Application of Biotechnology Biotechnology for Crop Improvement 1

Application of Biotechnology

Biotechnology for Crop Improvement 1. Pest Resistance Ø A variety of insects, insects mites and nematodes reduce the yield and quality of the crop plants Ø Conventional method use synthetic pesticides severe effects on human health and environment Ø Biotechnology uses an innovative and eco-friendly method to improve pest control management Ø About 40 genes obtained from microorganisms of higher plants and animals used to provide insect resistance in crop plants

Ø The first genes available for genetic engineering of crop plants for pest resistance were Cry genes (popularly known as Bt genes) genes from a bacterium Bacillus thuringiensis. Ø These are specific to particular group of insect pests, and are not harmful to other useful insects like butter flies and silk worms Ø The most notable example is Bt cotton (which contains Cry 1 Ac gene) gene that is resistant to a notorious insect pest Bollworm Ø Transgenic crops with Bt genes (e. g. cotton, rice, maize, potato, tomato, brinjal, cauliflower, cabbage, etc. ) have been developed.

Ø Certain genes from higher plants synthesis of products possessing insecticidal activity. Ø One of the examples is the Cowpea trypsin inhibitor gene (Cp. Ti) which was introduced into tobacco, potato, and oilseed Ø the insecticidal protein was a trypsin inhibitor that was capable of destroying insects belonging to the orders Lepidoptera, Orthaptera etc. Lepidoptera, Orthaptera Ø Cowpea trypsin inhibitor (Cp. Ti) has no effect on mammalian trypsin, hence it is non-toxic to mammals

2. Herbicide Tolerance Ø Weeds are unwanted plants which decrease the crop yields and by competing with crop plants for light, water and nutrients. Ø Several biotechnological strategies for weed control are being used – (a) Over-expression of the target protein to permit normal metabolism despite the presence of herbicide by herbicide integrating multiple copies of the gene or by using a strong promoter, strong promoter (b) Detoxifying the herbicide by using a foreign gene. , and

c) Modification of the target protein of protein herbicide action to insensitive the herbicide by the introduction of a modified gene that encodes for a resistant form of form the enzyme targeted by enzyme targeted the herbicide in weeds Ø Roundup Ready crop plants tolerant to herbicide. Ø Roundup, is already being used commercially.

Glyphosate resistanceØ Glyphosate is a glycine derivative and is a herbicide which is found to be effective against the 76 of the world’s worst weeds Ø It kills the plant by inhibitor of the enzyme 5 -enoylpyruvylshikimate 3 - phosphate synthase (EPSPS) in the shikimic acid pathway. shikimic acid Ø Glyphosate binds more tightly with EPSPS and thus blocks the shikimic acid pathway Ø EPSPS gene was over-expressed in Petunia due to gene amplification. EPSPS gene was isolated from Petunia and gene introduced in to the other plants.

3. Disease Resistance �Disease caused by viruses, fungi and bacteria are among the most serious limitation in crop production. a) Virus resistance � No chemical spray to kill virus � There are several strategies for engineering plants for viral resistance and these utilizes the genes from virus itself (e. g. the viral coat protein gene). � The induction of virus resistance is done by employing virus-encoded genes-virus coat proteins, movement proteins, satellite RNA, antisense RNAs, and ribozymes.

Ø The virus coat protein-mediated approach is the most successful one to provide virus resistance to plants. Ø It was in 1986, transgenic tobacco plants expressing tobacco mosaic virus (TMV) coat protein gene were first developed. These plants exhibited high levels of resistance to TMV. Ø The transgenic plant providing coat protein-mediated resistances to virus are rice, potato, peanut, sugar beet, alfalfa etc.

b) Resistance against Fungal and bacterial infections Ø As a defense strategy against the invading pathogens (fungi and bacteria) the plants accumulate low molecular weight proteins which are collectively known as pathogenesis-related (PR) proteins. Ø Several transgenic crop plants with increased resistance to fungal pathogens are being raised with genes coding for the different compounds e. g. Glucanase enzyme that degrades the cell wall of many fungi. The gene encoding for beta-1, 4 glucanase has been isolated from barley, introduced, and expressed in transgenic tobacco plants. This gene provided good protection against soil-borne fungal pathogen Rhizoctonia solani.

4. Abiotic Stress Resistance Ø The abiotic stresses due to temperature, drought, and salinity are collectively also known as water deficit stresses. Ø The plants produce osmolytes or osmoprotectants to overcome the osmotic stress. Ø The attempts are to increase the production of osmoprotectants in the plants. Ø The biosynthetic pathways for the production of many osmoprotectants have been established and genes coding the key enzymes have been isolated. E. g. Glycine betaine is a cellular osmolyte which is produced by the participation of a number of key enzymes like choline dehydrogenase, choline monooxygenase etc. Ø The choline oxidase gene from Arthrobacter sp. was used to produce transgenic rice with high levels of glycine betaine giving tolerance against water deficit stress.

5. Male Sterility Ø The plants may inherit male sterility either from the nucleus or cytoplasm. Ø In tobacco plants, male sterility are created by introducing a gene coding for an enzyme (barnase) that inhibits pollen formation. This gene is expressed specifically in the tapetal cells of anther using tapetal specific promoter TA 29 to restrict its activity only to the cells involved in pollen production. Ø The restoration of male fertility is done by introducing another gene barstar that suppresses the activity of barnase. Ø By using this approach, transgenic plants of tobacco, cauliflower, cotton, tomato, corn, lettuce etc. with male sterility have been developed.

6. Improvement of Nutritional quality Ø Transgenic crops with improved nutritional quality have already been produced by introducing genes involved in the metabolism of vitamins, minerals and amino acids. Ø A transgenic Arabidopsis thaliana that can produce ten-fold higher vitamin E (alpha-tocopherol) than the native plant Ø In A. thaliana a gene that can finally produce alpha-tocopherol is also present, but is not expressed. This dormant gene was activated by inserting a regulatory gene from a bacterium which resulted in an efficient production of vitamin E. Ø Glycinin is a lysine-rich protein of soybean and the gene encoding glycinin has been introduced into rice and successfully expressed. The transgenic rice plants produced glycinin with high contents of lysine.

Ø Using genetic engineering Prof Potrykus and Dr. Peter Beyer have developed rice which is enriched in pro-vitamin A by introducing three genes involved in the biosynthetic pathway for carotenoid, the precursor for vitamin A. i. psy (phytoene synthase) from daffodil ii. crt. I (phytoene desaturase) from the soil bacterium Erwinia uredovora iii. lyc (lycopene cyclase) gene from daffodil Ø The aim was to help millions of people who suffer from night blindness due to Vitamin A deficiency. The presence of beta-carotene in the rice gives a characteristic yellow /orange colour, hence this pro-vitamin A enriched rice is named as Golden Rice.

A overview of the carotenoid biosynthesis pathway in golden rice Geranyl geranyl di-phosphate Phytoene synthase Phytoene desaturase Lycopene β cyclase β- carotene

7. Delay Fruit Ripening Ø Ethylene is a natural plant hormone associated with the growth, development, ripening and aging of many plants. Ø There are several ways by which scientists can control the ripening process by genetic modification. a) Regulation of Ethylene Production: The amount of ethylene produced can be controlled primarily by “switching off” or decreasing the production of ethylene in the fruit b) Control of Ethylene Perception c) Suppression of Polygalacturonase Activity: The main strategy used was the antisense RNA approach. In the normal tomato plant, the PG gene (for the enzyme polygalacturonase) encodes a normal m. RNA that produces the enzyme polygalacturonase which is involved in the fruit ripening. The complimentary DNA of PG encodes for antisense m. RNA, which is complimentary to normal (sense) m. RNA. The hybridization between the sense and antisnse m. RNAs renders the sense m. RNA ineffective. Ø The most common example is the 'Flavr Savr' transgenic tomatoes, which were commercialized in U. S. A in 1994.

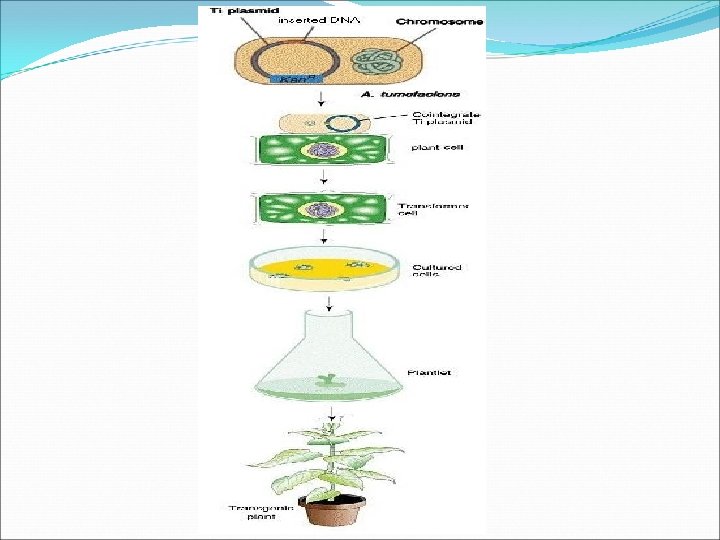
8. Edible Vaccine Ø Crop plants offer cost-effective bioreactors to express antigens which can be used as edible vaccines. Ø The approach is to isolate genes encoding antigenic proteins from the pathogens and then expressing them in plants. Such transgenic plants or their tissues producing antigens can be eaten for vaccination/immunization (edible vaccines). Ø The expression of such antigenic proteins in crops like banana and tomato are useful for immunization of humans since banana and tomato fruits can be eaten raw. Ø The process of making of edible vaccines involves the incorporation of a plasmid carrying the antigen gene and an antibiotic resistance gene, into the bacterial cells e. g. Agrobacterium tumefaciens.

Ø The small pieces of potato leaves are exposed to an antibiotic which can kill the cells that lack the new genes. The surviving cells with altered genes multiply and form a callus. This callus is allowed to grow and subsequently transferred to soil to form a complete plant. In about a few weeks, the plants bear potatoes with antigen vaccines Ø Another strategy to produce a plant-based vaccine, is to infect the plants with recombinant virus carrying the desired antigen that is fused to viral coat protein. The infected plants are reported to produce the desired fusion protein in large amounts. Ø In 1990, the first report of the production of edible vaccine (a surface protein from Streptococcus) in tobacco. Ø The first clinical trials in humans, using a plant derived vaccine were conducted in 1997.

9. N 2 Fixation Ø Molecular nitrogen in the atmosphere is converted into biologically converted forms by nitrogen fixing microorganisms e. g. Rhizobium. Ø Gene involved in nitrogen fixation. I. Nod genes- plant root produce various type of flavonoids that stimulates the release of Nod factor by Rhizobium for nodulation. The eight nod genes present in the Rhizobium bacteria. II. Nif gene – Nif genes are involved in nitrogen fixation that are isolate from k. pneumoniae and transfer into E. coli.

10. Photosynthesis Ø The unicellular cyanobacterium Synechocystis sp. was photosynthetic organism whose genome was completely sequenced. Ø Cyanobacteria developed the biotechnology of photosynthesis. Ø It was a remarkable achievement that the necessary and vital carbon could then be seized directly from the dense CO 2 concentration in the atmosphere, then combined with water to be converted into usable suger food.

Recombinant DNA (r. DNA) is a form of artificial DNA that is created by combining two or more sequences that would not normally occur together. Or Genetically engineered DNA prepared by splicing genes from one species into the cells of a host organism of a different species. Such DNA becomes part of the host's genetic makeup and is replicated

Recombinant DNA technology A series of procedures that are used to join together (recombine) DNA segments. A recombinant DNA molecule is constructed from segments of two or more different DNA molecules. Under certain conditions, a recombinant DNA molecule can enter a cell and replicate there, either on its own or after it has been integrated into a chromosome Department of Biotechnology

Applications of recombinant DNA technology 1. Recombinant chymosin 2. Recombinant human insulin 3. Recombinant human growth hormone somatotropin) 4. Recombinant blood clotting factor VIII 5. Recombinant hepatitis B vaccine 6. Diagnosis of infection with HIV 7. Golden rice 8. Herbicide-resistant crops 9. Insect-resistant crops Department of Biotechnology (HGH,

Vector is a vehicle used to transfer genetic material to a target cell. In molecular cloning, a vector is……. q. A DNA molecule used as a vehicle q. A vector containing foreign gene/DNA is termed recombinant DNA q Vector transfers the recombinant DNA into another cell q. It can be replicated and/or expressed Department of Biotechnology
Vector Department of Biotechnology

. The major types of vectors are- • Plasmids • Cosmids • Viral vectors Department of Biotechnology

Plasmids Vector ØDouble-stranded ØCircular DNA sequences ØCapable of automatically replicating in a host cell ØMultiple cloning site" Department of Biotechnology
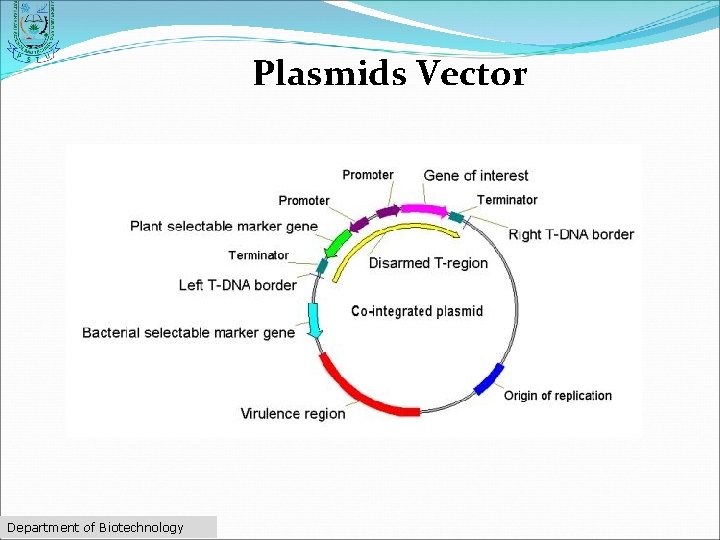
Plasmids Vector Department of Biotechnology
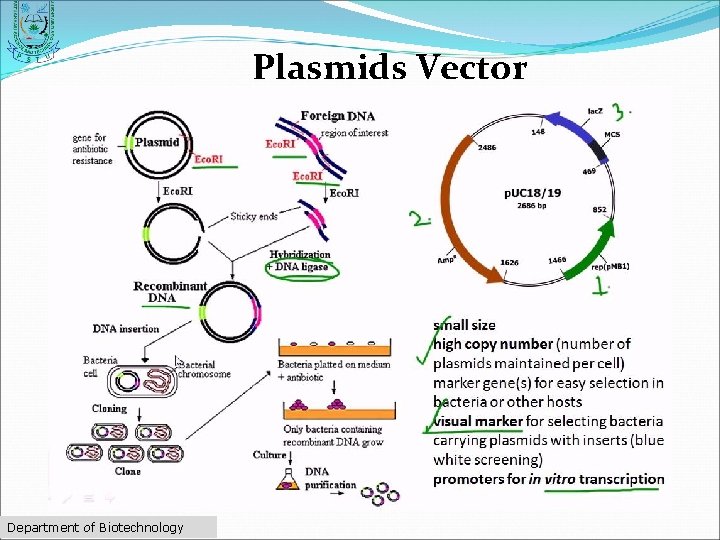
Plasmids Vector Department of Biotechnology
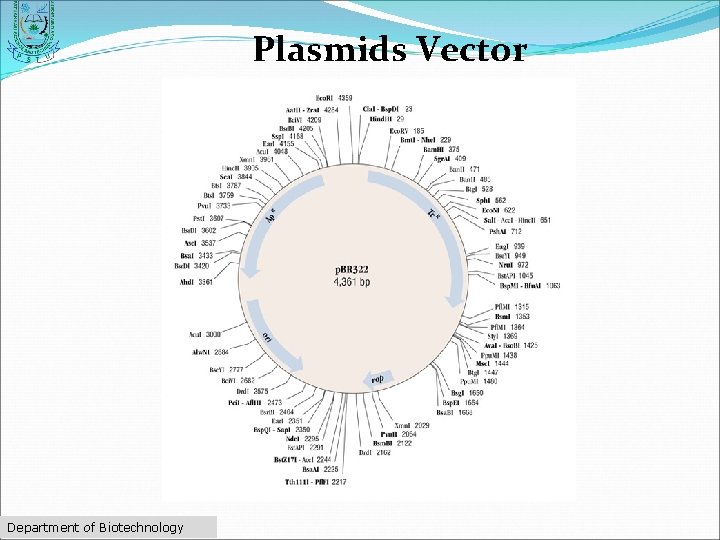
Plasmids Vector Department of Biotechnology

Cosmid üFirst described by Collins and Hohn in 1978 ü Hybrid plasmid (often used as a cloning vector) üContains a Lambda phage cos sequence. üCosmids' (cos sites + plasmid = cosmid) üCosmids can be used to build genomic libraries Department of Biotechnology
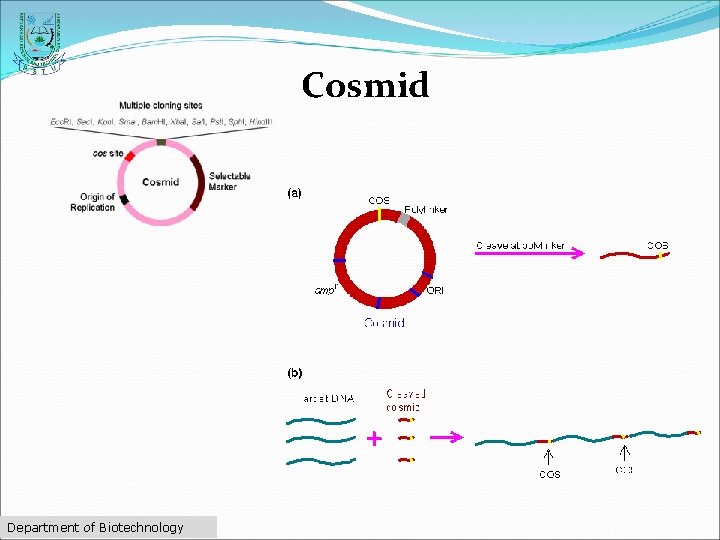
Cosmid Department of Biotechnology
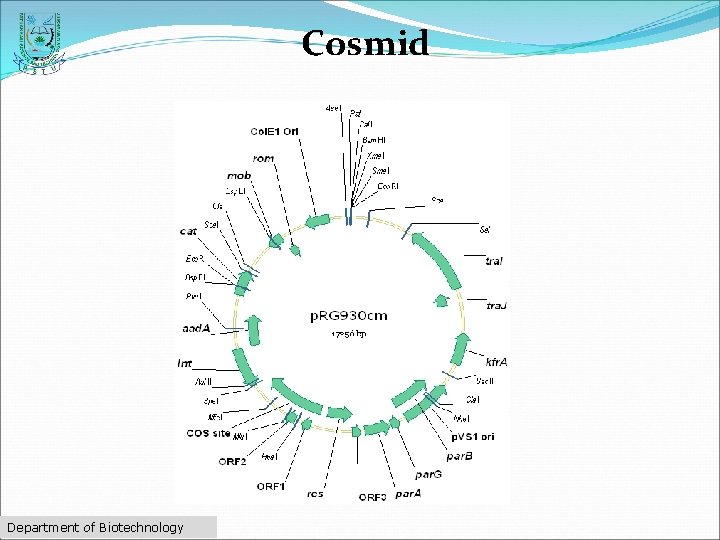
Cosmid Department of Biotechnology

Viral vectors q. Genetically-engineered viruses carrying modified viral DNA or RNA q. Viral vectors frequently are lacking infectious sequences q. Viral vectors are often designed for permanent incorporation of incorporation the insert into the host genome. Department of Biotechnology

Common features of modern vectors i. Origin of replication, ii. Promoter, iii. Multiple cloning site, iv. Genetic markers, v. Antibiotic resistance, vi. Epitope, vii. Reporter genes, viii. Targeting sequence and ix. Protein purification tags. Department of Biotechnology
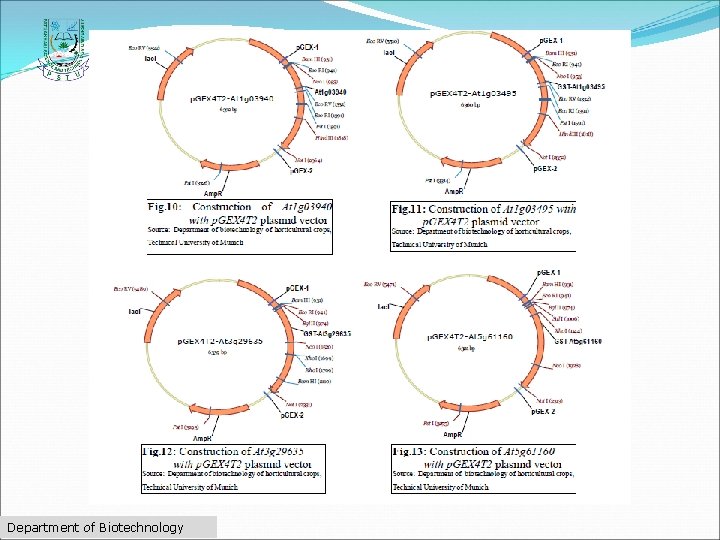
Department of Biotechnology

GENE ISOLATION METHODS OF GENE TRANSFER

Gene Isolation 1. Breaking the cells, cells commonly referred to as cell disruption or cell lysis, to expose the DNA. This is commonly achieved by grinding or sonicting the sample 2. Removing membrane lipids by adding a detergent. 3. Removing proteins by adding a protease. 4. Precipitating the DNA with an alcohol — usually icecold ethanol or isopropanol. 5. Targeted gene is then cut by specific endunuclease enzyme. 6. Preserving the gene under -200 C for further use.

Methods of gene transfer 1. Indirect gene transfer: §Vector-mediated (Agrobacterium tumefaciens) 2. Direct gene transfer §Microinjection § Electroporation § Particle gun/Particle bombardment §Liposome mediated gene transfer or Lipofection §Chemical mediated gene transfer § Conjunction Department of Biotechnology
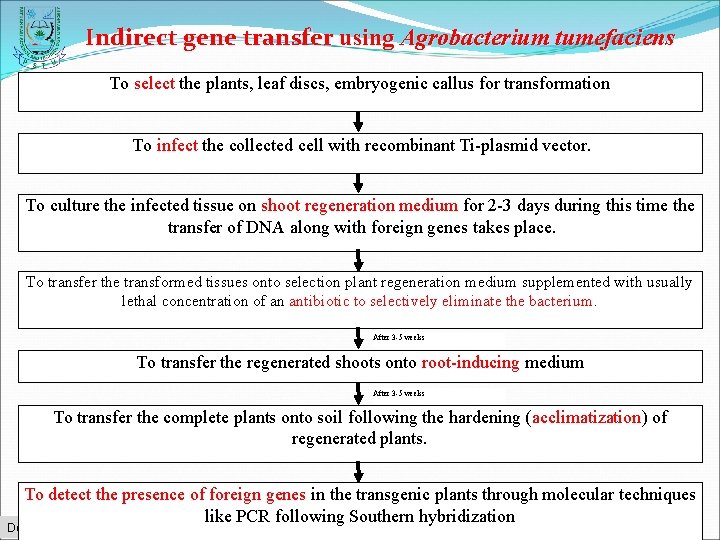
Indirect gene transfer using Agrobacterium tumefaciens To select the plants, leaf discs, embryogenic callus for transformation To infect the collected cell with recombinant Ti-plasmid vector. To culture the infected tissue on shoot regeneration medium for 2 -3 days during this time the transfer of DNA along with foreign genes takes place. To transfer the transformed tissues onto selection plant regeneration medium supplemented with usually lethal concentration of an antibiotic to selectively eliminate the bacterium After 3 -5 weeks To transfer the regenerated shoots onto root-inducing medium After 3 -5 weeks To transfer the complete plants onto soil following the hardening (acclimatization) of regenerated plants. To detect the presence of foreign genes in the transgenic plants through molecular techniques like PCR following Southern hybridization Department of Biotechnology

Direct gene Transfer Microinjection Ø Microinjection is a technique of delivering foreign DNA into a living cell (a cell, egg, oocyte, embryos of animals) through a glass micropipette. ØThe tip of the pipette attains to about 0. 5 mm diameter which resembles an injection needle. ØThe tip of the micropipette is injected through the membrane of the cell. Contents of the needle are delivered into the cytoplasm and the empty needle is taken out. Department of Biotechnology
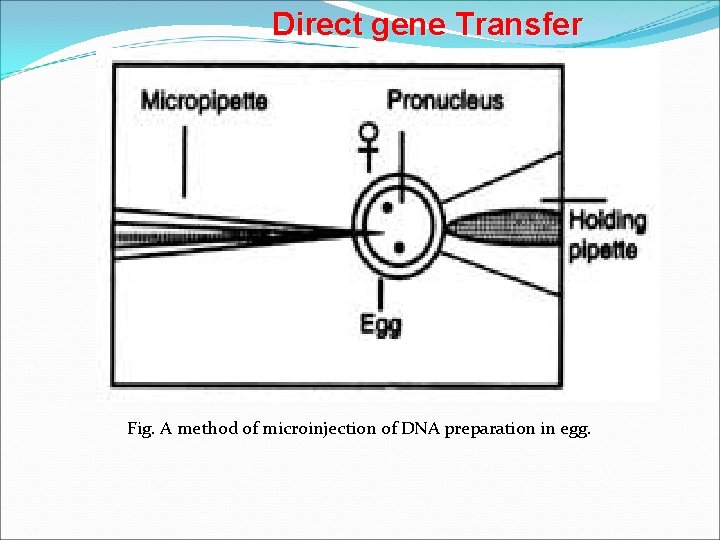
Direct gene Transfer Fig. A method of microinjection of DNA preparation in egg.

Direct gene Transfer Gene gun/Particle bombardment ØIn this method, the foreign DNA containing the genes to be transferred is coated onto the surface of minute gold or tungsten particles (13 micrometers) and bombarded onto the target tissue or cells using a particle gun (also called as gene gun/shot gun/microprojectile gun). ØSelectable markers are used to identify the cells that take up the transgene. ØTwo types of plant tissue are commonly used for particle bombardment- Primary explants and the proliferating embryonic tissues Department of Biotechnology
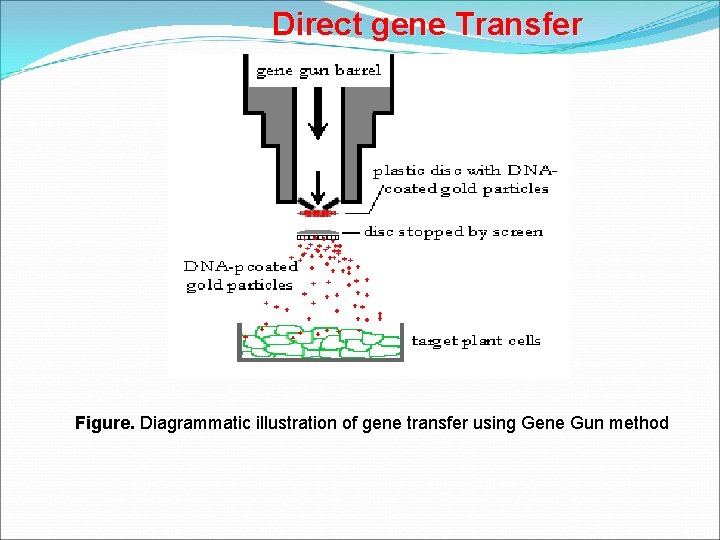
Direct gene Transfer Figure. Diagrammatic illustration of gene transfer using Gene Gun method

Direct gene Transfer Electroporation ØElectroporation involves a pulse of high voltage applied to protoplasts/ cells/ tissues to make transient (temporary) pores in the plasma membrane which facilitates the uptake of foreign DNA. ØThe cells are placed in a solution containing DNA and subjected to electrical shocks to cause holes in the membranes. ØThe foreign DNA fragments enter through the holes into the cytoplasm and then to nucleus. Department of Biotechnology
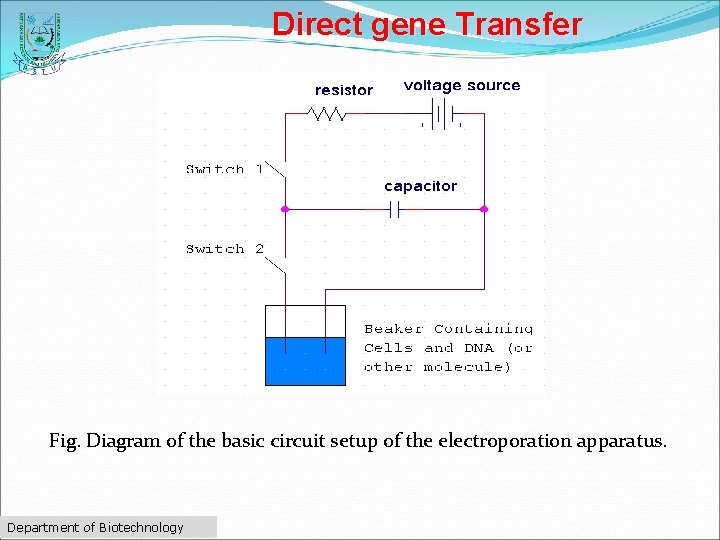
Direct gene Transfer Fig. Diagram of the basic circuit setup of the electroporation apparatus. Department of Biotechnology

Direct gene Transfer Liposome mediated gene transfer or Lipofection: Ø Liposomes are circular lipid molecules with an aqueous interior that can carry nucleic acids. Ø Liposomes encapsulate the DNA fragments and then adhere to the cell membranes and fuse with them to transfer DNA fragments. ØThus, the DNA enters the cell and then to the nucleus. ØLipofection is a very efficient technique used to transfer genes in bacterial, animal and plant cells. Department of Biotechnology

Direct gene Transfer Chemical mediated gene transfer: Chemicals like polyethylene glycol (PEG) and dextran sulphate induce DNA uptake into plant protoplasts. Calcium phosphate is also used to transfer DNA into cultured cells. Department of Biotechnology

Direct gene Transfer Conjunction It is a natural microbial recombination process and is used as a method for gene transfer. In conjuction, two live bacteria come together and the single stranded DNA is transferred via cytoplasmic bridges from the donor bacteria to the recipient bacteria. Department of Biotechnology

Thanks For The Attention Department of Biotechnology

Restriction Enzymes

Enzymes �Enzymes are proteins �biological catalysts help drive biochemical reactions �Enzyme names end with an ase (eg. , endonuclease)

Restriction Enzymes A restriction enzyme (or restriction endonuclease) is an enzyme that cuts DNA at or near specific recognition nucleotide sequences known as restriction sites.

Nomenclature �Smith and Nathans (1973) proposed enzyme naming scheme �three-letter acronym for each enzyme derived from the source organism �First letter from genus �Next two letters represent species �Additional letter or number represent the strain or serotypes �For example. the enzyme Eco. RI was isolated from Escherichia coli serotype R and numeric number I.

Nomenclature Derivation of the Eco. RI name Abbreviation Meaning E Escherichia co coli R RY 13 I First identified Description genus species strain order of identification in the bacterium

Few Restriction Enzymes Target sequence Enzyme Organism from which derived (cut at *) 5' -->3' Bam. HI Bacillus amyloliquefaciens G* G A T C C Eco. RI Escherichia coli RY 13 G* A A T T C Hind. III Haemophilus inflenzae Rd A* A G C T T Mbo. I Moraxella bovis *G A T C Pst. I Providencia stuartii CTGCA*G Sma. I Serratia marcescens CCC*GGG Taq. I Thermophilus aquaticus T*CGA Xma. I Xanthamonas malvacearum C*CCGGG

Few Restriction Enzymes Target sequence Enzyme Organism from which derived (cut at *) 5' -->3' Bam. HI Bacillus amyloliquefaciens G* G A T C C Eco. RI Escherichia coli RY 13 G* A A T T C Hind. III Haemophilus inflenzae Rd A* A G C T T Mbo. I Moraxella bovis *G A T C Pst. I Providencia stuartii CTGCA*G Sma. I Serratia marcescens CCC*GGG Taq. I Thermophilus aquaticus T*CGA Xma. I Xanthamonas malvacearum C*CCGGG

Classification �Synonymous to Restriction Endonuclease � Endonuclease: Cut DNA from inside �Highly heterogeneous �Evolved independently rather than diverging form a common ancestor �Broadly classified into four Types

R-M System Restriction-modification (R-M) system �Endonuclease activity: cuts foreign DNA at the recognition site �Methyltransferase activity: protects host DNA from cleavage by the restriction enzyme. �Methyleate one of the bases in each strand � Restriction enzyme and its cognate modification system constitute the R-M system

Protection of Self DNA �Bacteria protect their self DNA from restriction digestion by methylation of its recognition site. �Methylation is adding a methyl group (CH 3) to DNA. �Restriction enzymes are classified based on recognition sequence and methylation pattern.

Type I �Multi-subunit proteins � Function as a single protein complex � Contain � two R (restriction) subunits, � two M (methylation) subunits and � one S (specificity) subunit � Cleave DNA at random length from recognition site �Type I enzymes cleave at sites remote from recognition site; require both ATP and S-adenosyl-L-methionine to function; multifunctional protein with both restriction and methylase activities.

Type III � Large enzymes � Combination restriction-and-modification � Cleave outside of their recognition sequences � Require two recognition sequences in opposite orientations within the same DNA molecule � No commercial use or availability � Type III enzymes cleave at sites a short distance from recognition site; require ATP (but do not hydrolyse it); S-adenosyl-L-methionine stimulates reaction but is not required; exist as part of a complex with a modification methylase.

Type IV �Cleave only modified DNA (methylated, hydroxymethylated and glucosyl-hydroxymethylated bases). �Recognition sequences have not been well defined �Cleavage takes place ~30 bp away from one of the sites. �Sequence similarity suggests many such systems in other bacteria and archaea. �Type IV enzymes target modified DNA, e. g. methylated, hydroxymethylated and glucosyl-hydroxymethylated DNA.

Type II � Most useful for gene analysis and cloning �More than 3500 REs � Recognize 4 -8 bp sequences � Need Mg 2+ as cofactor � Cut in close proximity of the recognition site � Homodimers �ATP hydrolysis is not required �Type II enzymes cleave within or at short specific distances from recognition site; most require magnesium; single function (restriction) enzymes independent of methylase.

Thanks For The Attention Department of Biotechnology
- Slides: 66