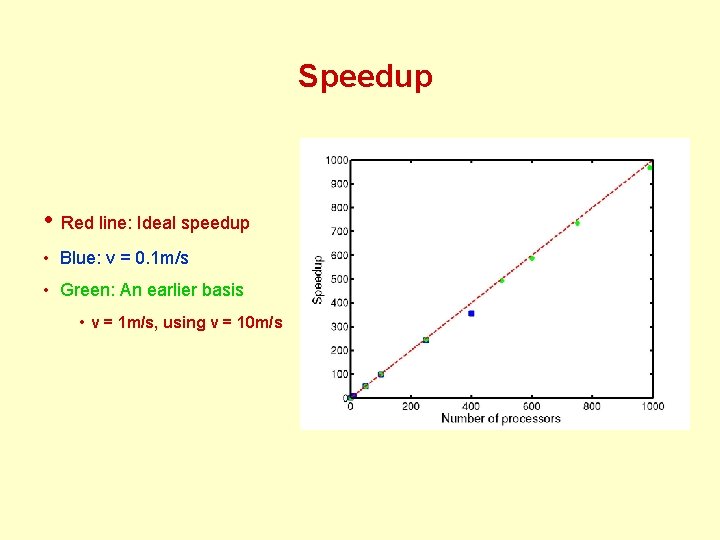
Speedup • Red line: Ideal speedup • Blue: v = 0. 1 m/s • Green: An earlier basis • v = 1 m/s, using v = 10 m/s

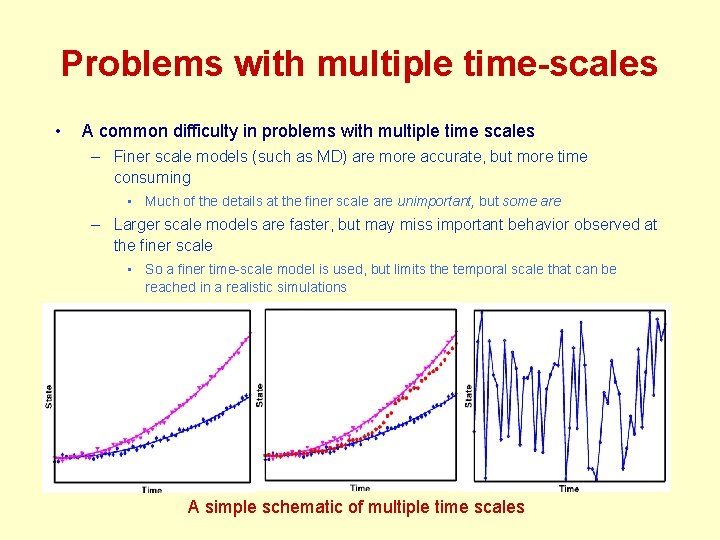
Problems with multiple time-scales • A common difficulty in problems with multiple time scales – Finer scale models (such as MD) are more accurate, but more time consuming • Much of the details at the finer scale are unimportant, but some are – Larger scale models are faster, but may miss important behavior observed at the finer scale • So a finer time-scale model is used, but limits the temporal scale that can be reached in a realistic simulations A simple schematic of multiple time scales

Solution • Use results of related finer scale simulations to model the significant effects on the larger scale – Example: (long time, high temperature/strain rate) -> (short time, low temperature/strain rate) – Technique: Reduced order modeling • Identify important modes of behavior • Relationships between simulation parameters – Technique: clustering – Interpolate from existing simulation results, to predict behavior when possible – Parallelization of time when unexpected modes might be significant • Technique: Learning

Conclusions • Time parallelization shows significant improvement in speed, without sacrificing accuracy significantly – This suggests that time can be considered, effectively, as a parallelizable domain • Direct prediction can yield several orders of magnitude improvement in performance when applicable

Future Work • More complex problems – Better prediction • POD is good for representing data, but not necessarily for identifying patterns • Use better dimensionality reduction / reduced order modeling techniques • Better learning in the predictor will also be useful – Simulations with multiple parameters • Example: Predict based on simulations that differ in temperature and strain rate • Such simulations may differ significantly from those in the database