Antimicrobial drug resistance Prof Aparna Mane Subject Microbiology

Antimicrobial drug resistance Prof. Aparna Mane Subject: Microbiology

DEFINITION Drug resistance is the reduction in effectiveness of a medication such as an antimicrobial or an antineoplastic in treating a disease or condition. Prof. Aparna Mane Subject: Microbiology

How does it occur? To survive, germs are constantly finding new defense strategies, called “resistance mechanisms, ” to avoid the effects of antibiotics. Bacteria develop resistance mechanisms by using instructions provided by their DNA. Often, resistance genes are found within plasmids, small pieces of DNA that carry genetic instructions from one germ to another. This means that some bacteria can share their DNA and make other germs become resistant. Prof. Aparna Mane Subject: Microbiology

How does it spread? Antibiotics are given to animals. Antibiotic resistance may develop. Resistant bacteria spread to humans and other animals through poorly prepared food, close proximity and poor hygiene. Resistant bacteria spread to the environment and food through water contaminated by faeces or through wildlife. Research into how antibiotic resistance is spread in Australia is ongoing and new information is continuing to emerge. What is happening in other countries may not be applicable to Australia. Antibiotics are given to a person in hospital or the community. Antibiotic resistance may develop. Resistant bacteria spread to other people through poor hygiene and close proximity. Resistant bacteria spread to humans and other animals through the environment (water, soil, air). Antibiotic resistance occurs when bacteria change to protect themselves from an antibiotic. The more we use antibiotics, the more chance bacteria have to become resistant to them. Prof. Aparna Mane Subject: Microbiology
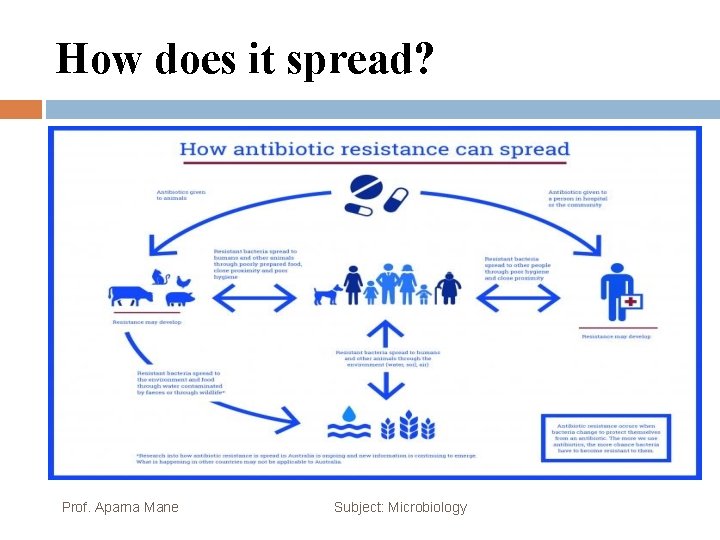
How does it spread? Prof. Aparna Mane Subject: Microbiology

Drug Resistance Mechanisms Efflux pumps Decreased permealibility against antibiotics Inactivating Enzymes Use of alternative enzymes Target alterations Prof. Aparna Mane Subject: Microbiology

EFFLUX PUMPS Efflux pumps are transport proteins involved in the extrusion of toxic substrates (including virtually all classes of clinically relevant antibiotics) from within cells into the external environment. Pumps may be specific for one substrate or may transport a range of structurally dissimilar compounds (including antibiotics of multiple classes); such pumps can be associated with multiple drug resistance (MDR). In the prokaryotic kingdom there are five major families of efflux transporter: 2 MF (major facilitator), MATE (multidrug and toxic efflux), RND (resistance-nodulation-division), SMR (small multidrug resistance) and ABC (ATP binding cassette). All these systems utilize the proton motive force as an energy source, 3 apart from the ABC family, which utilizes ATP hydrolysis to drive the export of substrates. The broad substrate range of efflux systems is of concern, as often over-expression of a pump will result in resistance to antibiotics of more than one class as well as some dyes, detergents and disinfectants (including some commonly used biocides). Cross-resistance is also a problem; exposure to any one agent that belongs to the substrate profile of a pump would favour over-expression of that pump and consequent cross-resistance to all other substrates of the pump. Prof. Aparna Mane Subject: Microbiology
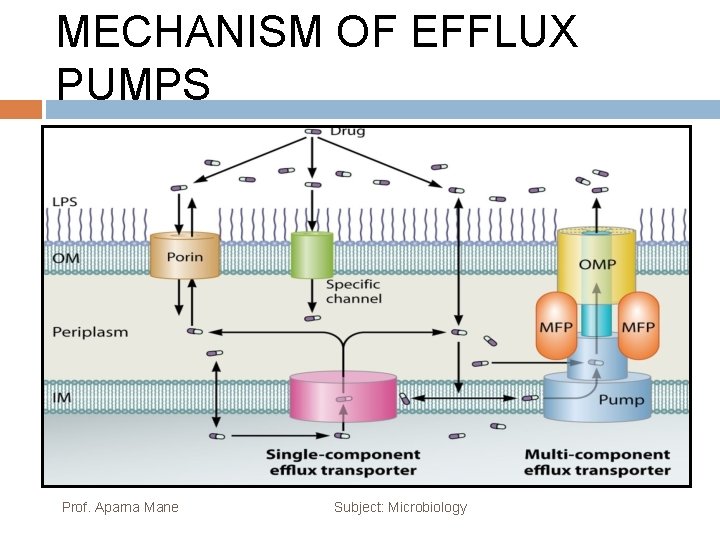
MECHANISM OF EFFLUX PUMPS Prof. Aparna Mane Subject: Microbiology

DECREASED PERMEABILITY AGAINST ANTIBIOTICS The outer membrane (OM) of Gram-negative bacteria performs the crucial role of providing an extra layer of protection to the organism without compromising the exchange of material required for sustaining life. By combining a highly hydrophobic lipid bilayer with pore-forming proteins of specific size-exclusion properties, the OM acts as a selective barrier. The permeability properties of this barrier, therefore, have a major impact on the susceptibility of the microorganism to antibiotics, which, to date, are essentially targeted at intracellular processes. There are essentially two pathways that antibiotics can take through the outer membrane: a lipid-mediated pathway for hydrophobic antibiotics, and general diffusion porins for hydrophilic antibiotics. The lipid and protein compositions of the outer membrane have a strong impact on the sensitivity of bacteria to many types of antibiotics, and drug resistance involving modifications of these macromolecules is common. Prof. Aparna Mane Subject: Microbiology
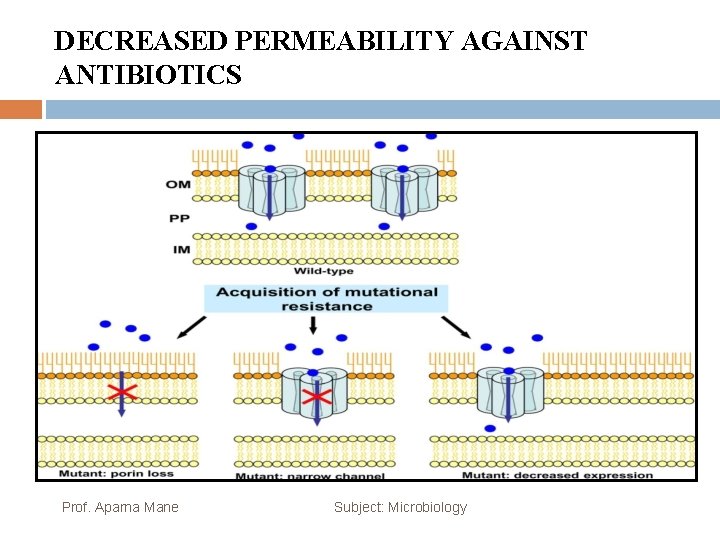
DECREASED PERMEABILITY AGAINST ANTIBIOTICS Prof. Aparna Mane Subject: Microbiology
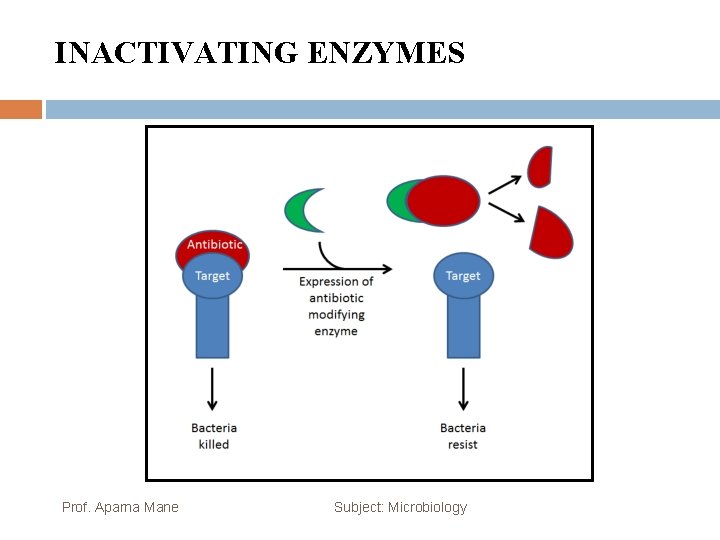
INACTIVATING ENZYMES Prof. Aparna Mane Subject: Microbiology

INACTIVATING ENZYMES Resistance genes may code for enzymes that chemically modify an antimicrobial, thereby inactivating it, or destroy an antimicrobial through hydrolysis. Resistance to many types of antimicrobials occurs through this mechanism. For example, aminoglycoside resistance can occur through enzymatic transfer of chemical groups to the drug molecule, impairing the binding of the drug to its bacterial target. For β-lactams, bacterial resistance can involve the enzymatic hydrolysis of the β-lactam bond within the β-lactam ring of the drug molecule. Once the β-lactam bond is broken, the drug loses its antibacterial activity. This mechanism of resistance is mediated by β-lactamases, which are the most common mechanism of β-lactam resistance. Inactivation of rifampin commonly occurs through glycosylation, phosphorylation, or adenosine diphosphate (ADP) ribosylation, and resistance to macrolides and lincosamides can also occur due to enzymatic inactivation of the drug or modification. Prof. Aparna Mane Subject: Microbiology
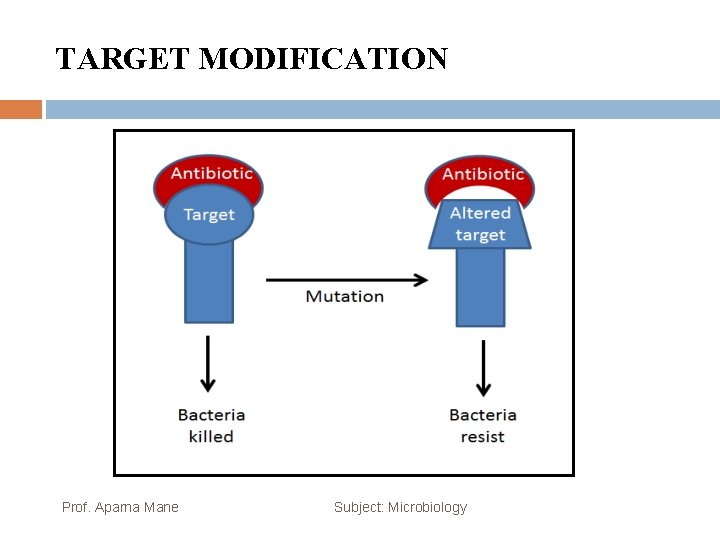
TARGET MODIFICATION Prof. Aparna Mane Subject: Microbiology

TARGET MODIFICATION Because antimicrobial drugs have very specific targets, structural changes to those targets can prevent drug binding, rendering the drug ineffective. Through spontaneous mutations in the genes encoding antibacterial drug targets, bacteria have an evolutionary advantage that allows them to develop resistance to drugs. This mechanism of resistance development is quite common. Examples of this resistance strategy include alterations in ribosome subunits, providing resistance to macrolides, tetracyclines, and aminoglycosides; lipopolysaccharide (LPS) structure, providing resistance to polymyxins; RNA polymerase, providing resistance to rifampin; DNA gyrase, providing resistance to fluoroquinolones; metabolic enzymes, providing resistance to sulfa drugs, sulfones, and trimethoprim; and peptidoglycan subunit peptide chains, providing resistance to glycopeptides. Prof. Aparna Mane Subject: Microbiology
- Slides: 14