Antibiotic resistance criteria Komal Pareek 1 Key Terms

Antibiotic resistance criteria Komal Pareek 1

Key Terms • Antibiotic = A drug that kills or inhibits the growth of microorganisms • Resistant = Somewhat arbitrary designation that implies that an antimicrobial will not inhibit bacterial growth at clinically achievable concentrations • Susceptible = Somewhat arbitrary designation that implies that an antimicrobial will inhibit bacterial growth at clinically achievable concentrations 2

Key Terms • MIC = Minimal inhibitory concentration. Lowest concentration of antimicrobial that inhibits growth of bacteria. Commonly used in clinical lab • MBC = Minimal bactericidal concentration. Concentration of an antimicrobial that kills bacteria. Used clinically only in special circumstances • Breakpoint = The MIC that is used to designate between susceptible and resistant. Arbitrarily set by a committee 3
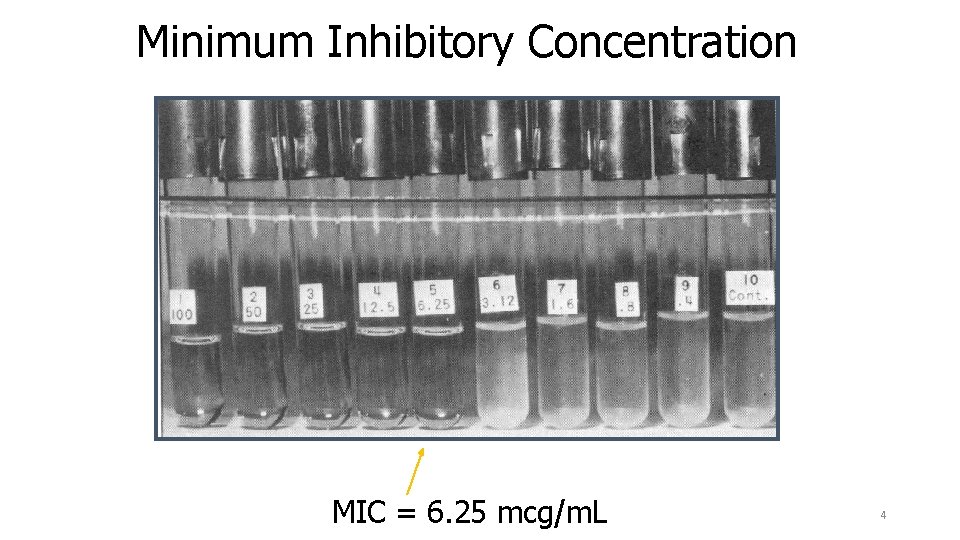
Minimum Inhibitory Concentration MIC = 6. 25 mcg/m. L 4

Automated Methods Well Plate for MIC Testing Many Labs Use Automated Testing 5
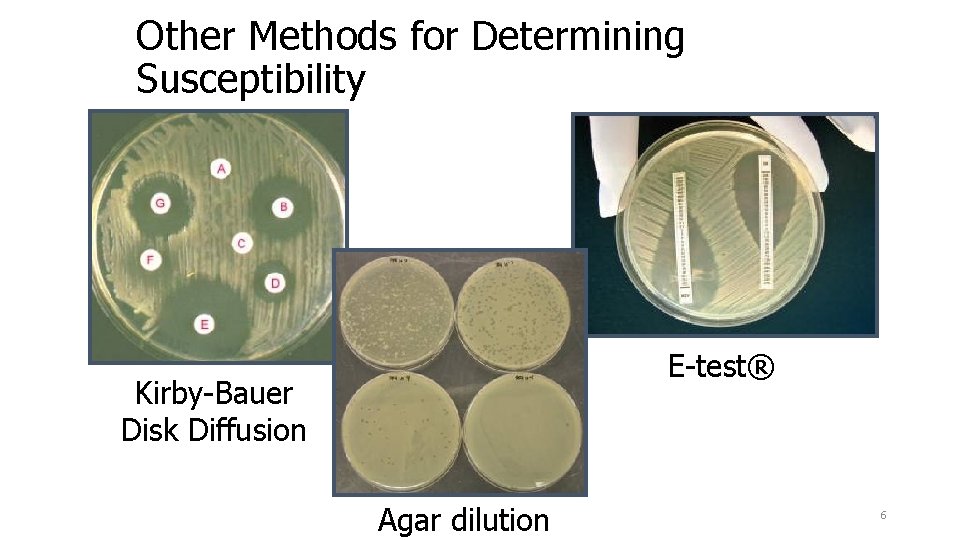
Other Methods for Determining Susceptibility E-test® Kirby-Bauer Disk Diffusion Agar dilution 6
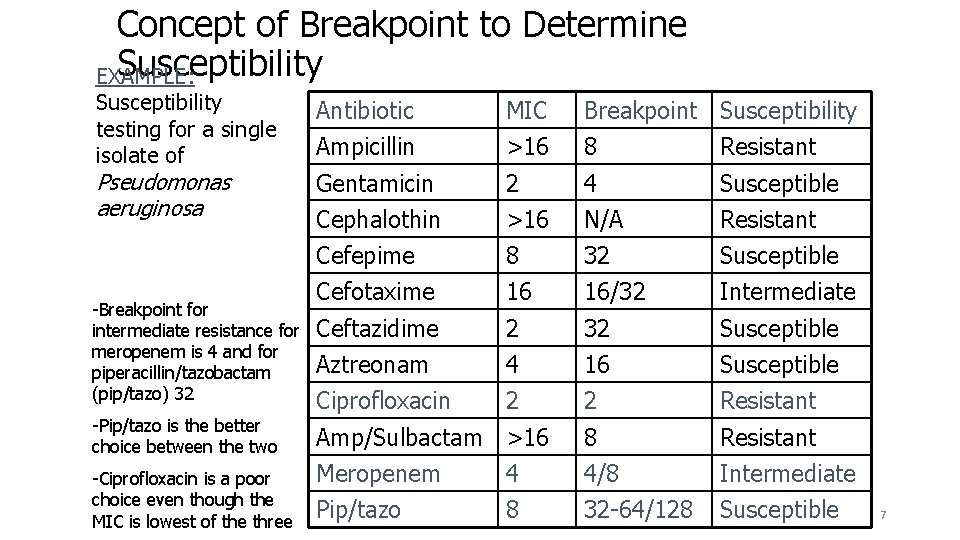
Concept of Breakpoint to Determine Susceptibility EXAMPLE: Susceptibility testing for a single isolate of Pseudomonas aeruginosa -Breakpoint for intermediate resistance for meropenem is 4 and for piperacillin/tazobactam (pip/tazo) 32 -Pip/tazo is the better choice between the two -Ciprofloxacin is a poor choice even though the MIC is lowest of the three Antibiotic MIC Breakpoint Susceptibility Ampicillin >16 8 Resistant Gentamicin 2 4 Susceptible Cephalothin >16 N/A Resistant Cefepime 8 32 Susceptible Cefotaxime 16 16/32 Intermediate Ceftazidime 2 32 Susceptible Aztreonam 4 16 Susceptible Ciprofloxacin 2 2 Resistant Amp/Sulbactam >16 8 Resistant Meropenem 4 4/8 Intermediate Pip/tazo 8 32 -64/128 Susceptible 7
Antibiotic Use Leads to Antibiotic Resistance Inpatient Agriculture Outpatient 8

Reasons for Antibiotic Overuse : Conclusions from 8 Focus Groups Patient Concerns • Want clear explanation • Green nasal discharge • Need to return to work Physician Concerns • Patient expects antibiotic • Diagnostic uncertainty • Time pressure Antibiotic Prescription Barden L. S. Clin Pediatr 1998; 37: 665 9
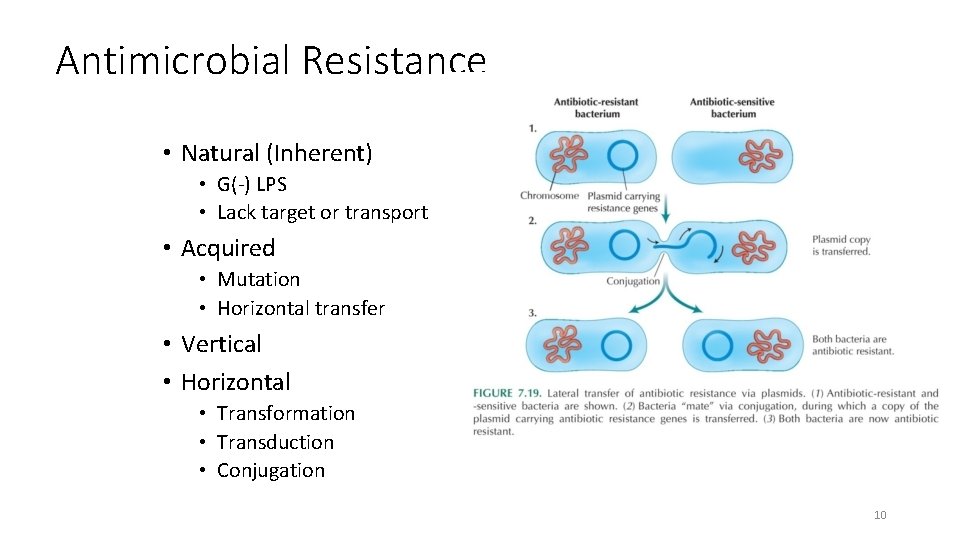
Antimicrobial Resistance • Natural (Inherent) • G(-) LPS • Lack target or transport • Acquired • Mutation • Horizontal transfer • Vertical • Horizontal • Transformation • Transduction • Conjugation 10
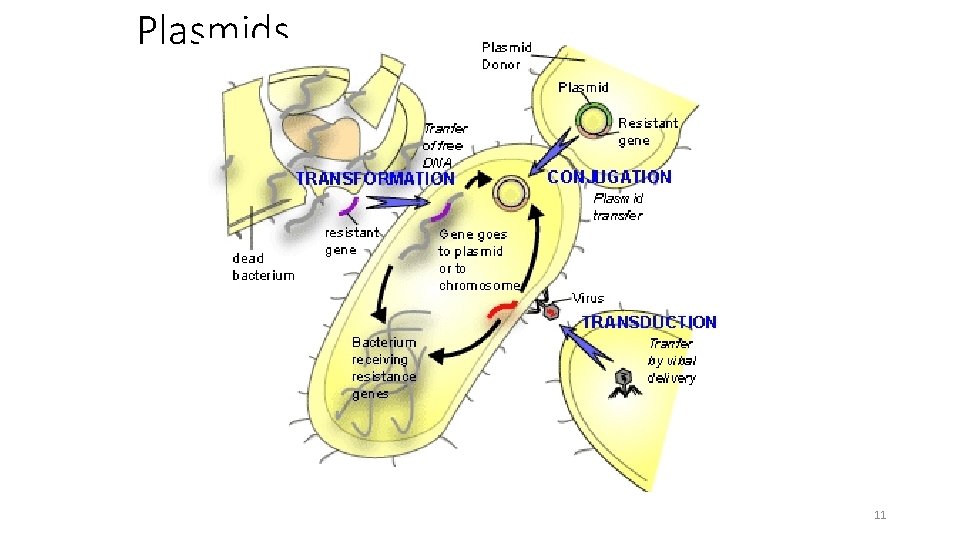
Plasmids 11
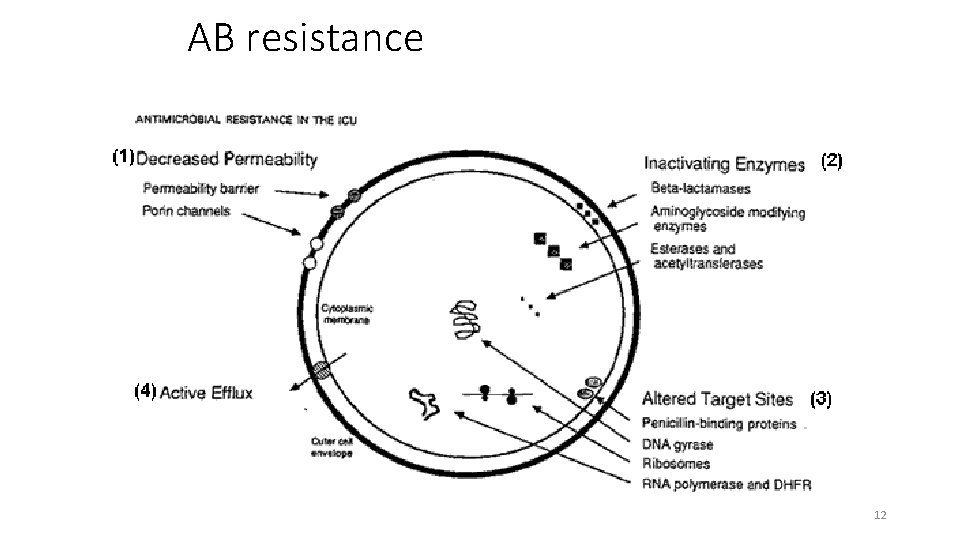
AB resistance 12

“Super Bugs” 13

Resistance to Antimicrobial Drugs • Mechanisms of resistance • Enzymes that cleave or otherwise inactivate antibiotics • β-lactamases • Changes in bacterial permeabilities • Prevents entry of antibiotic into cell • Mutation in target molecule • Alter binding characteristics of the antibiotics • Alteration of metabolic pathways • Some resistant bacteria can acquire PABA from the environment • Molecular pumps (efflux systems) • Secretion systems that export antibiotics faster than the rate of import 14

Nongenetic Origins of Drug Resistance • Low replication rates • Antibiotic is metabolized or neutralized before it act • Mycobacteria spp. • Alteration of cellular physiology • Bacterial L forms are cell wall-free • Streptococcus spp. , Treponema spp. , Bacillius spp. , others • Colonization of sites where antibiotics cannot reach • Gentamicin cannot enter cells • Salmonella are thus resistant to gentamicin 15

Genetic Origins of Drug Resistance • Chromosomal Resistance • Genes that regulate susceptibility • Often found in enzymes, r. RNA and secretion system genes • Mutations in RNApol render it resistant to the effects of rifampin • Efflux pumps with specificity for antibiotics • Found in all bacteria • All possess large hydrophobic cavity for binding antibiotics Five efflux pumps (“antiporters”) that regulate antibiotic resistance (Paulsen, 2003) 16

Genetic Origins of Drug Resistance • Extrachromosomal Resistance • Often account for interspecies acquisition of resistance • Contribute to multi-drug resistance (MDR) • Genetic elements are: • Plasmids • Transposons • Conjugation • Transduction • Transformation 17
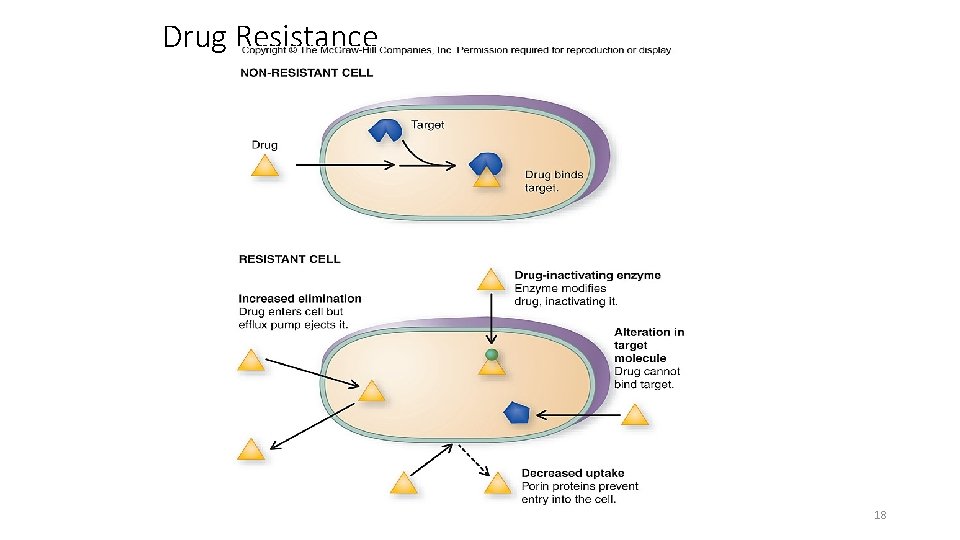
Drug Resistance 18

Antimicrobial Activity In Vivo • Drug-Pathogen Relationships • Environment • State of metabolic activity: slow-growing or dormant bacteria less susceptible • Distribution of drug: CNS is often exclusionary • Location of organisms: Some drugs do not enter host cells • Interfering substances: p. H, damaged tissues, etc. • Concentration • Absorption: some cannot be taken orally • Distribution: some accumulate in certain tissues • Variability of concentration: peaks and troughs • Postantibiotic effect: delayed regrowth of bacteria 19

Mechanisms of Resistance Overview Specific Examples 1. 2. 3. 4. Antibiotic Degrading Enzymes Decreased Permeability Efflux Pumps Target Alterations 20

Mechanisms of Resistance Antibiotic Degrading Enzymes • Sulfonation, phosphorylation, or esterifictation • Especially a problem for aminoglycosides • β-lactamases • Simple, Extended spectrum β-lactamases (ESBL), cephalosporinases, carbapenemases • Confer resistance to some, many, or all beta-lactam antibiotics • May be encoded on chromosome or plasmid • More potent in Gram-negative bacteria • Examples: S. aureus, H. influenzae, N. gonorrhoeae, E. coli, Klebsiella sp. , Enterobacter sp. , Serratia sp. , other enteric bacteria, anaerobes 21

Extended Spectrum -lactamases • -lactamases capable of hydrolysing extended spectrum cephalosporins, penicillins, and aztreonam • Most often associated with E. coli and Klebsiella pneumoniae but spreading to other bacteria • Usually plasmid mediated • Aminoglycoside, ciprofloxacin and trimethoprimsulfamethoxazole resistance often encoded on same plasmid • Has become a significant resistance determinate in acute and long-term care facility enteric pathogens 22

Class A Carbapenemases • Most common in Klebsiella pneumoniae (KPC) • Also seen in E. coli, Enterobacter, Citrobacter, Salmonella, Serratia, Pseudomonas and Proteus spp. • Very often with multiple other drug resistance mechanisms, resistance profile similar to ESBL but also carbapenem resistant • Became problem in New York City first in 2002 -2003 and is being increasingly recognized in Mid-Atlantic US. • Spreading across species to other Gram-negatives and Enterobacteriaceae • Emerging in long-term care facilities 23

Mechanisms of Resistance Decreased Permeability • Pseudomonas spp. • Affects many antibiotics including carbapenems Efflux Pumps • Pseudomonas spp. (multiple antibiotics) • Tetracyclines • Macrolides 24

Mechanisms of Resistance Target Alteration • DNA gyrase • Fluoroquinolones • Many Gram-negatives, S. pneumoniae • Penicillin-binding protein • Methicillin-resistant S. aureus (MRSA) • Penicillin-resistant S. pneumoniae • Gram positive cell wall • Vancomycin • Enterococcus spp. 25

Mechanisms of Resistance Target Alteration • Ribosome • Tetracyclines • Macrolides • S. pneumoniae, Staphylococcus sp. , N. gonorrhoeae, enteric Gram-negative rods 26

VRE-Vancomycin resistant Enterococci • VRE- found first time in France, 1986 • Vancomycin n glycopeptide (avaporcin) used in feed-Europe • VRE found in horse, pigs, dogs and poulty- used avaporcin in feed • Use of avaporcin – leads development of VRE • Food products from Europe contain VRE • 1996, use of avaporcin is banned and drop of VRE rate • Enterococcus faecium and Enterococcus faecalis • Vancomycin is the only antibiotic for MRSA strain 27
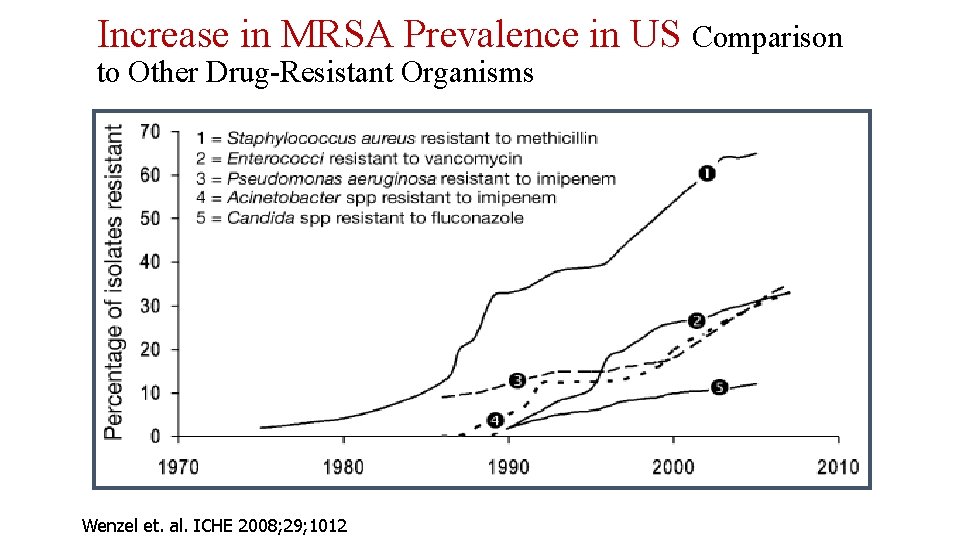
Increase in MRSA Prevalence in US Comparison to Other Drug-Resistant Organisms Wenzel et. al. ICHE 2008; 29; 1012

Conclusion • Inappropriate and excessive use of antibiotics is a major factor contributing to emerging antibiotic resistance • Determinants of resistance are selected for by antibiotic use • Multiple mechanisms exist for bacteria to become resistant to antibiotics • Antibiotic resistance is a problem in outpatient and inpatient settings and is a factor in a wide variety of infections • Antibiotic resistance continues to emerge as a serious threat to public health 29

30
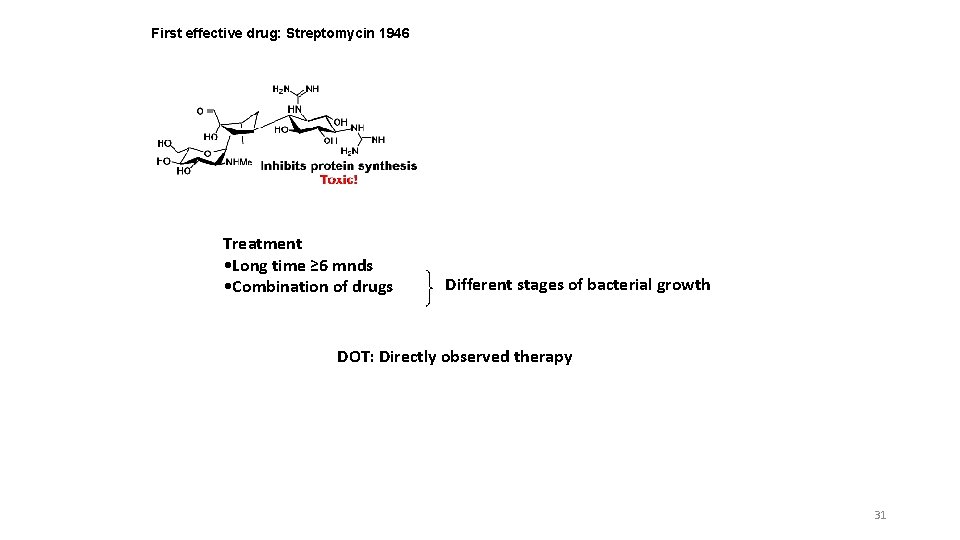
First effective drug: Streptomycin 1946 Treatment • Long time ≥ 6 mnds • Combination of drugs Different stages of bacterial growth DOT: Directly observed therapy 31
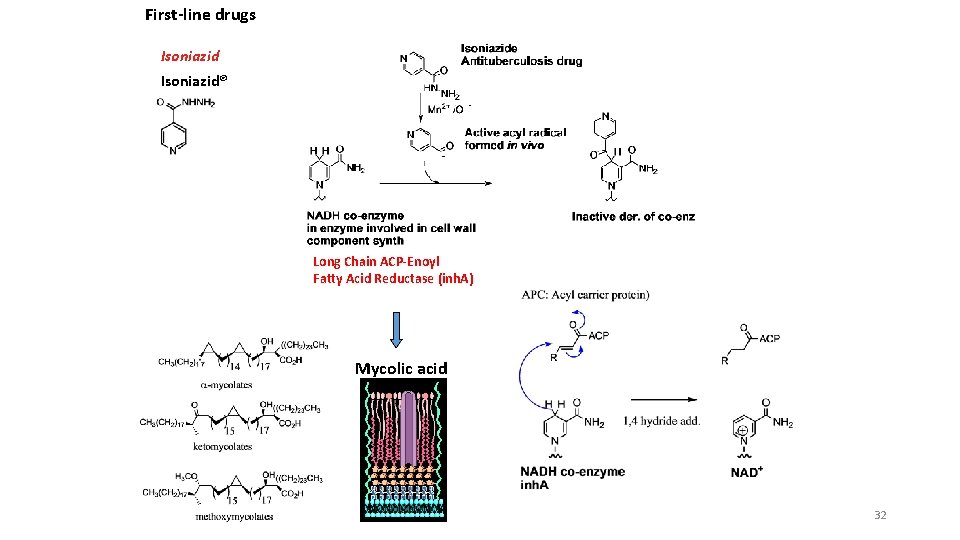
First-line drugs Isoniazid® Long Chain ACP-Enoyl Fatty Acid Reductase (inh. A) Mycolic acid 32
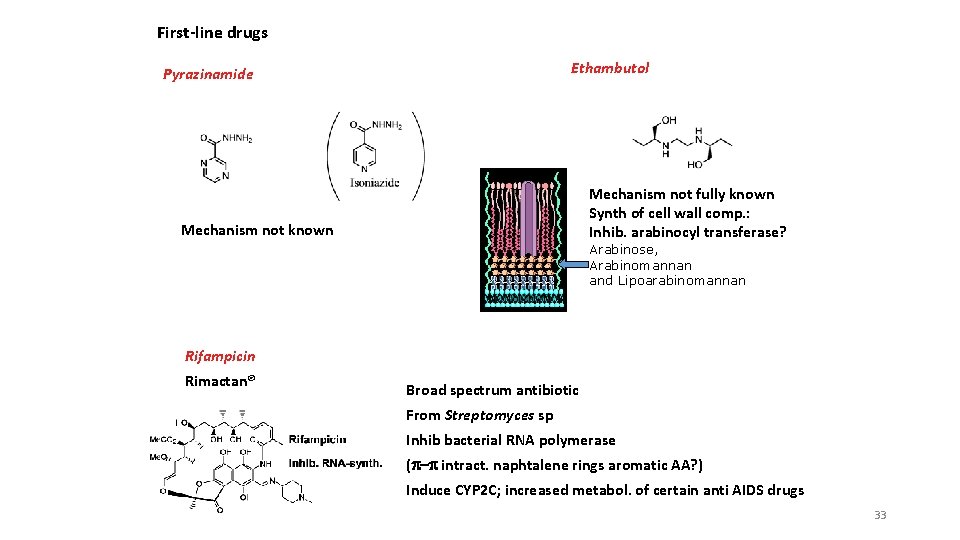
First-line drugs Ethambutol Pyrazinamide Mechanism not fully known Synth of cell wall comp. : Inhib. arabinocyl transferase? Mechanism not known Arabinose, Arabinomannan and Lipoarabinomannan Rifampicin Rimactan® Broad spectrum antibiotic From Streptomyces sp Inhib bacterial RNA polymerase (p-p intract. naphtalene rings aromatic AA? ) Induce CYP 2 C; increased metabol. of certain anti AIDS drugs 33
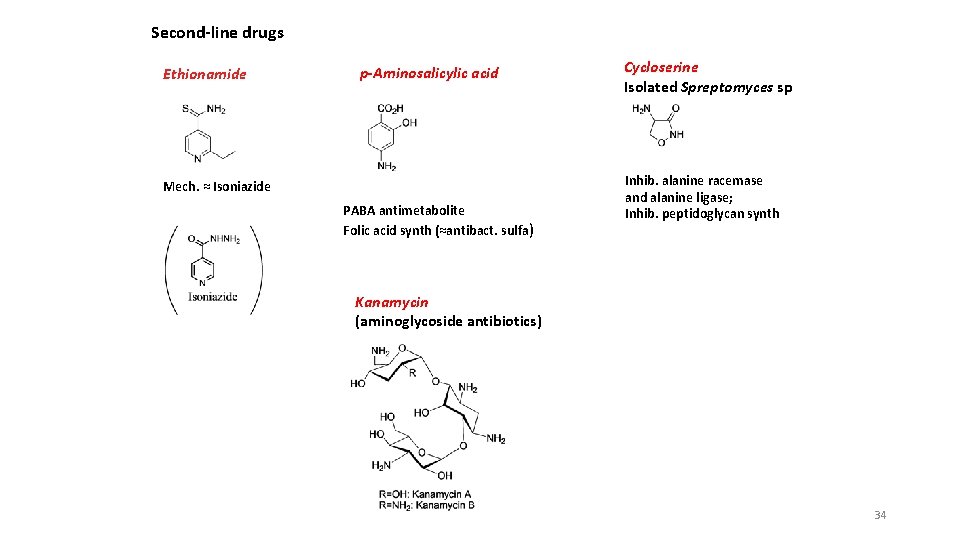
Second-line drugs Ethionamide p-Aminosalicylic acid Mech. ≈ Isoniazide PABA antimetabolite Folic acid synth (≈antibact. sulfa) Cycloserine Isolated Spreptomyces sp Inhib. alanine racemase and alanine ligase; Inhib. peptidoglycan synth Kanamycin (aminoglycoside antibiotics) 34
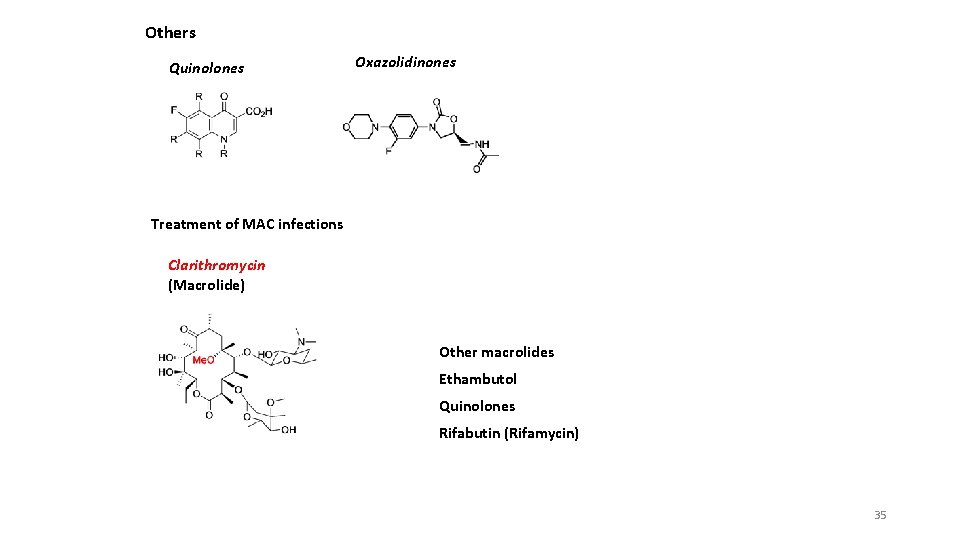
Others Quinolones Oxazolidinones Treatment of MAC infections Clarithromycin (Macrolide) Other macrolides Ethambutol Quinolones Rifabutin (Rifamycin) 35

Inhibition of AFB • MOA • Disrupt cell wall synthesis • Cycloserine • Prevents replication • Clofazimine • Disrupt Waxy layer • Mycolic acid and Arabinogalactan • Examples • Isoniazid • Ethambutal • Mycobacterium • Multidrug Therapies 36

37
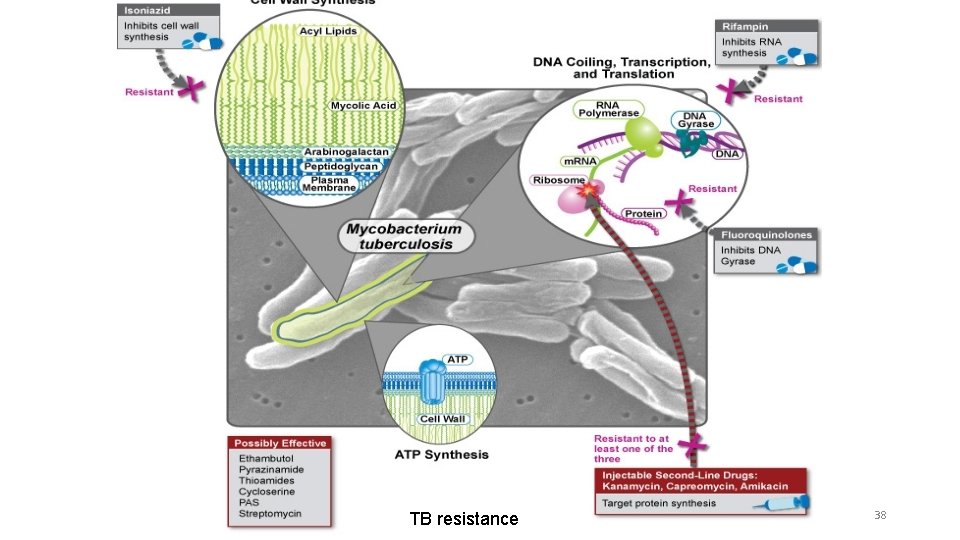
TB resistance 38

Antibacterial Antibiotics Inhibitors of Cell Wall Synthesis • Antimycobacterium antibiotics • Isoniazid (INH) • Inhibits mycolic acid synthesis • Ethambutol • Inhibits incorporation of mycolic acid Multidrug-Resistant Tuberculosis Infection 39
- Slides: 39