Animal Improvement Program AIP A big data project

Animal Improvement Program (AIP) A “big data” project of the Animal Genomics and Improvement Laboratory (AGIL) George R. Wiggans Animal Genomics and Improvement Laboratory Agricultural Research Service, USDA Beltsville, MD 20705‐ 2350 george. wiggans@ars. usda. gov http: //aipl. arsusda. gov/ ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (1) Wiggans

AGIL mission l l Discover and develop improved methods for the genetic and genomic evaluation of economically important traits of dairy animals and small ruminants Conduct fundamental genomics‐based research aimed at improving their health and productive efficiency ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (2) Wiggans

AGIL staff l Dr. Erin E. Connor, Research Leader l 10 senior scientists l 2 postdoctoral associates l 9 support scientists l 2 chemists l 5 laboratory technicians l 3 information technology specialists l 2 administrative assistants l Visiting scientists and students ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (3) Wiggans

AGIL appropriated projects l l Enhancing genetic merit of ruminants through genome selection and analysis Understanding genetic and physiological factors affecting nutrient use efficiency of dairy cattle Development of genomic tools to study ruminant resistance to gastrointestinal nematodes Improving genetic predictions in dairy animals using phenotypic and genomic information “Animal Improvement Program” (AIP) ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (4) Wiggans

AIP staff l 4 senior scientists Dr. George Wiggans Dr. Paul Van. Raden Dr. John Cole l 6 support scientists l 3 information technology specialists l 1 administrative assistant l 2 visiting scientists ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (5) Dr. Derek Bickhart Wiggans

AIP objectives l l l Expand national and international collection of phenotypic and genotypic data Develop a more accurate genomic evaluation system with advanced, efficient methods to combine pedigrees, genotypes, and phenotypes Use economic analysis to maximize genetic progress and financial benefits from collected data ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (6) Wiggans

Genetic evaluation l Improve future performance through selection l Possible data w Animal’s own measurable traits w Pedigrees and phenotypes of relatives w Genomic information ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (7) Wiggans

Phenotypic data l l Records for milk yield, fat percentage, protein percentage, and somatic cell count (1/month) Appraiser‐assigned scores for 16 body and udder characteristics related to conformation (e. g. , stature) Breeding records that include indicator for conception success Calving difficulty scores and stillbirth occurrences ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (8) Wiggans

Primary traits evaluated l Yield (milk, fat, and protein) l Conformation (overall and individual traits) l Longevity (productive life) l Fertility (conception and pregnancy rates) l Calving (dystocia and stillbirth) l Disease resistance (somatic cell score) ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (9) Wiggans

Data amounts (as of July 2015) l Pedigree records 71, 974, 045 l Animal genotypes 1, 035, 590 l Lactation records (since 1960) 132, 629, 200 l Daily yield records (since 1990) 641, 864, 015 l Reproduction event records 176, 559, 035 l Calving difficulty scores 29, 528, 607 l Stillbirth scores 19, 567, 198 ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (10) Wiggans

Value of incoming data Data Phenotypes (2014) 4 million cows × $1. 25/cow/month Genotypes (2014) 15, 000 medium‐density × $125 258, 000 low‐density × $45 Whole‐genome sequence (2015) 200+ bulls × $1, 000+ bulls × $3, 000 Total ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (11) Annual value $60 million $2 million $12 million $0. 2 million $3 million $77. 2 million Wiggans

Genomics and SNP l l l Genomics – Applies DNA technology and bioinformatics to sequence, assemble and analyze the function and structure of genomes SNP – Single nucleotide polymorphisms; serve as markers to track inheritance of chromosomal segments Genomic selection – Selection using genomic predictions of economic merit early in life ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (12) Wiggans

Benefit of genomics l Determine value of bull at birth l Increase accuracy of selection l Reduce generation interval l Increase selection intensity l Increase rate of genetic gain ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (13) Wiggans

Why genomics works for dairy cattle l Extensive historical data available l Well‐developed genetic evaluation program l Widespread use of artificial‐insemination (AI) sires l Progeny‐test programs l High‐value animals worth the cost of genotyping l Long generation interval that can be reduced substantially by genomics ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (14) Wiggans

Evaluation transition to dairy industry l l Council on Dairy Cattle Breeding (CDCB) w Database maintenance w Calculation and distribution of genetic merit estimates w Interface with evaluation users and data suppliers AGIL w Research and development using data made available by CDCB ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (15) Wiggans
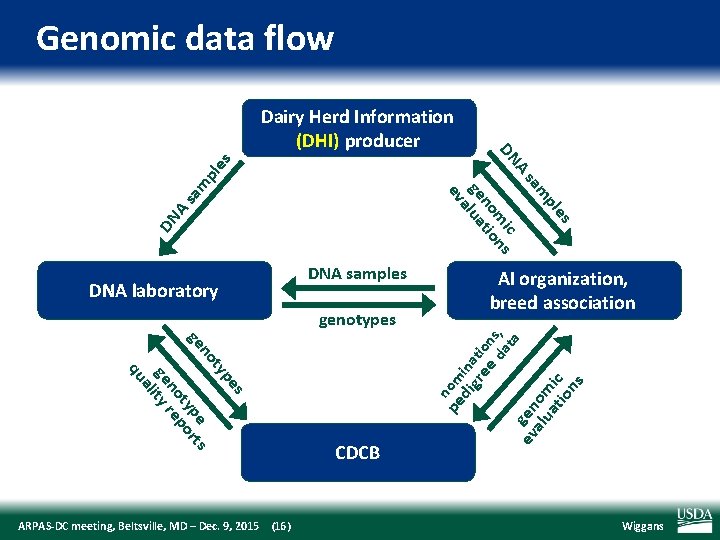
Genomic data flow DN A sa m pl es es pl m sa A ic DN m ns no tio ge lua a ev Dairy Herd Information (DHI) producer ta da s pe ty no pe ts ge ty or no rep ge lity a qu ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (16) pe nom di ina gr ti ee on s, genotypes AI organization, breed association CDCB ev gen alu om at ic io ns DNA laboratory DNA samples Wiggans

Evaluation flow l l Animal nominated for genomic evaluation by approved nominator DNA source sent to genotyping lab (2014) Source Blood Hair Nasal swab Semen Tissue Unknown Samples (no. ) 10, 727 113, 455 2, 954 3, 432 149, 301 12, 301 ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (17) Samples (%) 4 39 1 1 51 4 Wiggans

Evaluation flow l l (continued) DNA extracted and placed on chip w Marker panels that range from 2, 900 to 777, 962 SNPs w 3‐day genotyping process Genotypes sent from genotyping lab for accuracy review ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (18) Wiggans
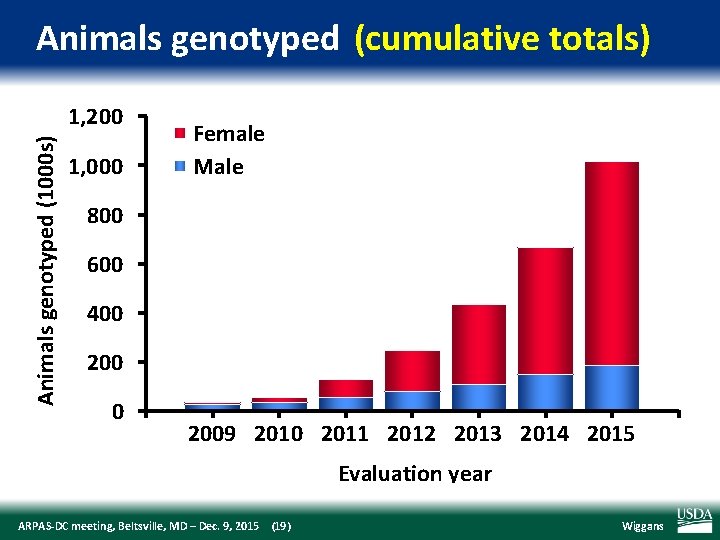
Animals genotyped (cumulative totals) Animals genotyped (1000 s) 1, 200 1, 000 Female Male 800 600 400 2009 2010 2011 2012 2013 2014 2015 Evaluation year ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (19) Wiggans

Laboratory quality control l Each SNP evaluated for w Call rate w Portion heterozygous w Parent‐progeny conflicts l Clustering investigated if SNP exceeds limits l Number of failing SNPs indicates genotype quality l Target of <10 SNPs in each category ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (20) Wiggans

Evaluation flow (continued) l Genotype calls modified as necessary l Genotypes loaded into database l Nominators receive reports of parentage and other conflicts l Pedigree or animal assignments corrected l Genotypes extracted and imputed to 61 K l SNP effects estimated l Final evaluations calculated ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (21) Wiggans

Parentage validation and discovery l l Parent‐progeny conflicts detected w Animal checked against all other genotypes w Reported to breeds and requesters w Correct sire usually detected Maternal grandsire checking w SNP at a time checking w Haplotype checking more accurate ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (22) Wiggans

Evaluation flow l (continued) Evaluations released to dairy industry w Download from FTP site with separate files for each nominator w Weekly release of evaluations of new animals w Monthly release for females and bulls not marketed w All genomic evaluations updated 3 times each year with traditional evaluations ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (23) Wiggans
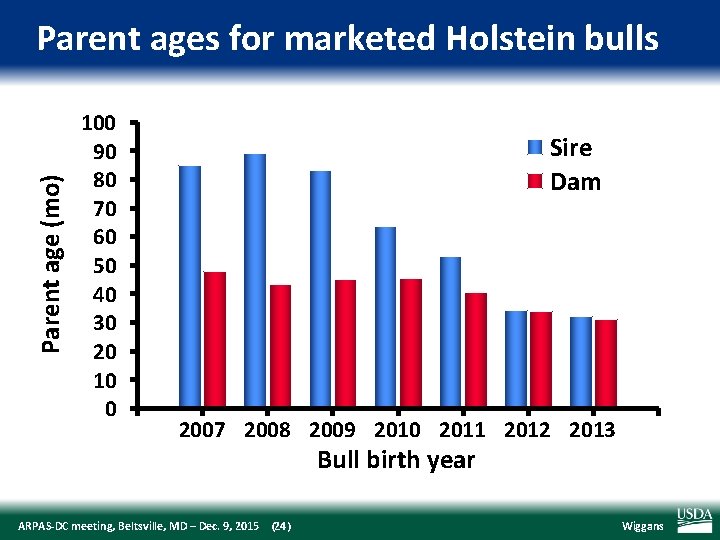
Parent age (mo) Parent ages for marketed Holstein bulls 100 90 80 70 60 50 40 30 20 10 0 Sire Dam 2007 2008 2009 2010 2011 2012 2013 Bull birth year ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (24) Wiggans
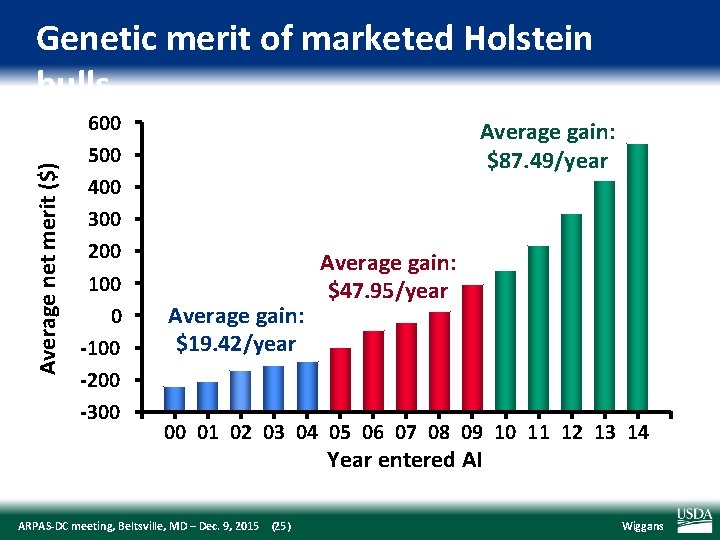
Average net merit ($) Genetic merit of marketed Holstein bulls 600 500 400 300 200 100 0 ‐ 100 ‐ 200 ‐ 300 Average gain: $87. 49/year Average gain: $19. 42/year Average gain: $47. 95/year 00 01 02 03 04 05 06 07 08 09 10 11 12 13 14 Year entered AI ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (25) Wiggans

Improving accuracy l l Increase size of predictor population w Share genotypes across country w Young bulls receive progeny test Use more or better SNPs Account for effect of genomic selection on traditional evaluations Reduce cost to reach more selection candidates ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (26) Wiggans
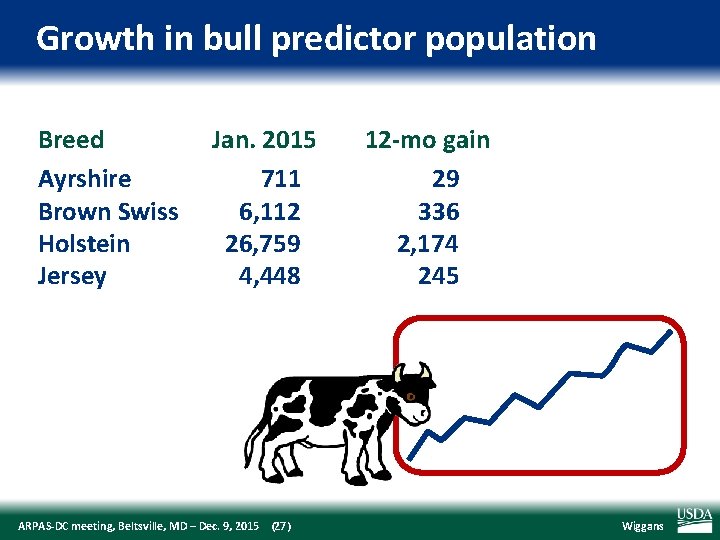
Growth in bull predictor population Breed Ayrshire Brown Swiss Holstein Jersey Jan. 2015 711 6, 112 26, 759 4, 448 ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (27) 12‐mo gain 29 336 2, 174 245 Wiggans

Haplotypes affecting fertility l l Rapid discovery of new recessive defects w Large numbers of genotyped animals w Affordable DNA sequencing Determination of haplotype location w Significant number of homozygous animals expected, but none observed w Narrow suspect region with fine mapping w Use sequence data to find causative mutation ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (28) Wiggans

Current research areas l l Improve evaluation methodology Develop applications for sequence data Acquire data for additional traits Develop evaluations for new traits ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (29) Wiggans

Mating programs l Genomic relationships of genotyped females with available bulls provided l Determination of best mate possible l Dominance effects could be considered ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (30) Wiggans

Working with sequence data l l l Sequence data available from 1000 Bull Genomes Project hosted in Australia Project funded by industry to sequence over 200 bulls to create a haplotype library A posteriori granddaughter design to locate chromosomal segments of interest from 71 bulls each with over 100 genotyped and progeny‐tested sons ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (31) Wiggans
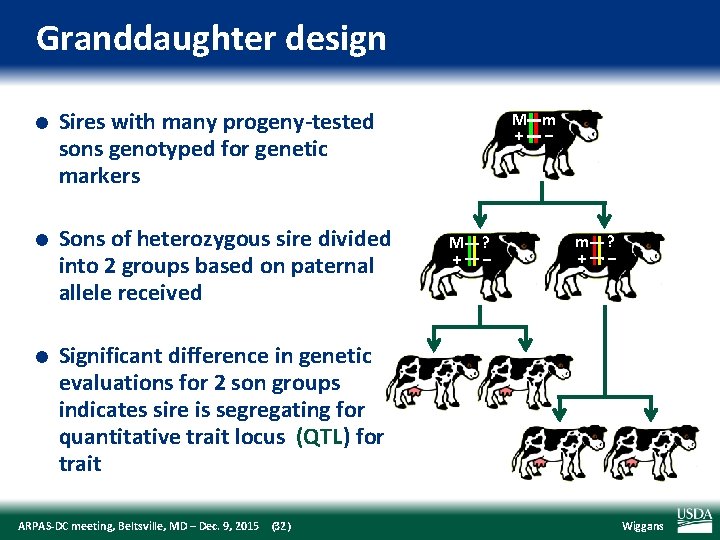
Granddaughter design l l l Sires with many progeny‐tested sons genotyped for genetic markers Sons of heterozygous sire divided into 2 groups based on paternal allele received M m + – M + ? – m + ? – Significant difference in genetic evaluations for 2 son groups indicates sire is segregating for quantitative trait locus (QTL) for trait ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (32) Wiggans

Alignment of sequence data l l Alignment – determining location of chromosomal segments provided by sequencer Findmap – matches segment against library of haplotypes l Preserves low‐frequency variants l Does not identify new variants l Uses a hash table to find variant enabling rapid processing ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (33) Wiggans

Further use of sequence data l Discovery of causative genetic variants l Refinement of SNPs used in genomic evaluation l w Add discovered causative variants w Use some SNPs for imputation but not for estimation of SNP effects Create genotypes for genomic evaluation from sequence data to enable immediate use through imputation of any new SNPs ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (34) Wiggans

Additional traits requiring data l l l l l Clinical mastitis Displaced abomasum Ketosis Hoof health Immune response Other health traits Feed efficiency Methane production Milk fatty acid compositio n from mid‐infrared spectroscopy ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (35) Wiggans

Evaluation of new traits l Mortality l Days to first breeding l Gestation length l Persistency l Resistance to h eat stress (predicting genotype × environment interactions) ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (36) Wiggans

Benefits to dairy industry l l l Low‐cost genotyping tools for genomic predictions of genetic merit Identification of gene mutations for cow fertility Genetic evaluations for more than 30 traits of U. S. dairy cows Genetic‐economic indexes to help dairy farmers choose parents of future generations Genomic mating programs for dairy cattle ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (37) Wiggans

Impact on breeders l l Haplotype and gene tests in selection and mating programs Trend towards a small number of elite breeders that are investing heavily in genomics About 30% of young males genotyped directly by breeders since April 2013 Prices for top genomic heifers can be very high (e. g. , $265, 000 ) ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (38) Wiggans

Impact on dairy producers l l General w Reduced generation interval w Increased rate of genetic gain w More inbreeding/homozygosity? Sires w Higher average genetic merit of available bulls w More rapid increase in genetic merit for all traits w Larger choice of bulls for traits and semen price w Greater use of young bulls ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (39) Wiggans

Summary l l Highly successful program leading to annual increases in genetic merit for production efficiency Large database of phenotypic and genomic data provided by industry Research projects to determine mechanism of genetic control of economically important traits Data processing techniques developed so that rapid turnaround could be realized ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (40) Wiggans

Funding acknowledgments l U. S. taxpayers (USDA appropriated project) l Council on Dairy Cattle Breeding l Binational Agricultural Research & Development l National Institute of Food and Agriculture l Washington State University (NIFA grant) ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (41) Wiggans

Questions? AIP web site: http : //aipl. arsusda. gov Holstein and Jersey crossbreds graze on American Farm Land Trust’s Cove Mountain Farm in south‐central Pennsylvania Source: ARS Image Gallery, image #K 8587 -14; photo by Bob Nichols ARPAS‐DC meeting, Beltsville, MD – Dec. 9, 2015 (42) Wiggans
- Slides: 42