Application of Data Independent Acquisition Techniques Optimized for


- Slides: 42

Application of Data Independent Acquisition Techniques Optimized for Improved Precursor Selectivity Jarrett D. Egertson, Ph. D. Mac. Coss Lab Department of Genome Sciences University of Washington 6/8/2013

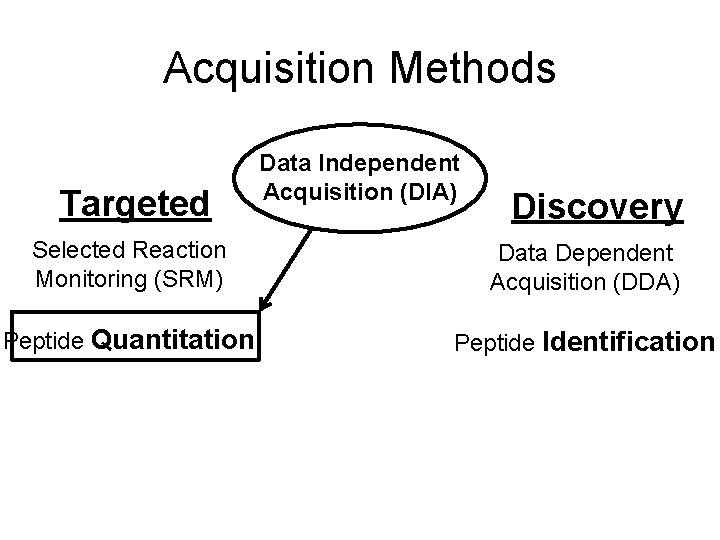
Acquisition Methods Targeted Data Independent Acquisition (DIA) Discovery Selected Reaction Monitoring (SRM) Data Dependent Acquisition (DDA) Peptide Quantitation Peptide Identification

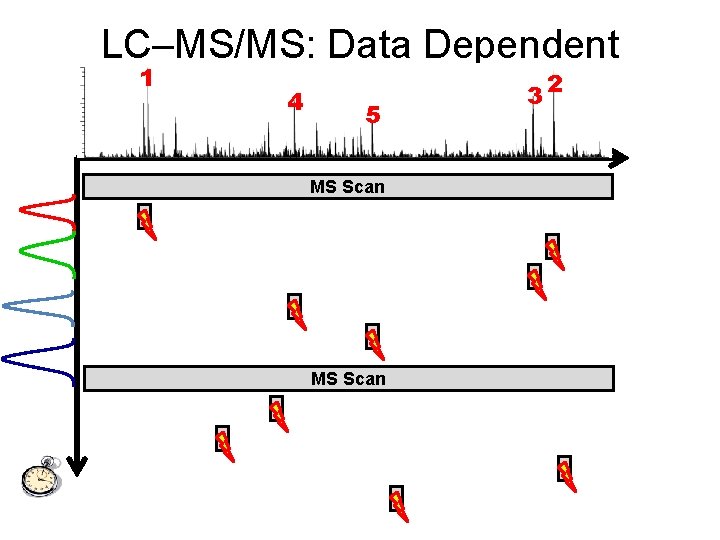
LC–MS/MS: Data Dependent 1 2 3 Acquisition 4 5 m/z MS Scan

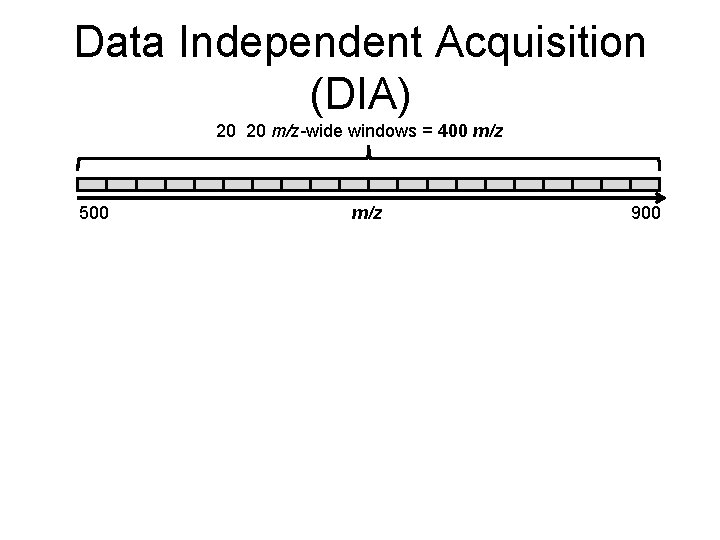
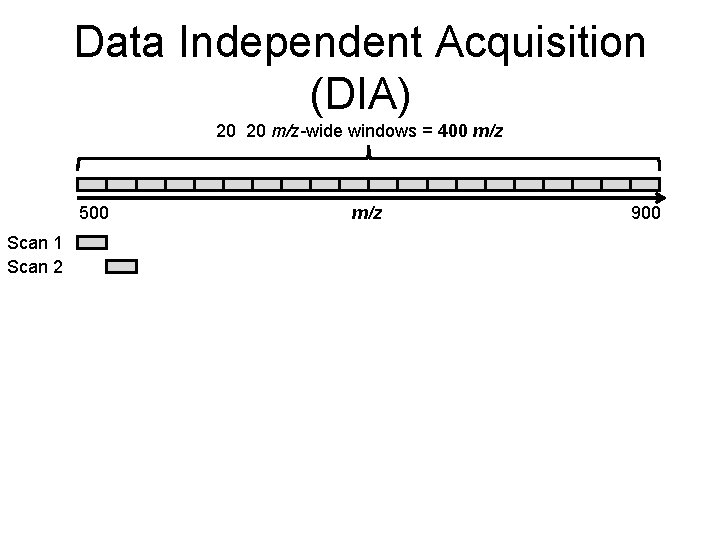
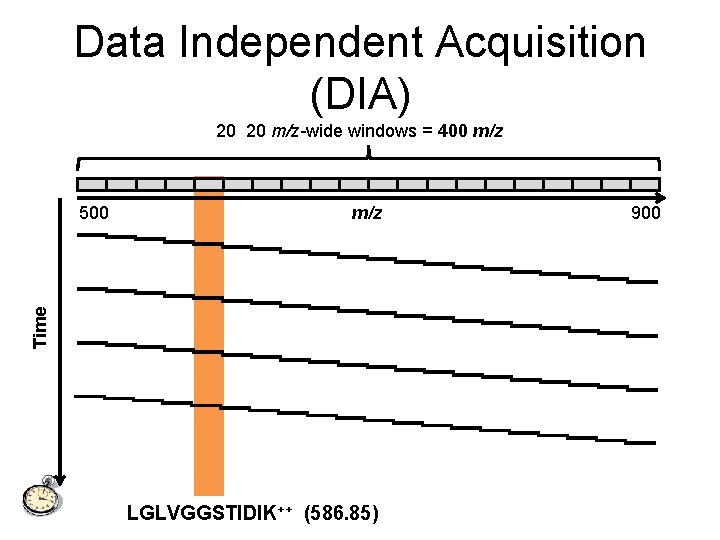
Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z 500 m/z 900

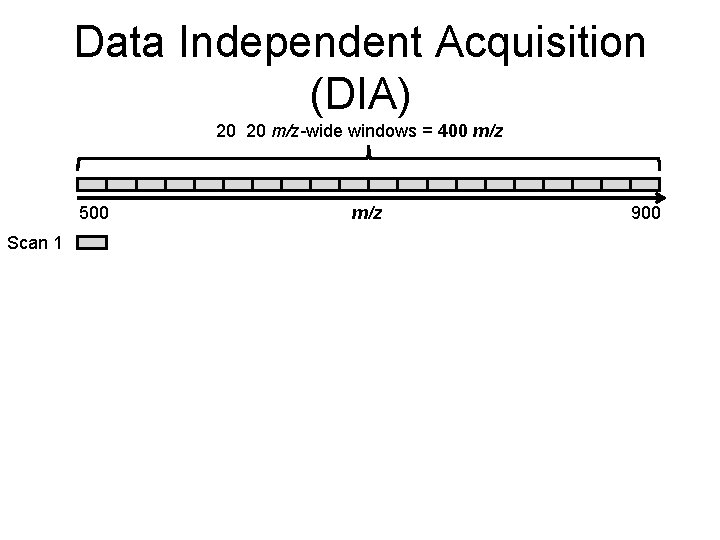
Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z 500 Scan 1 m/z 900

Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z 500 Scan 1 Scan 2 m/z 900

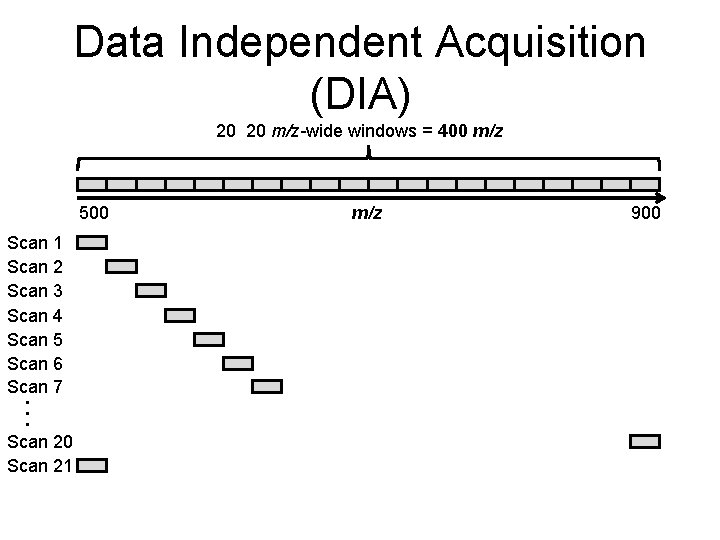
Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z 500 Scan 1 Scan 2 Scan 3 Scan 4 Scan 5 Scan 6 Scan 7 … Scan 20 Scan 21 m/z 900

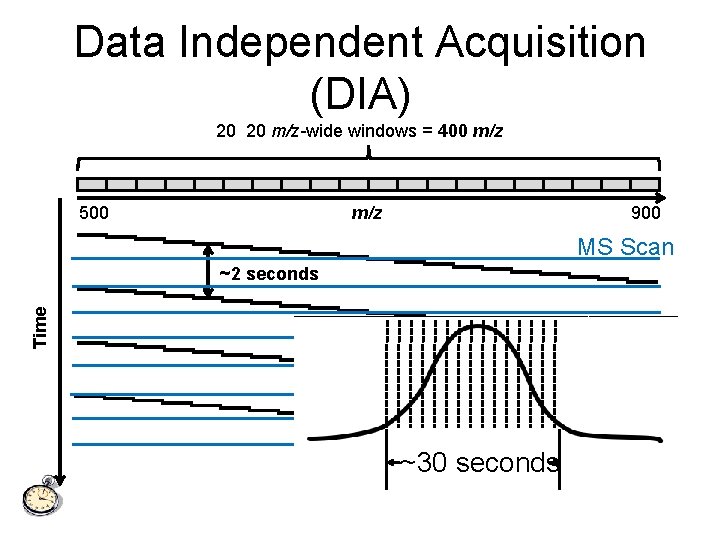
Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z 500 m/z 900 MS Scan Time ~2 seconds ~30 seconds

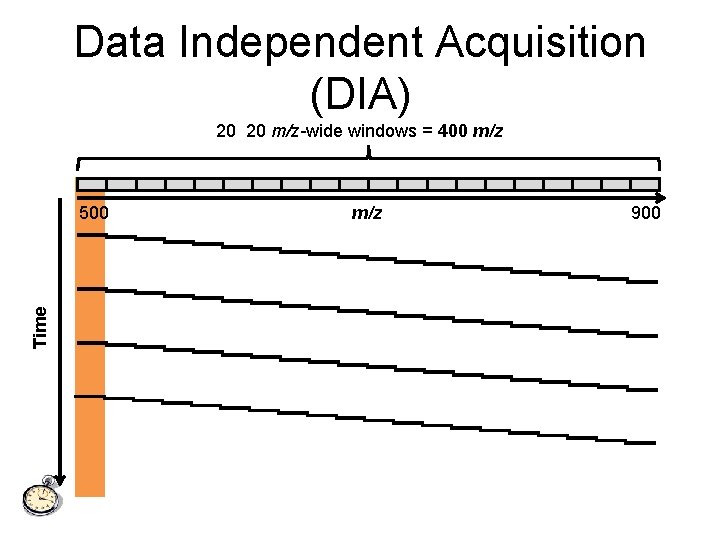
Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z Time 500 m/z 900

Data Independent Acquisition (DIA) 20 20 m/z-wide windows = 400 m/z Time 500 LGLVGGSTIDIK++ (586. 85) 900

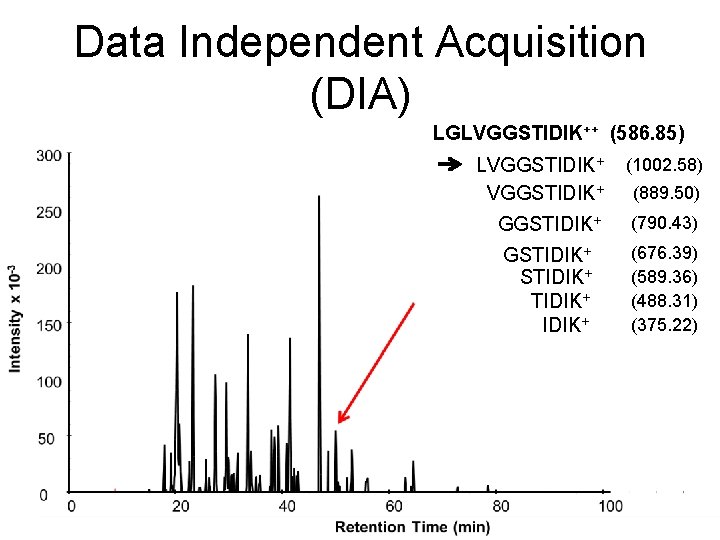
Data Independent Acquisition (DIA) LGLVGGSTIDIK++ (586. 85) LVGGSTIDIK+ (1002. 58) GGSTIDIK+ (790. 43) GSTIDIK+ (676. 39) (589. 36) (488. 31) (375. 22) (889. 50)

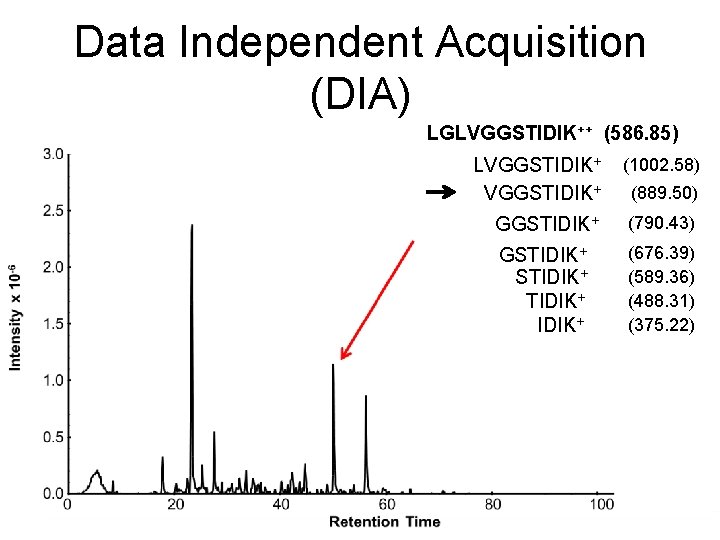
Data Independent Acquisition (DIA) LGLVGGSTIDIK++ (586. 85) LVGGSTIDIK+ (1002. 58) GGSTIDIK+ (790. 43) GSTIDIK+ (676. 39) (589. 36) (488. 31) (375. 22) (889. 50)

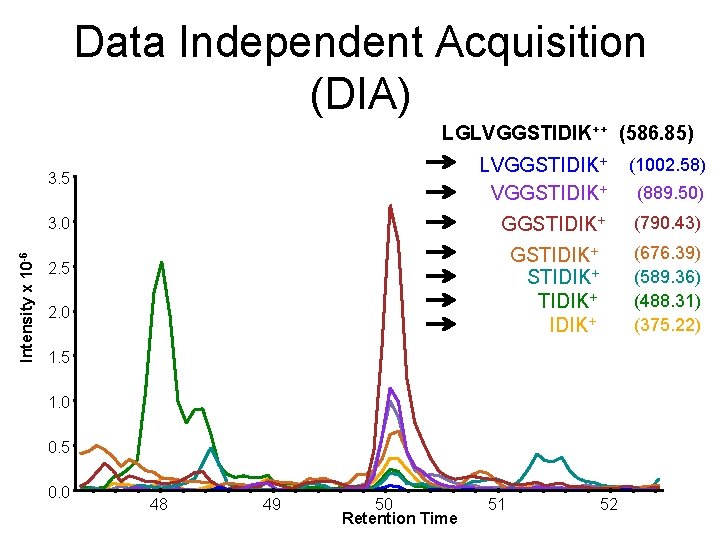
Data Independent Acquisition (DIA) Intensity x 10 -6 LGLVGGSTIDIK++ (586. 85) 3. 5 LVGGSTIDIK+ (1002. 58) 3. 0 GGSTIDIK+ (790. 43) GSTIDIK+ (676. 39) (589. 36) (488. 31) (375. 22) 2. 5 2. 0 1. 5 1. 0 0. 5 0. 0 48 49 50 Retention Time 51 52 (889. 50)

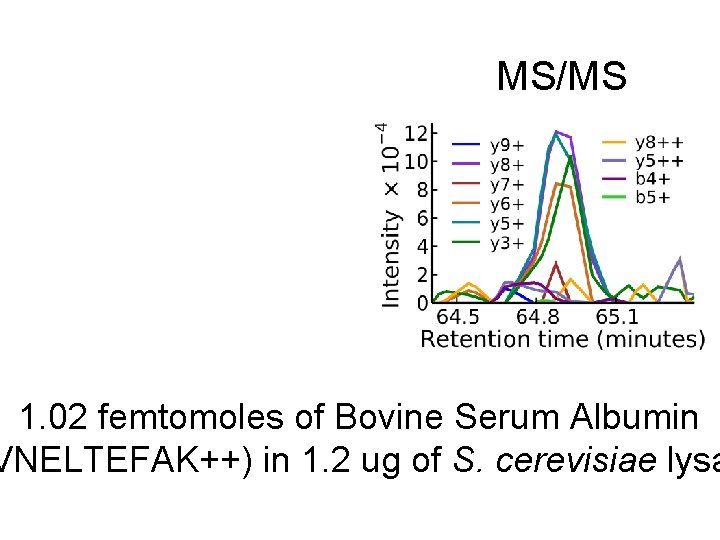
MS/MS 1. 02 femtomoles of Bovine Serum Albumin VNELTEFAK++) in 1. 2 ug of S. cerevisiae lysa

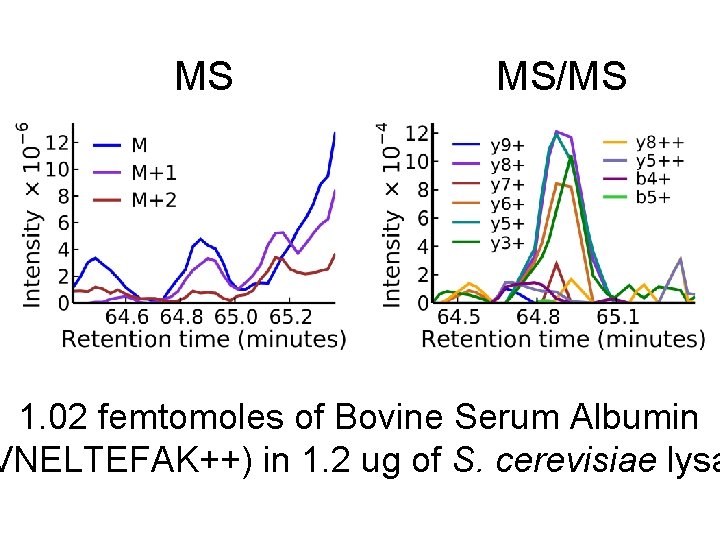
MS MS/MS 1. 02 femtomoles of Bovine Serum Albumin VNELTEFAK++) in 1. 2 ug of S. cerevisiae lysa

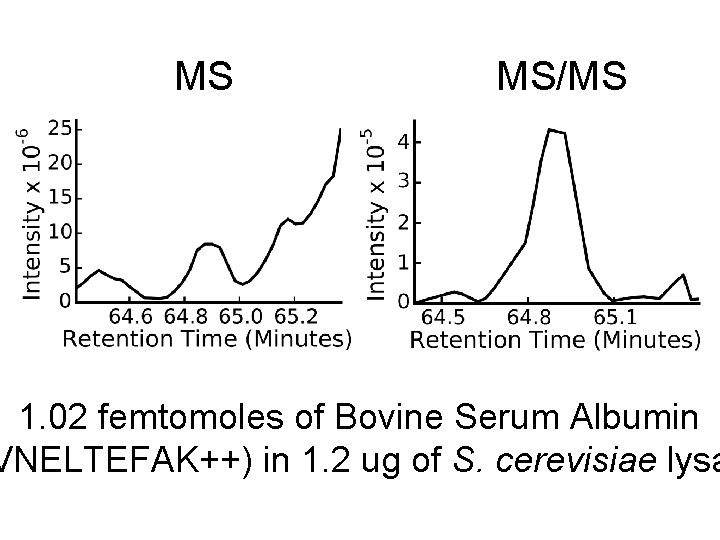
MS MS/MS 1. 02 femtomoles of Bovine Serum Albumin VNELTEFAK++) in 1. 2 ug of S. cerevisiae lysa

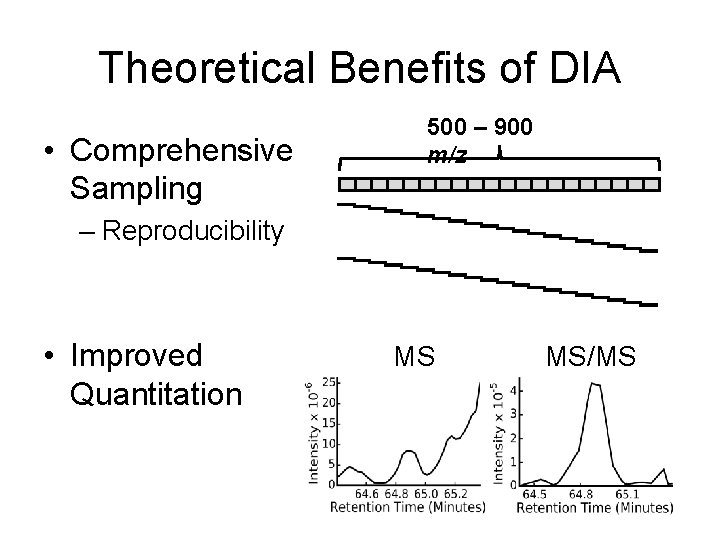
Theoretical Benefits of DIA • Comprehensive Sampling 500 – 900 m/z – Reproducibility • Improved Quantitation MS MS/MS

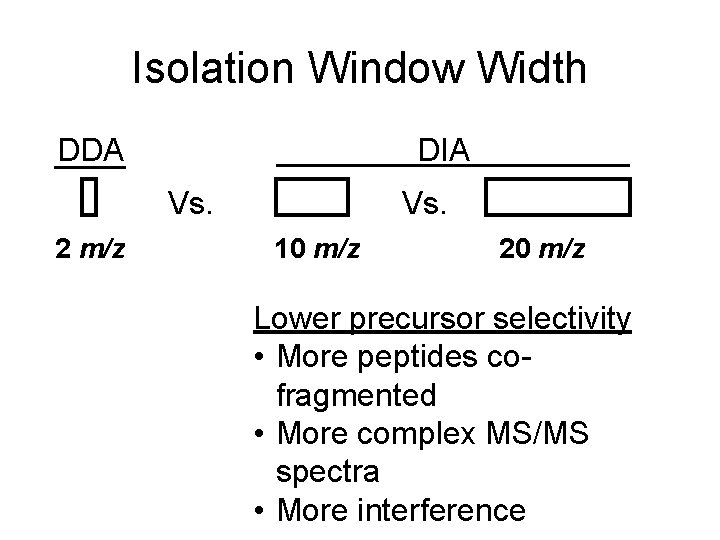
Isolation Window Width DDA DIA Vs. 2 m/z Vs. 10 m/z 20 m/z Lower precursor selectivity • More peptides cofragmented • More complex MS/MS spectra • More interference

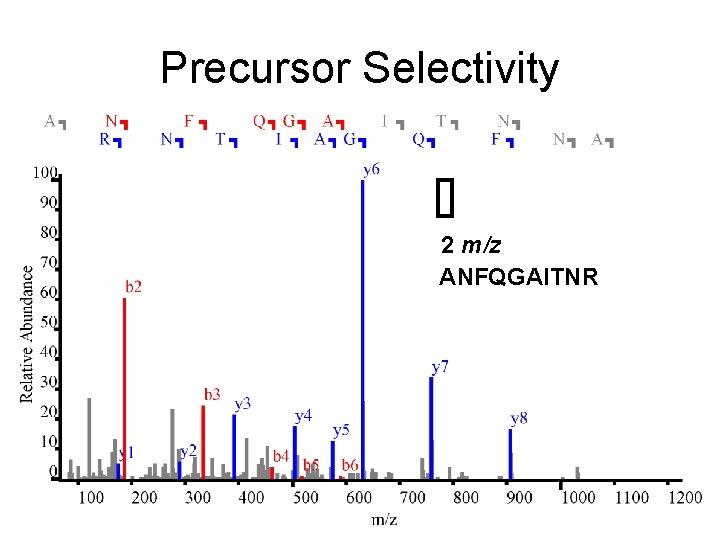
Precursor Selectivity 2 m/z ANFQGAITNR

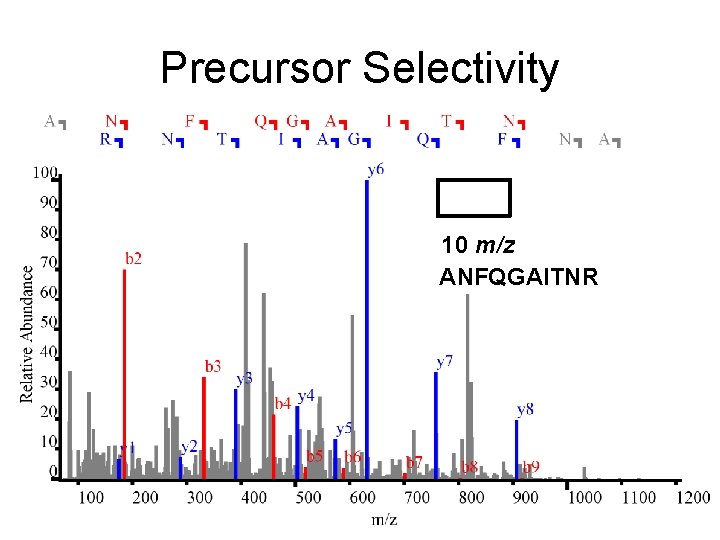
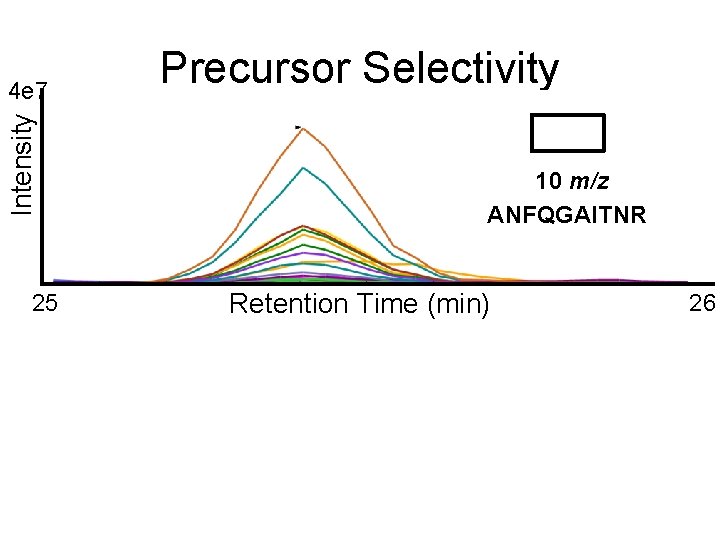
Precursor Selectivity 10 m/z ANFQGAITNR

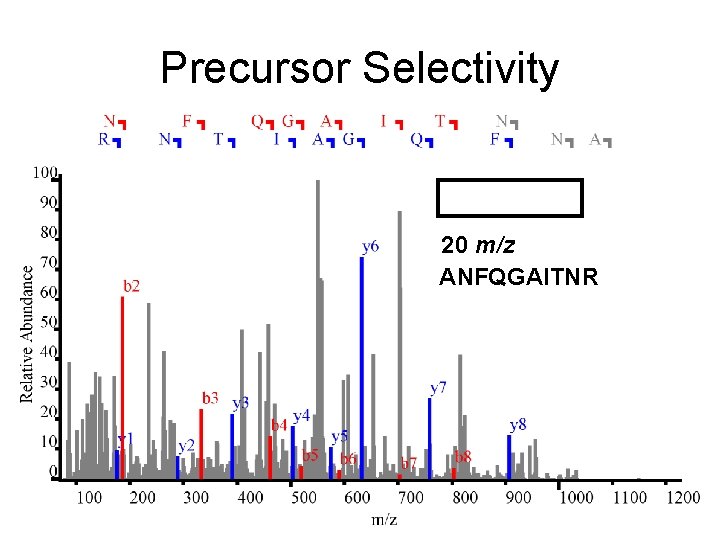
Precursor Selectivity 20 m/z ANFQGAITNR

Intensity 4 e 7 25 Precursor Selectivity 10 m/z ANFQGAITNR Retention Time (min) 26

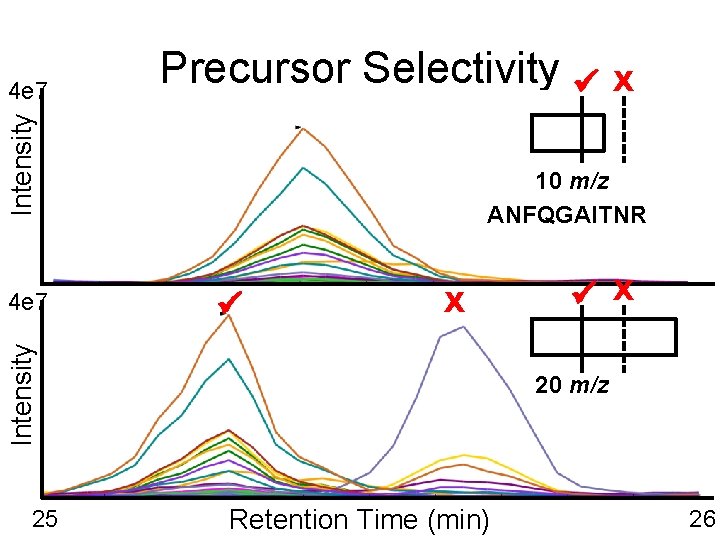
Intensity 4 e 7 Precursor Selectivity 10 m/z ANFQGAITNR X Intensity 4 e 7 25 X X 20 m/z Retention Time (min) 26

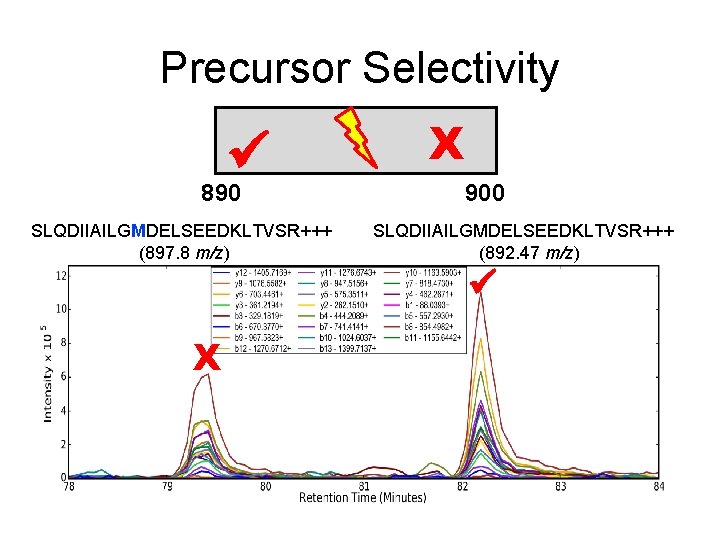
Precursor Selectivity 890 SLQDIIAILGMDELSEEDKLTVSR+++ (897. 8 m/z) X 900 SLQDIIAILGMDELSEEDKLTVSR+++ (892. 47 m/z) X

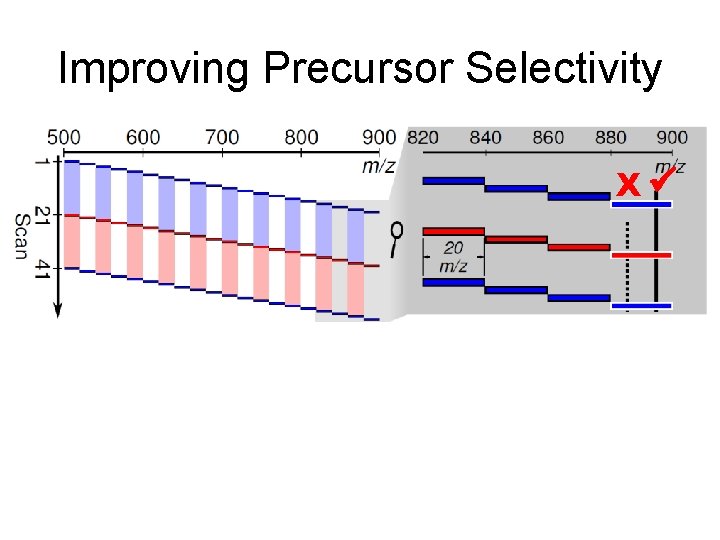
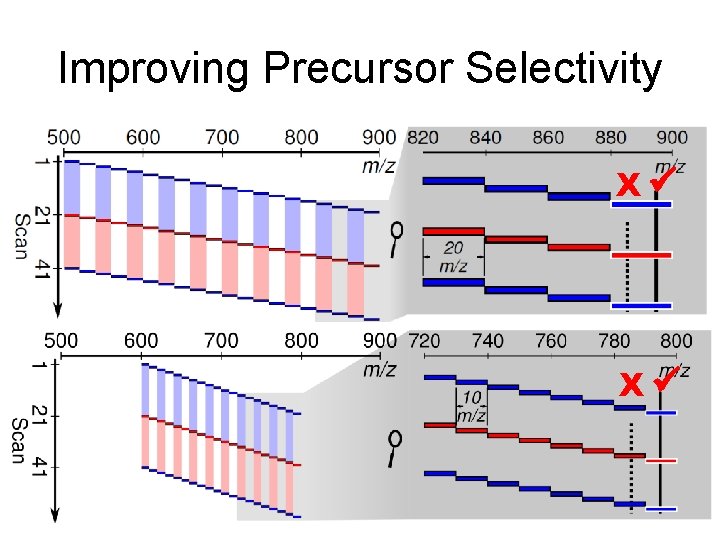
Improving Precursor Selectivity X

Improving Precursor Selectivity X X

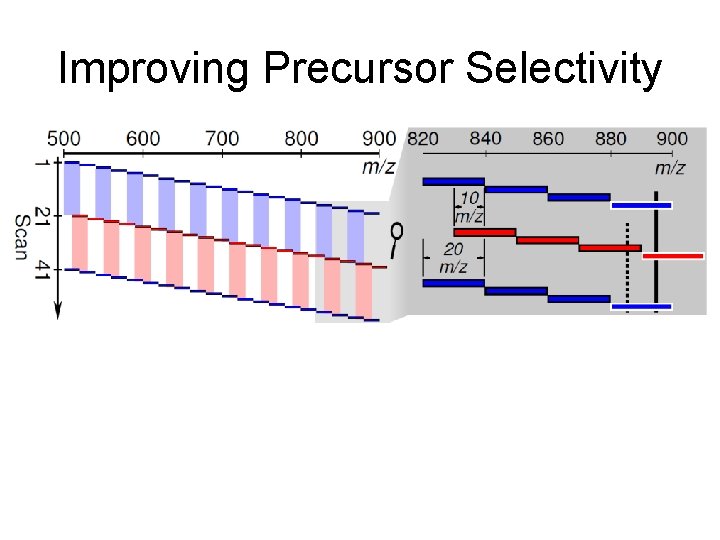
Improving Precursor Selectivity

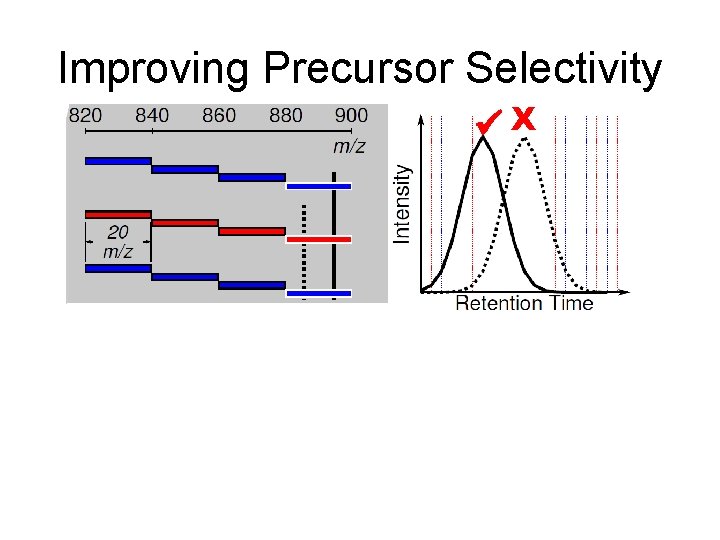
Improving Precursor Selectivity X

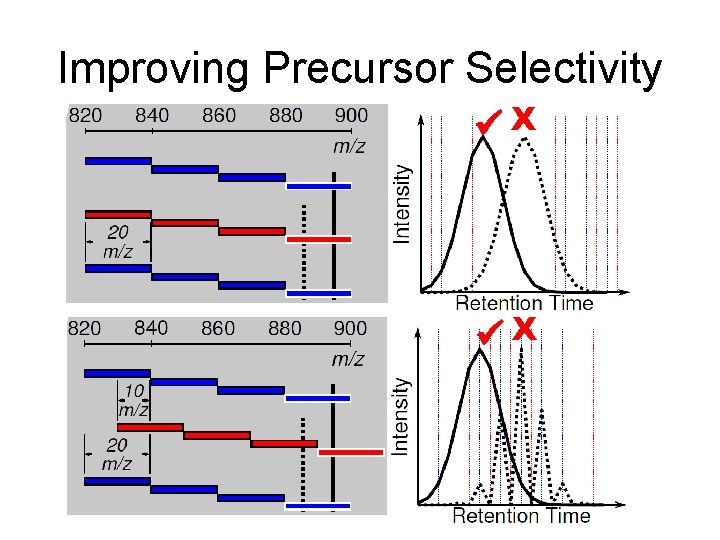
Improving Precursor Selectivity X X

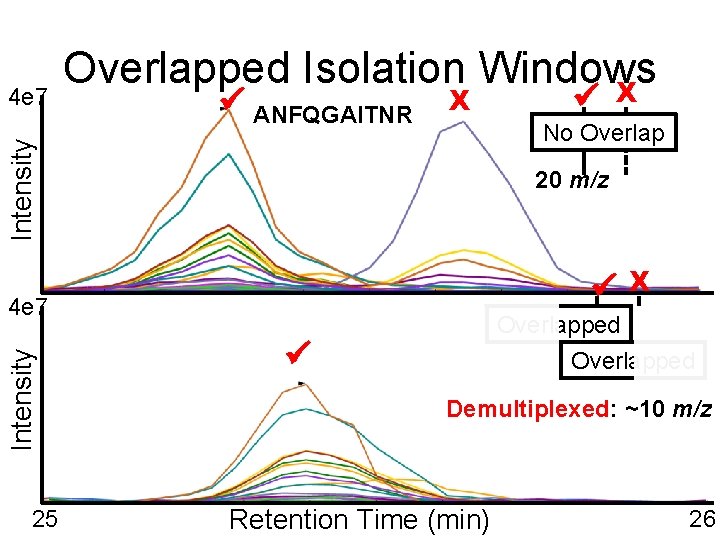
ANFQGAITNR X 20 m/z X Intensity 4 e 7 25 No Overlap Intensity 4 e 7 Overlapped Isolation Windows X Overlapped 20 m/z Demultiplexed: ~10 m/z X Retention Time (min) 26

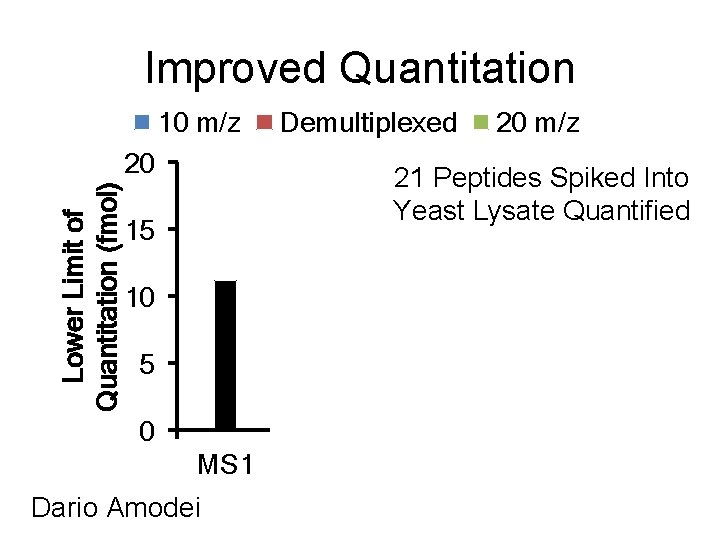
Improved Quantitation 10 m/z Demultiplexed Lower Limit of Quantitation (fmol) 20 20 m/z 21 Peptides Spiked Into Yeast Lysate Quantified 15 10 5 0 MS 1 Dario Amodei All Top 3 Top 5 Top 7 Transitions Integrated

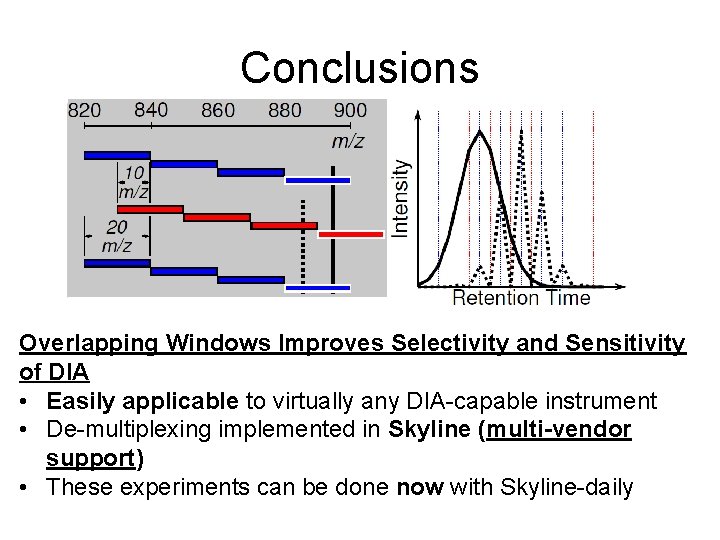
Conclusions Overlapping Windows Improves Selectivity and Sensitivity of DIA • Easily applicable to virtually any DIA-capable instrument • De-multiplexing implemented in Skyline (multi-vendor support) • These experiments can be done now with Skyline-daily

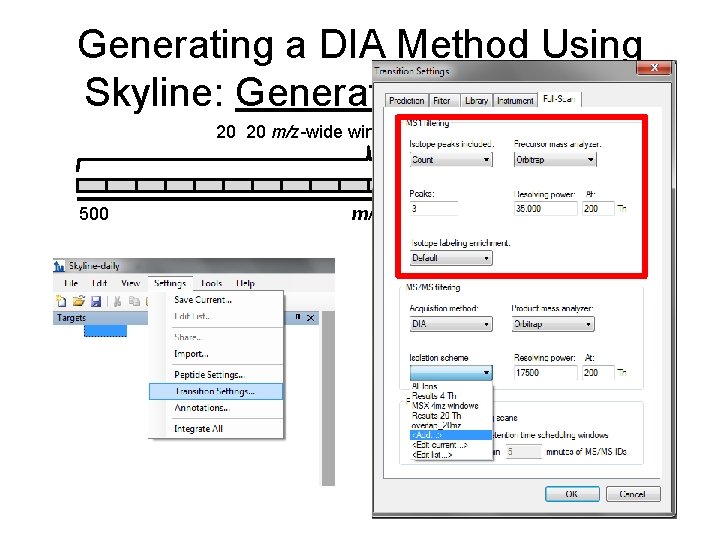
Generating a DIA Method Using Skyline: Generate a Target List 20 20 m/z-wide windows = 400 m/z 500 m/z 900

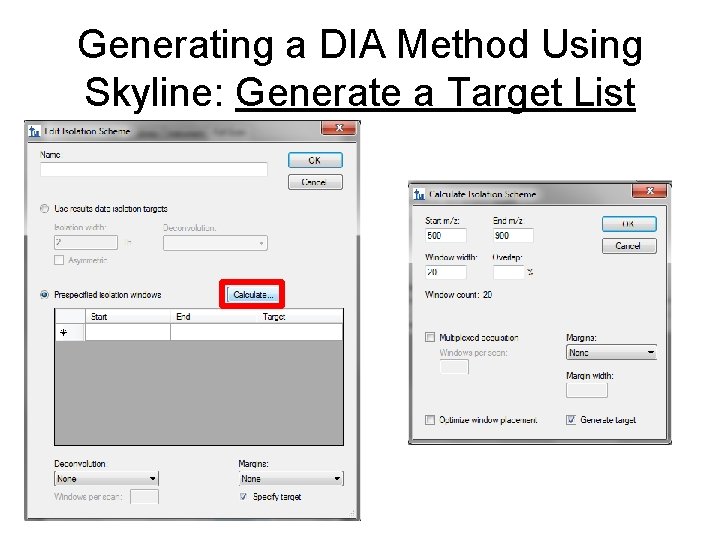
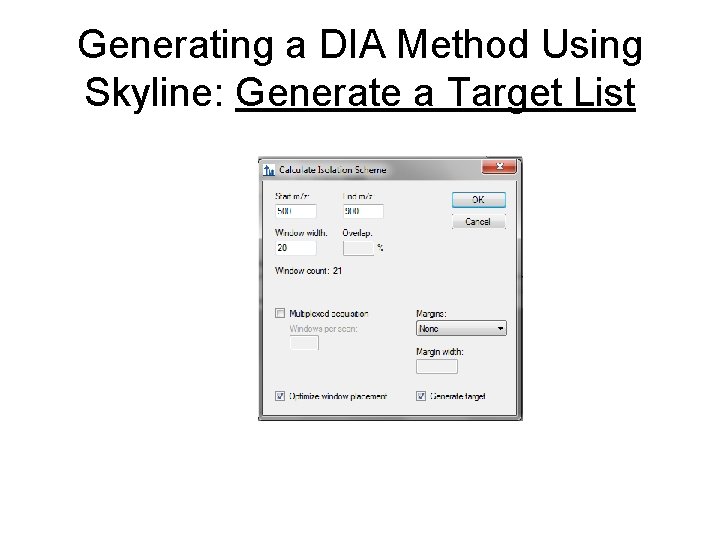
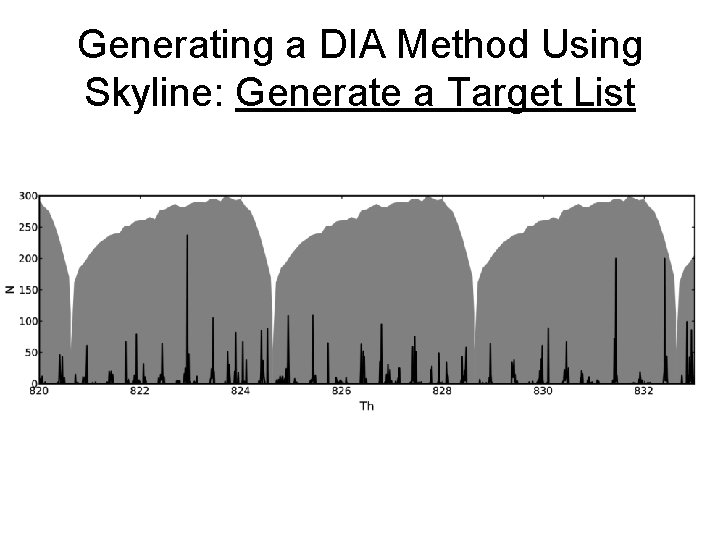
Generating a DIA Method Using Skyline: Generate a Target List

Generating a DIA Method Using Skyline: Generate a Target List

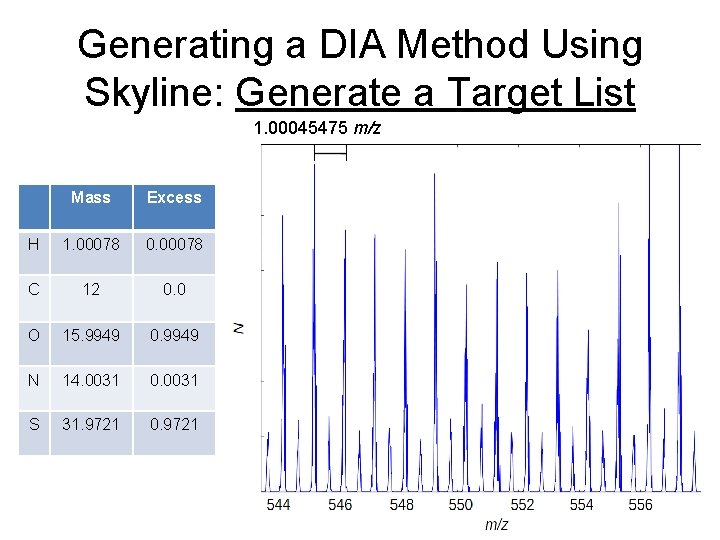
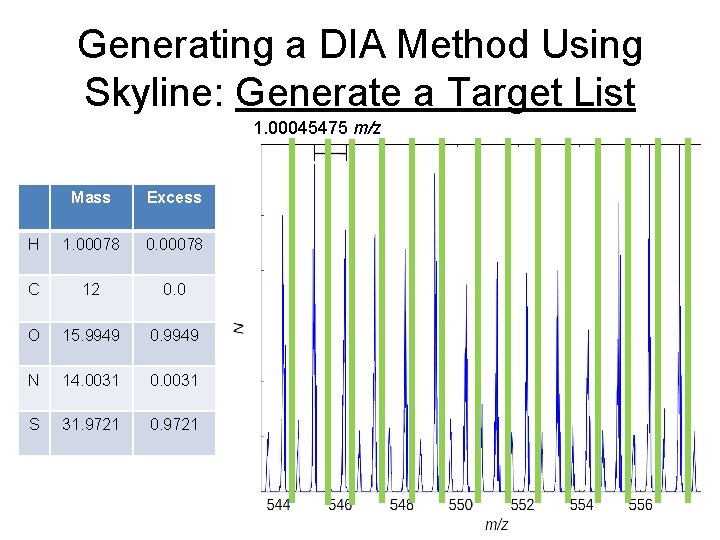
Generating a DIA Method Using Skyline: Generate a Target List 1. 00045475 m/z Mass Excess H 1. 00078 0. 00078 C 12 0. 0 O 15. 9949 0. 9949 N 14. 0031 0. 0031 S 31. 9721 0. 9721

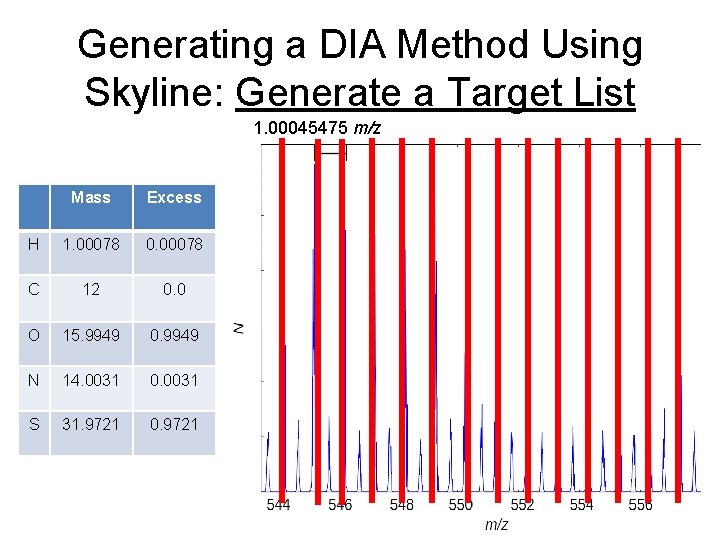
Generating a DIA Method Using Skyline: Generate a Target List 1. 00045475 m/z Mass Excess H 1. 00078 0. 00078 C 12 0. 0 O 15. 9949 0. 9949 N 14. 0031 0. 0031 S 31. 9721 0. 9721

Generating a DIA Method Using Skyline: Generate a Target List 1. 00045475 m/z Mass Excess H 1. 00078 0. 00078 C 12 0. 0 O 15. 9949 0. 9949 N 14. 0031 0. 0031 S 31. 9721 0. 9721

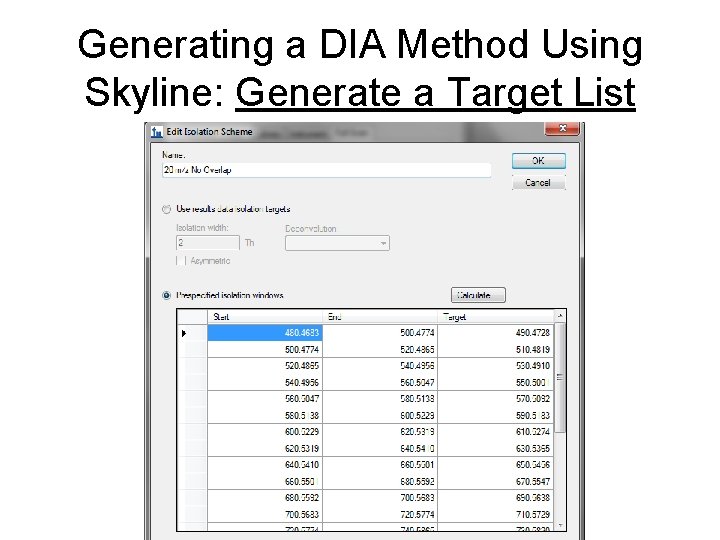
Generating a DIA Method Using Skyline: Generate a Target List

Generating a DIA Method Using Skyline: Generate a Target List

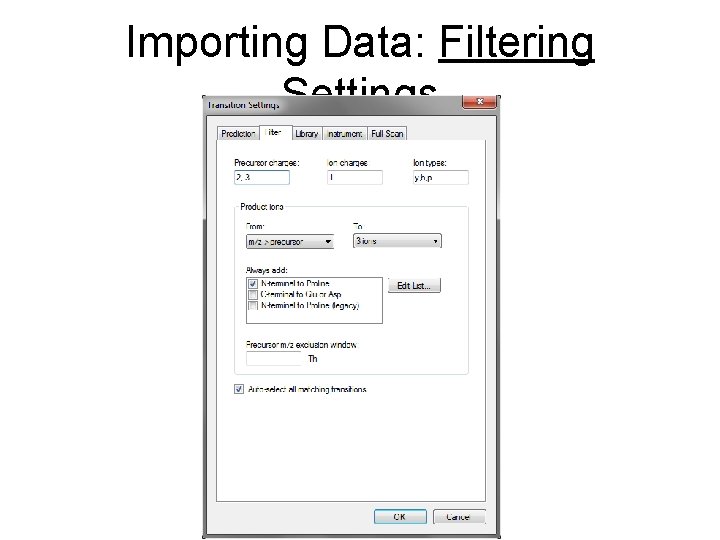
Importing Data: Filtering Settings

Acknowledgements Stanford University of Dario Amodei Washington Parag Mallick Mike Mac. Coss Brendan University Dario’s Poster: Purdue Tuesday June Mac. Lean 11 th Olga Vitek Don Marsh (#512) 10: 30 AM Thermo – 2: 30 PMScientific Jarrett’s Talk: Monday, June Gennifer Markus Kellmann th 10 8: 30 -8: 50 AM Exhibit Hall A Merrihew Andreas Kuehn Richard Johnson Reiko Kiyonami Sonia Ting Yue Xuan & the rest of the lab