Announcements for MCB 110 L Office Hours starting

Announcements for MCB 110 L Office Hours (starting Mon. , 28 Jan. ): M 12 -1 PM Jane GPB 203 Tu 2 -3 PM Dale GPB 109 W 12 -1 PM Jesse GPB 201 W 12 -1 PM Maumita GPB 203 F 12 -1 PM Joo Eun GPB 203 Th 3 -4 PM Jeremy 526 Barker
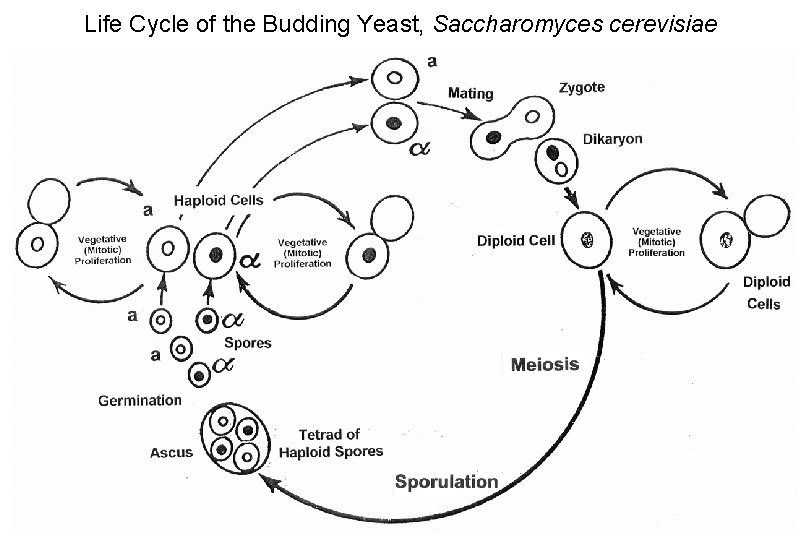
Life Cycle of the Budding Yeast, Saccharomyces cerevisiae

Genetic Nomenclature Conventions for Saccharomyces cerevisiae Normal (“wild-type”) locus: YFG 1 Loss-of-function (hypomorphic) allele: yfg 1 -1 (point mutation) yfg 1 -∆1 (null mutation) Gain-of-function (hypermorphic) allele: YFG 1 -20 Alteration-of-function (neomorphic) allele: YFG 1 -54 A normal, hypermorphic or neomorphic allele is dominant over a hypomorphic allele: YFG 1/yfg 1 Presence and status of an episome or non-Mendelian determinant in brackets: [YCp 352], introduced DNA plasmid [cir+], endogenous 2 m DNA circle [rho-plus = +], functional mitochondrial DNA [psi+], presence of prion form of Sup 35 (yeast e. RF 3) [KIL-k 1], endogenous ds. RNA viroid encoding secreted toxin
![FCP 1 [TFIIF-associated Rpo 21 carboxy-terminal domain (CTD) phosphatase] Strain YSA 1: MAT fcp FCP 1 [TFIIF-associated Rpo 21 carboxy-terminal domain (CTD) phosphatase] Strain YSA 1: MAT fcp](http://slidetodoc.com/presentation_image_h/28015f49b3b39bc6d40fadf0fae71630/image-4.jpg)
FCP 1 [TFIIF-associated Rpo 21 carboxy-terminal domain (CTD) phosphatase] Strain YSA 1: MAT fcp 1∆: : LEU 2 TRP 1: : fcp 1 -2 ts ade 2 his 3 leu 2 trp 1 ura 3 [YCp-FCP 1, URA 3] The FCP 1 gene is essential for S. cerevisiae cell viability. (mutations in the orthologous human gene product, CTDP 1, causes congenital cataracts with facial dysmorphism and neuropathy)

The Concept of a Genetic Cross e. g. , MATa mutation a X MAT mutation b What can we learn from such crosses? • Assess whether mutation a is dominant or recessive to the corresponding WT locus, and assess whether mutation b is dominant or recessive to the corresponding WT locus (dominance test) • Determine whether mutation a and mutation b are likely to be alterations of the same gene (complementation test) • Map the relative positions of locus a and locus b by examining their segregation behavior in meiosis (tetrad analysis) • Generate a potentially useful double mutant (epistasis test)
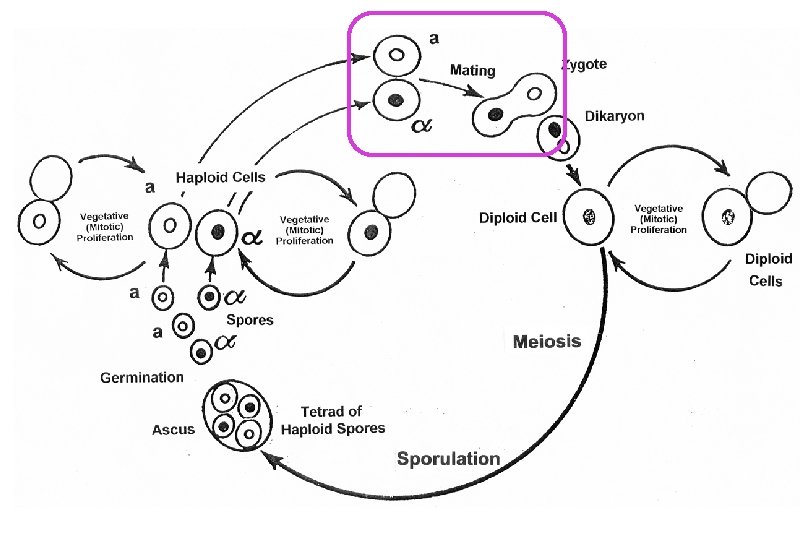
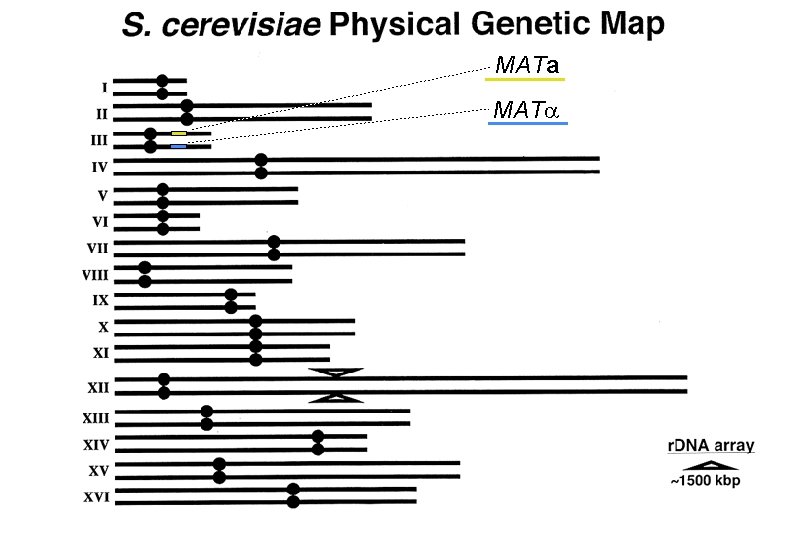
MATa MAT
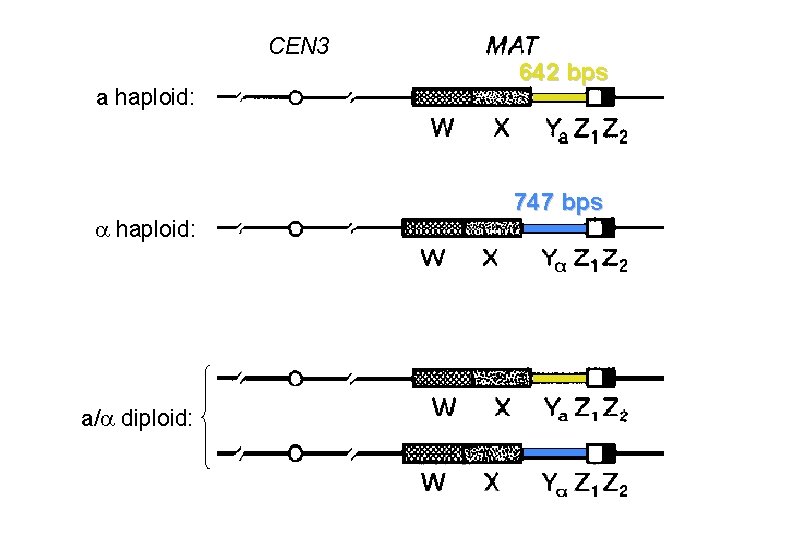
CEN 3 a haploid: a/ diploid: 642 bps 747 bps
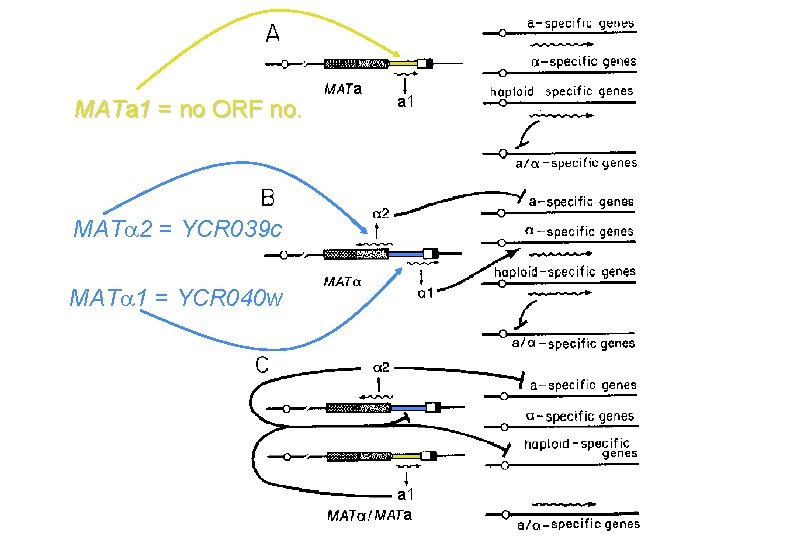
MATa 1 = no ORF no. a a 1 MAT 2 = YCR 039 c MAT 1 = YCR 040 w a 1 a
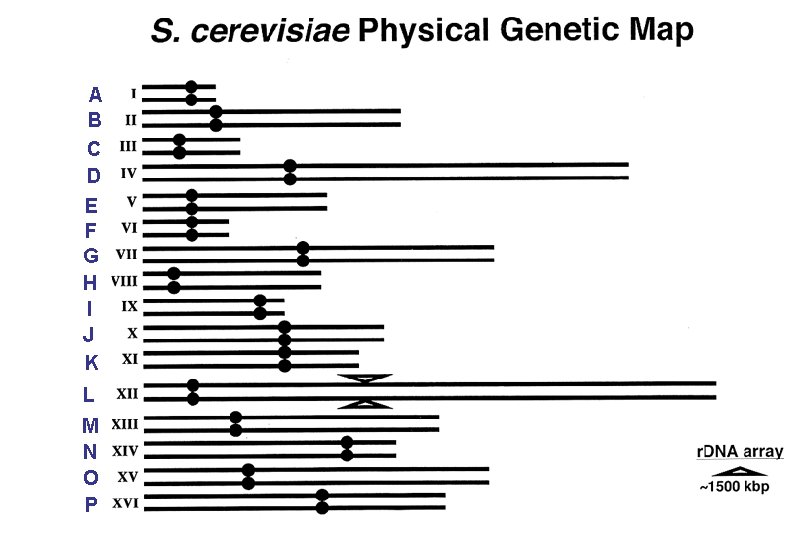
A B C D E F G H I J K L M N O P

Systematic name for the FCP 1 locus is YMR 277 w. Automatically tells you that the ORF encoded by the FCP 1 gene is situated on the right arm of Chromosome XIII and transcribed in the direction away from CEN 13.
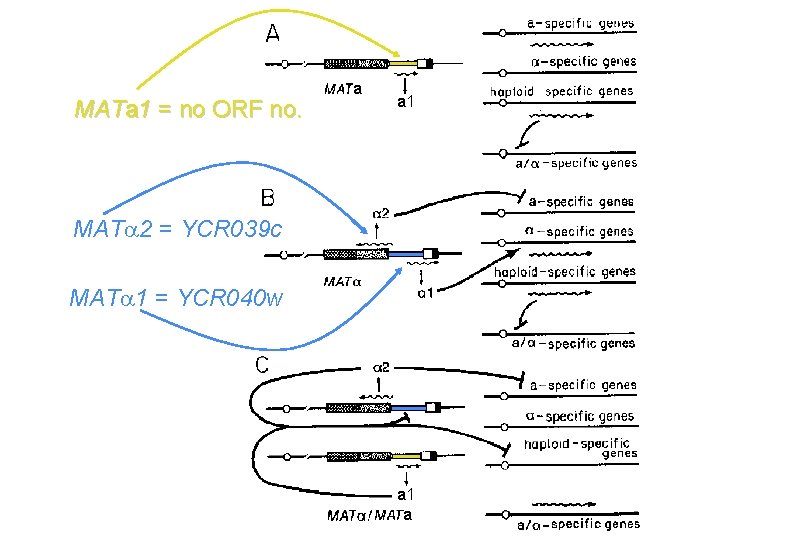
MATa 1 = no ORF no. a a 1 MAT 2 = YCR 039 c MAT 1 = YCR 040 w a 1 a

Available from Cold Spring Harbor Laboratory Press

The Concept of a Genetic Cross e. g. , MATa mutation a X MAT mutation b What can we learn from such crosses? • Assess whether mutation a is dominant or recessive to the corresponding WT locus, and assess whether mutation b is dominant or recessive to the corresponding WT locus (dominance test) • Determine whether mutation a and mutation b are likely to be alterations of the same gene (complementation test) • Map the relative positions of locus a and locus b by examining their segregation behavior in meiosis (tetrad analysis) • Generate a potentially useful double mutant (epistasis test)
- Slides: 14