Analyzing Patterns Using Principal Component Analysis BMSC 601

Analyzing Patterns Using Principal Component Analysis BMSC 601 W. Rose

PCA Hubley-Kozey et al. (2006) “Neuromuscular alterations during walking…”, J. Electromyo. Kinesiol. 16: 365 -378. EMGs and kinematic data are complex, or “highdimensional”, i. e. they have a lot of numbers: they vary with time and by site. Pattern analysis tries to find patterns in complex data, and thus to reduce complex data to a smaller set of numbers that can be quantitatively compared (“data reduction”). Principal component analysis (PCA) is one kind of pattern analysis. BMSC 601 W. Rose

PCA Assume we have multiple “copies” of a signal. The copies could be from multiple trials in one person, or from different people, etc. Goal Find patterns in the signals and quantify whethere are differences in patterns between subgroups. BMSC 601 W. Rose

PCA Approach to the Goal Find the pattern (wave shape) which, when scaled up or down by an adjustable scaling factor, accounts for as much of the power as possible in the signals. This is the first principal component. The scaling factor is adjusted for each trial individually, but the basic shape is not adjustable and cannot “slide” along the time axis. BMSC 601 W. Rose

PCA Approach to the Goal, cont. Best basic shape = 1 st principal component. Once 1 st PC is found, find scale factor for each trial. Scale factor tells us “how much” of the 1 st PC there is in each trial. (Scale factor could even be negative. ) Collect scale factors from “experimentals” & controls and compare them using standard statistics (t test, Wilcoxon ranksum, ANOVA, etc), to see if scale factors differ from group to group. If they do, it means that “size” of 1 st PC is different in diff. groups. BMSC 601 W. Rose

PCA for EMGs PCA Approach to the Goal, cont. Scaled versions of 1 st PC can be subtracted from each signal to get “first residual” for each trial, i. e. the part of signal not accounted for by 1 st PC. Now repeat what we did before, using 1 st residuals of each signal, instead of the actual signals. The shape which, when scaled separately for each trial, does the best job in accounting for the 1 st residuals, is the 2 nd PC. BMSC 601 W. Rose

PCA for EMGs PCA Approach to the Goal, cont. As with 1 st PC, we find a scale factor for each trial. The scale factor is “how much” of 2 nd PC there is in each trial, i. e. it’s the amount that 2 nd PC should be multiplied by to get best fit to that trial’s 1 st residual. Once again, we can use statistics to compare 2 nd PC scale factors across groups to see if “size” of 2 nd PC is different in diff. groups. BMSC 601 W. Rose

PCA Approach to the Goal, cont. We now compute “ 2 nd residuals” by subtracting the scaled version of 2 nd PC from each 1 st residual. We use the 2 nd residuals to get the third PC, the scaling factors for 3 rd PC, and so on. If there are N time points in each signal, we can keep going until we find N principal components. BMSC 601 W. Rose

PCA The high numbered PCs are probably not very significant. They may be just fitting the noise. How many PCs should we analyze? Standard Answers: Accept only enough PCs to account for 90% (or 95%) of the variance in the original signals Or Accept PCs until the next one accounts for less than 1% of the variance in the original signals. BMSC 601 W. Rose

PCA as described above is done on signals whose mean values have been subtracted off, so the signals analyzed WILL have zero mean value. If desired, the means can be added back on at the end. Hubley-Kozey et al. use Karhunen-Loeve transformation (KLT). Raptopoulos et al. (J Biomech 2006) use “KLT”; they DO subtract mean values first. Raptop 2006 use correlation mtx instead of covariance mtx. BMSC 601 W. Rose

PCA In PCA and KLT, start with matrix X whose columns are the signals (e. g. EMGs) from diff. subjects. Rows are different time points. KLT and PCA: compute Cov(X) (or Corr(X): Raptopoulos et al 2006; Schutte et al 2000; Deluzio et al 1997) Cov(X) = [X-E(X)]*[X-E(X)]T / (m-1) E(X) = matrix whose every column = the mean vector. Eigenvectors of Cov(X) are the PCs. Eigenvalues of Cov(X) indicate how much of the variance is accounted for by each PC. BMSC 601 W. Rose

PCA Acc. Gerbrands, 1981: KLT: compute y = TT x PCA: compute y = TT [x-E(x)] where E(x)=mean vector. Hall Müller Wang 2006 define KLT with de-meaned X; equate KLT with “functional PCA” BMSC 601 W. Rose

PCA BMSC 601 Simulate some EMGs. Each EMG is a weighted sum of 4 components (pulses). The weight (i. e. factor that multiplies the y -scale) of each component is a random number between -1, +1. The random numbers differ from trace to trace. Some random noise is also added at each time point. W. Rose
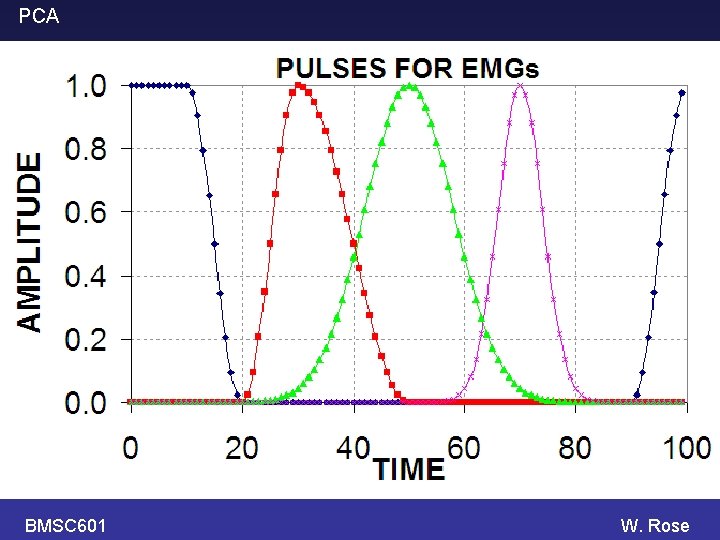
PCA BMSC 601 W. Rose
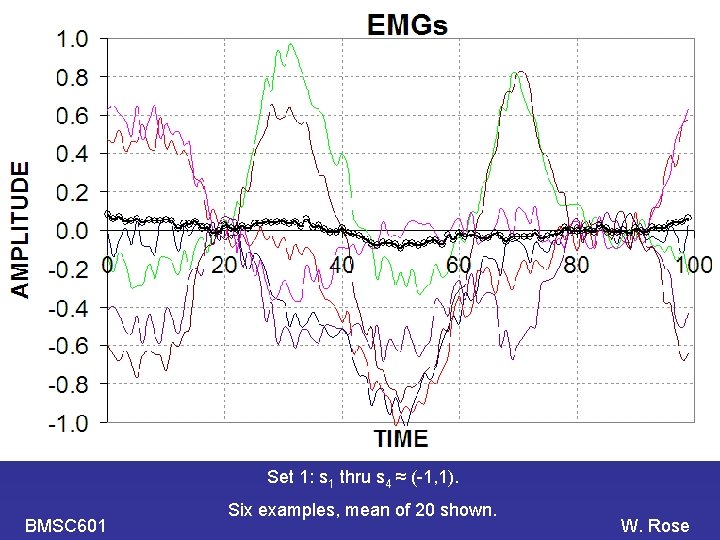
PCA Set 1: s 1 thru s 4 ≈ (-1, 1). BMSC 601 Six examples, mean of 20 shown. W. Rose

PCA Example: running a PCA program in Matlab >> PCA 02 Enter name of text file containing data (. txt will be added): EMG_set 1 100 rows (=times); 20 columns (=different signals) k(90%)=3, k(95%)=4, k(1%)=4 PC weight +- SE %Var accounted for 1 -0. 337 +- 0. 59 46. 4 2 -0. 101 +- 0. 47 30. 2 The program also 3 +0. 162 +- 0. 33 14. 7 4 -0. 046 +- 0. 23 7. 0 makes 3 plots. 5 +0. 011 +- 0. 04 0. 2 6 +0. 006 +- 0. 04 0. 2 1 thru 10 11 thru 20 PC weight +- SE 1 -0. 480 +- 0. 78 -0. 193 +- 0. 91 2 0. 515 +- 0. 72 -0. 716 +- 0. 58 3 -0. 142 +- 0. 31 0. 465 +- 0. 58 4 -0. 148 +- 0. 33 0. 055 +- 0. 33 Variables saved in EMG_set 1_PCA 2. mat. >> BMSC 601 W. Rose
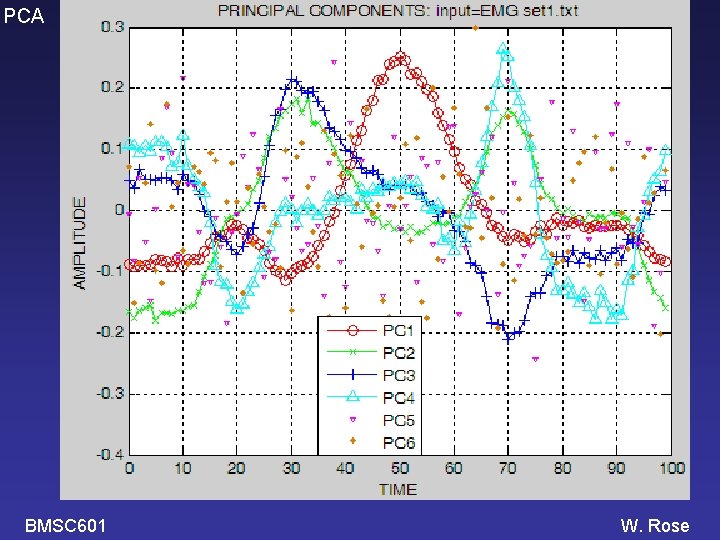
PCA BMSC 601 W. Rose
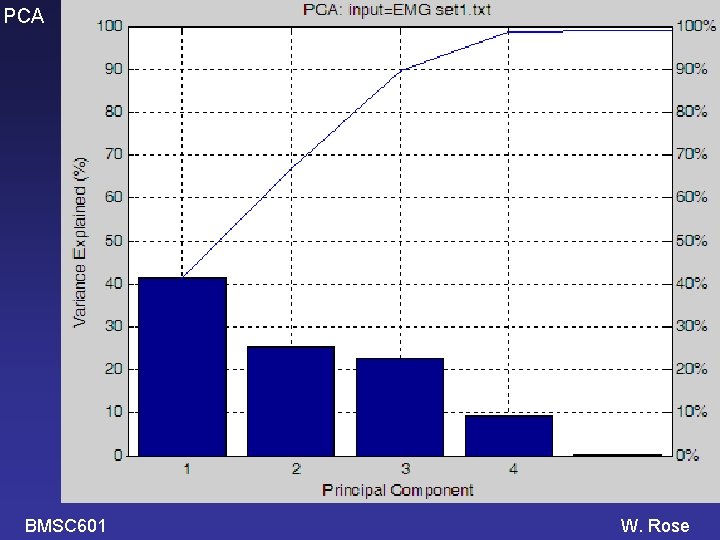
PCA BMSC 601 W. Rose
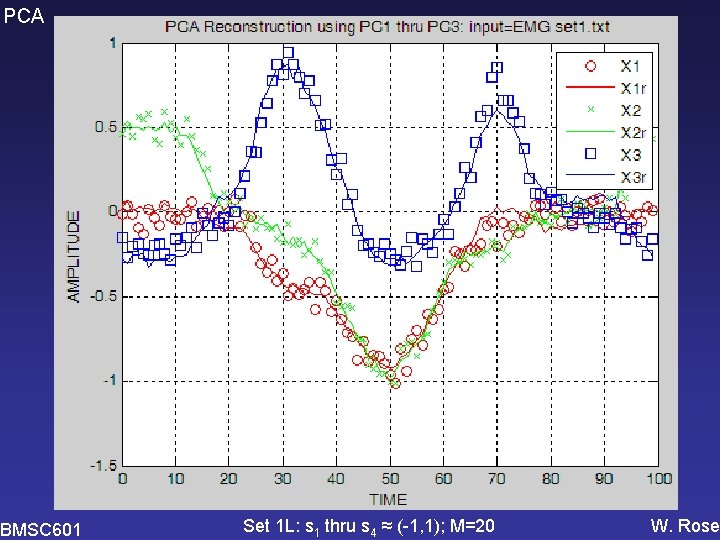
PCA BMSC 601 Set 1 L: s 1 thru s 4 ≈ (-1, 1); M=20 W. Rose
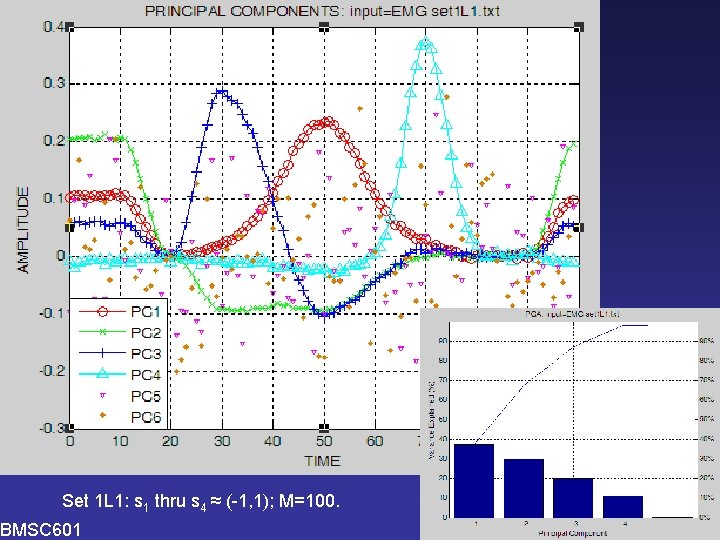
PCA Set 1 L 1: s 1 thru s 4 ≈ (-1, 1); M=100. BMSC 601 W. Rose
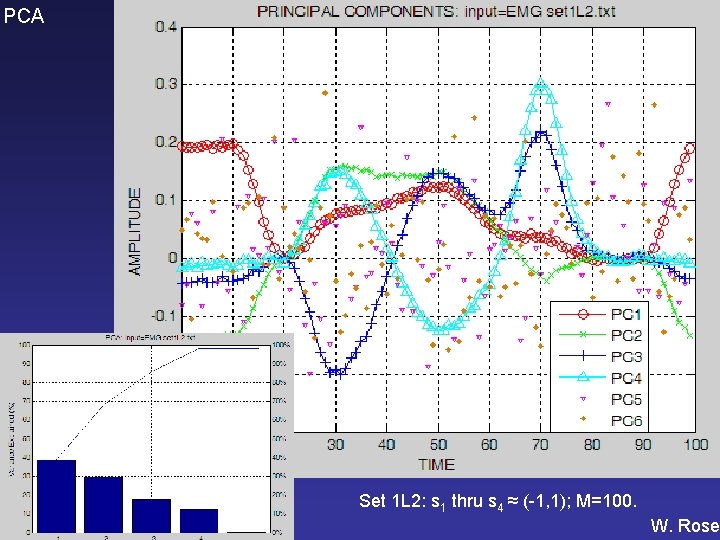
PCA BMSC 601 Set 1 L 2: s 1 thru s 4 ≈ (-1, 1); M=100. W. Rose
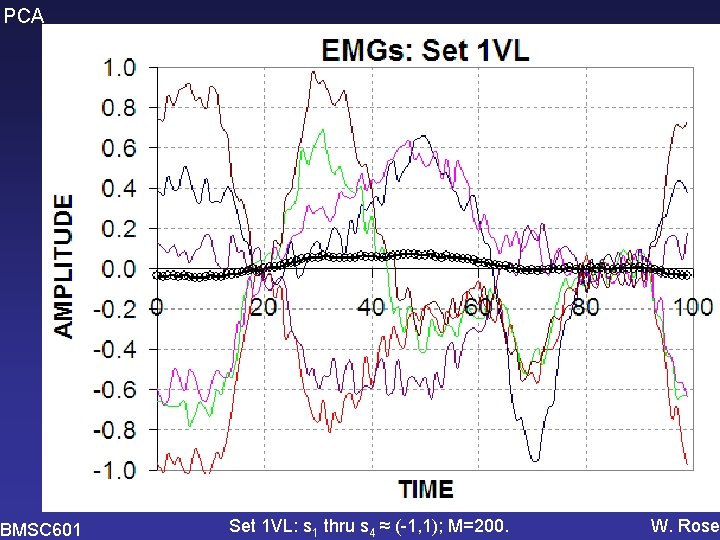
PCA BMSC 601 Set 1 VL: s 1 thru s 4 ≈ (-1, 1); M=200. W. Rose
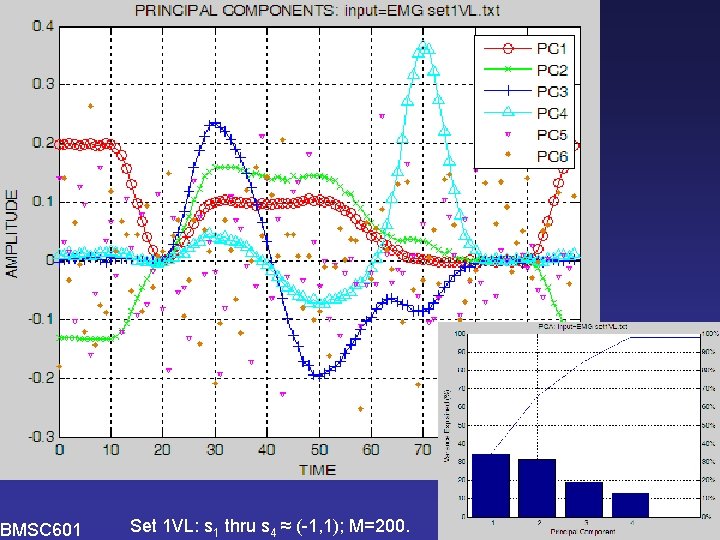
PCA BMSC 601 Set 1 VL: s 1 thru s 4 ≈ (-1, 1); M=200. W. Rose
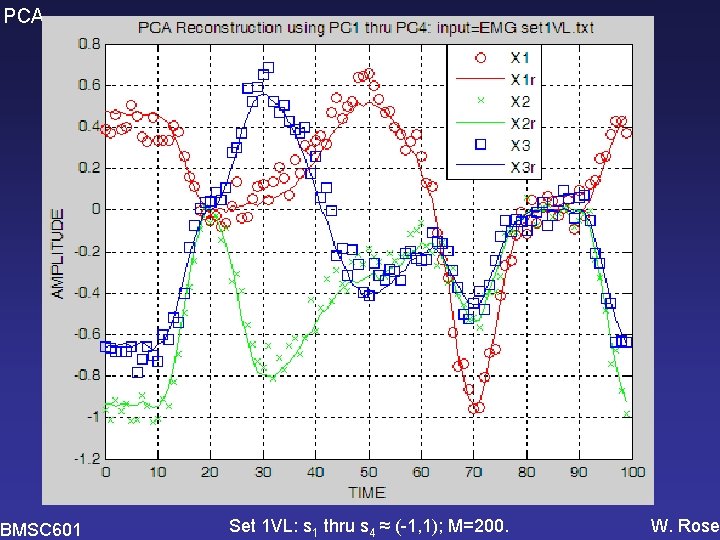
PCA BMSC 601 Set 1 VL: s 1 thru s 4 ≈ (-1, 1); M=200. W. Rose

PCA BMSC 601 Do it again using Hubley. Kozey’s version of KLT (i. e. without de-meaning) [Need to check KLT graphs and software to see if it correctly used the covariance matrix – WCR 2006 -11 -16] W. Rose
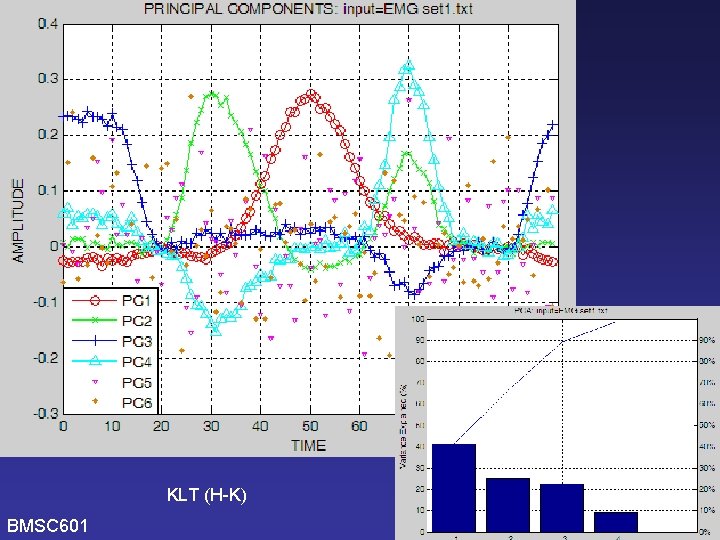
KLT (H-K) BMSC 601 W. Rose
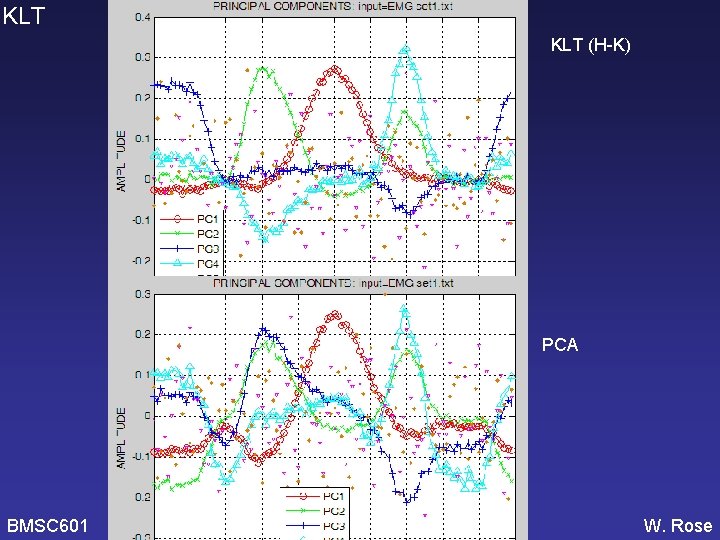
KLT (H-K) PCA BMSC 601 W. Rose
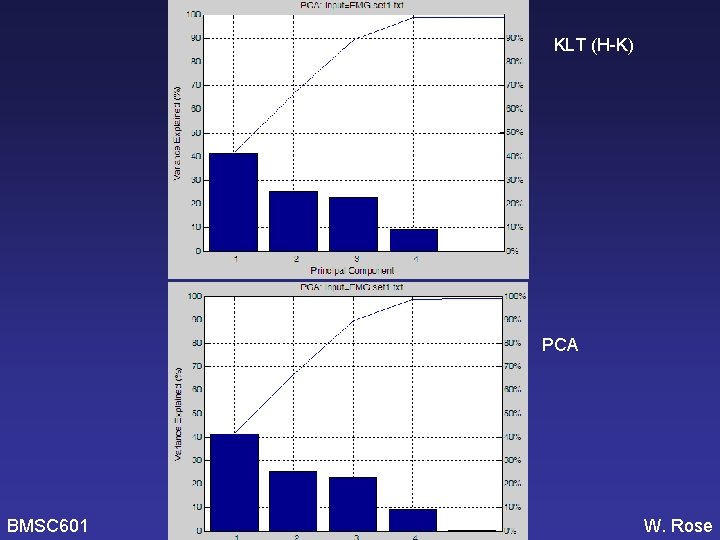
KLT (H-K) PCA BMSC 601 W. Rose
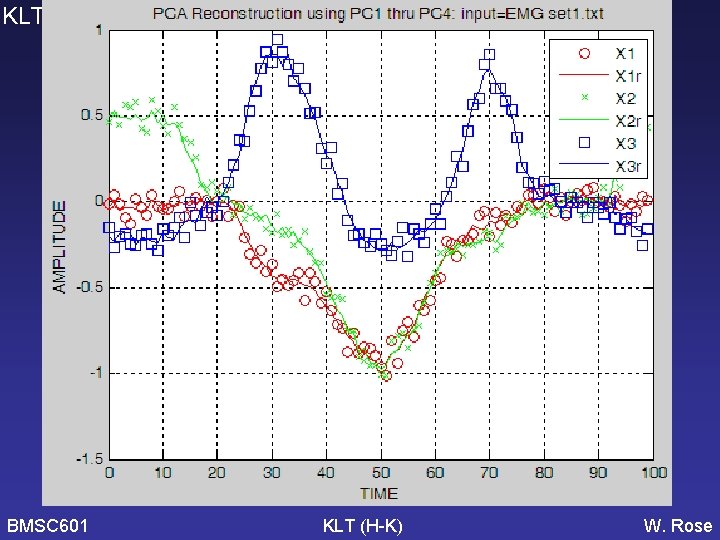
KLT BMSC 601 KLT (H-K) W. Rose
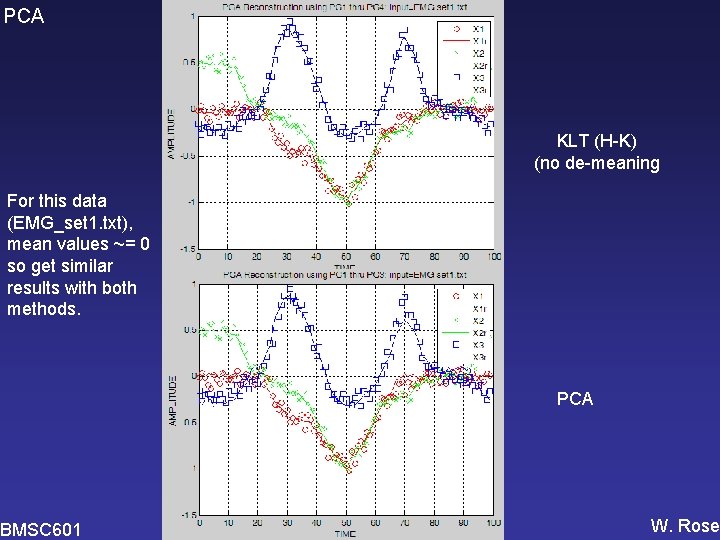
PCA KLT (H-K) (no de-meaning For this data (EMG_set 1. txt), mean values ~= 0 so get similar results with both methods. BMSC 601 PCA W. Rose

PCA BMSC 601 Simulate EMGs using same four components (pulses) as before. Make two populations whose random weights (s 1 to s 4) for the components are randomly distributed over the following intervals: s 1 Group 1 Rec. 1 -20 (1, 3) Group 2 Rec. 21 -40 (1, 3) s 2 (1, 3) (1, 2) s 3 (1, 3) s 4 (1, 3) (1, 4) W. Rose
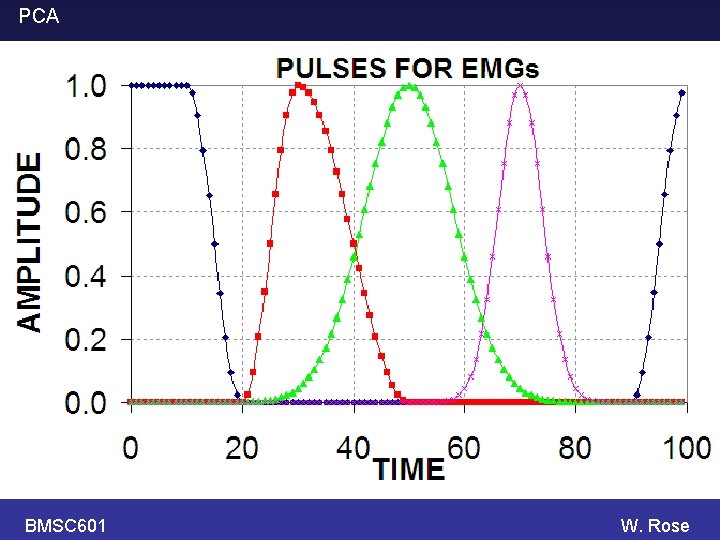
PCA BMSC 601 W. Rose
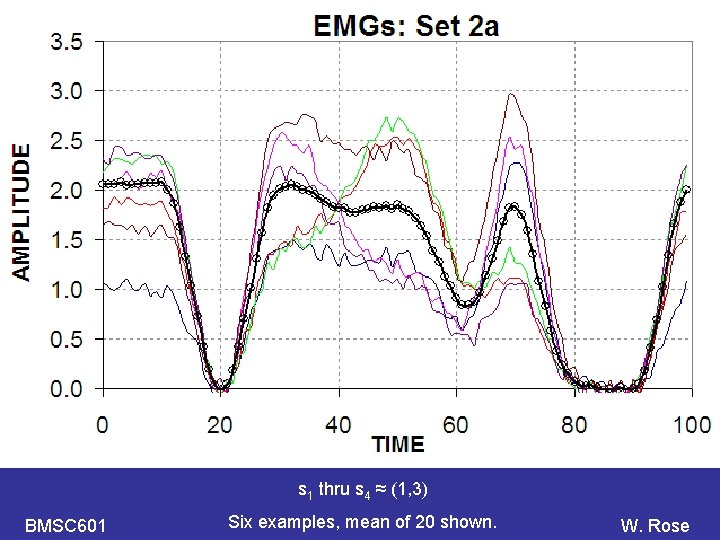
PCA s 1 thru s 4 ≈ (1, 3) BMSC 601 Six examples, mean of 20 shown. W. Rose
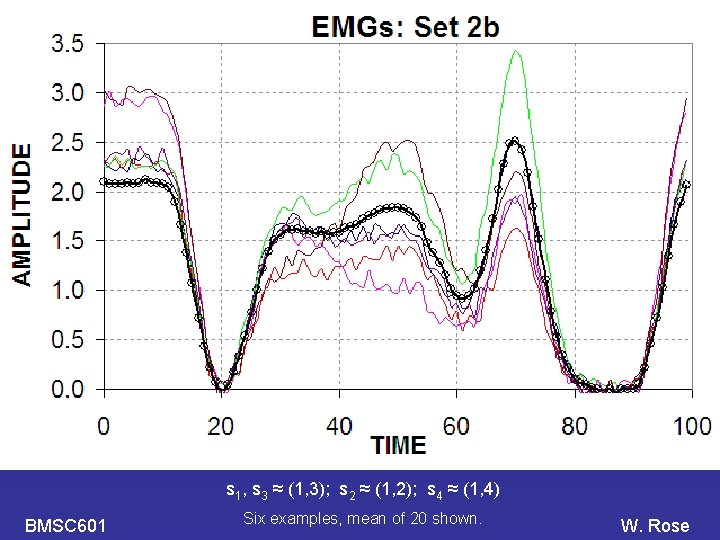
PCA s 1, s 3 ≈ (1, 3); s 2 ≈ (1, 2); s 4 ≈ (1, 4) BMSC 601 Six examples, mean of 20 shown. W. Rose
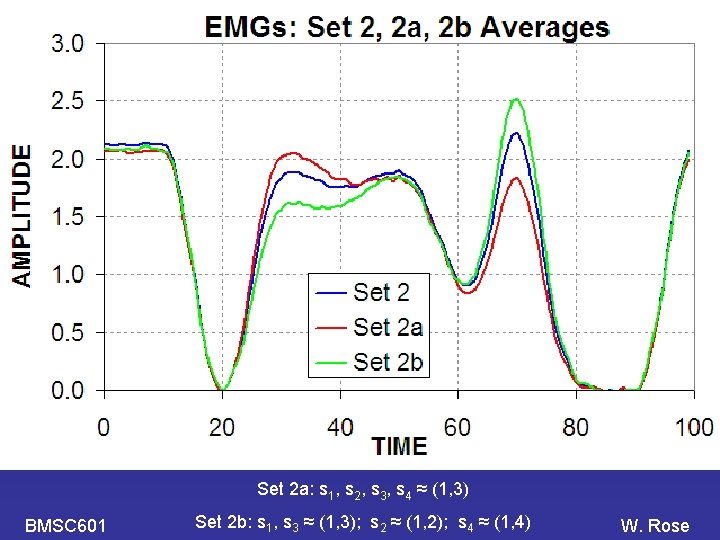
PCA Set 2 a: s 1, s 2, s 3, s 4 ≈ (1, 3) BMSC 601 Set 2 b: s 1, s 3 ≈ (1, 3); s 2 ≈ (1, 2); s 4 ≈ (1, 4) W. Rose

PCA Analysis of Simulated Data Set 2 >> PCA 02 Enter name of text file containing data (. txt will be added): EMG_set 2 100 rows (=times); 40 columns (=different signals) k(90%)=4, k(95%)=4, k(1%)=4 PC weight +- SE %Var accounted for 1 -0. 977 +- 0. 37 40. 4 2 -0. 163 +- 0. 32 28. 8 3 -0. 870 +- 0. 27 20. 8 4 -7. 389 +- 0. 17 8. 0 5 -0. 119 +- 0. 02 0. 1 6 -0. 032 +- 0. 02 0. 1 1 thru 20 21 thru 40 PC weight +- SE 1 -1. 556 +- 0. 49 -0. 397 +- 0. 55 2 -0. 981 +- 0. 42 0. 655 +- 0. 40 3 -0. 192 +- 0. 44 -1. 549 +- 0. 23 4 -7. 554 +- 0. 23 -7. 224 +- 0. 24 Variables saved in EMG_set 2_PCA 2. mat. >> BMSC 601 W. Rose
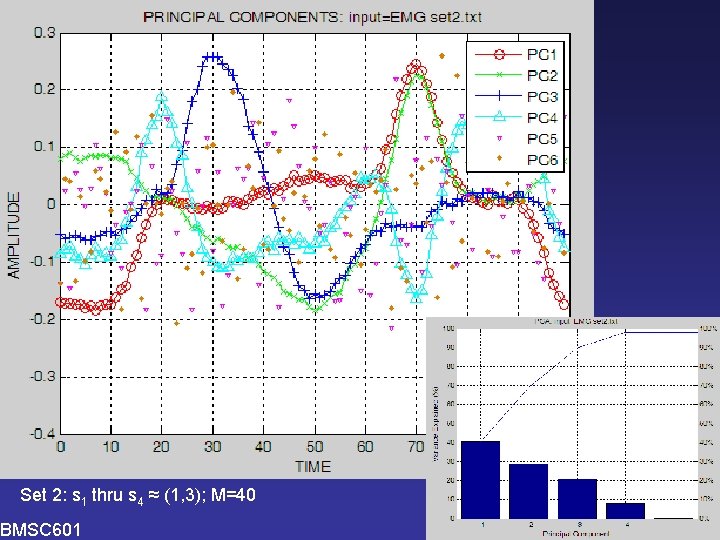
PCA Set 2: s 1 thru s 4 ≈ (1, 3); M=40 BMSC 601 W. Rose
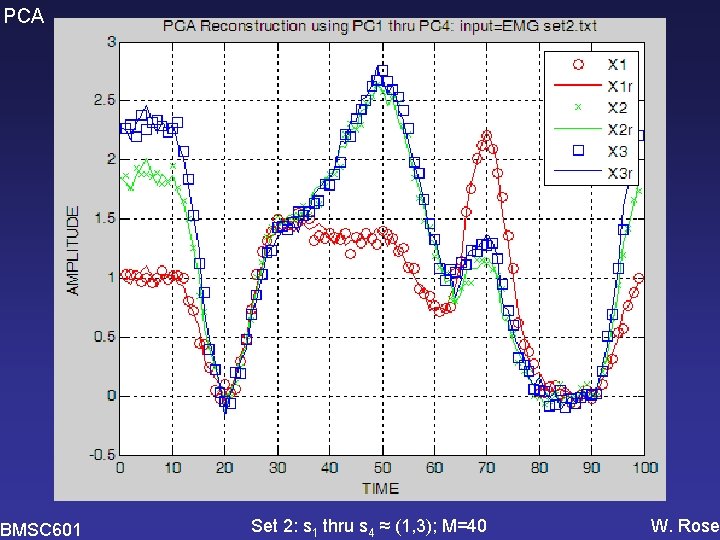
PCA BMSC 601 Set 2: s 1 thru s 4 ≈ (1, 3); M=40 W. Rose

PCA BMSC 601 Do it again using Hubley. Kozey’s version of KLT (i. e. without de-meaning) [Need to check KLT graphs and software to see if it correctly used the covariance matrix – WCR 2006 -11 -16] W. Rose
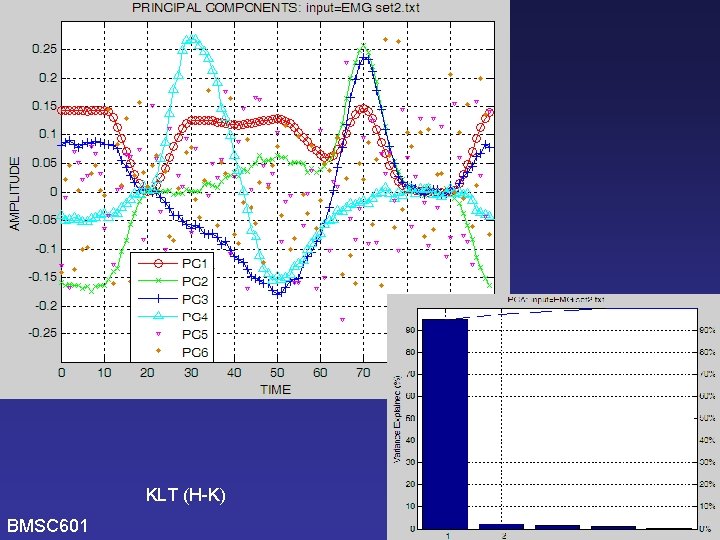
KLT (H-K) BMSC 601 W. Rose
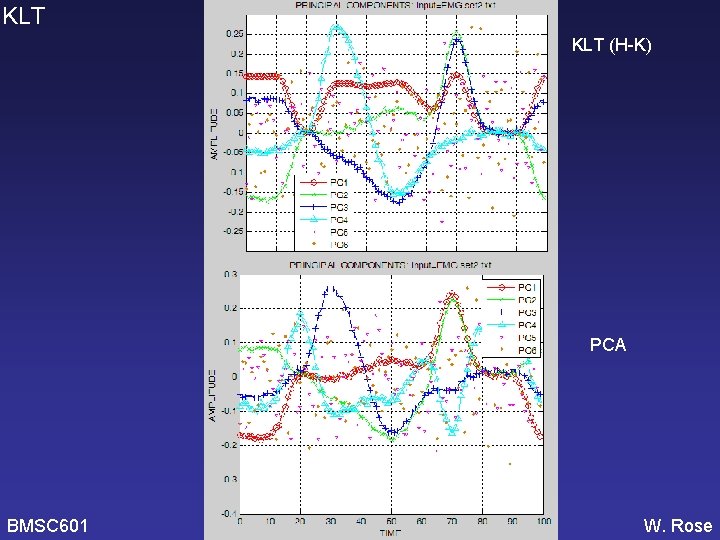
KLT (H-K) PCA BMSC 601 W. Rose
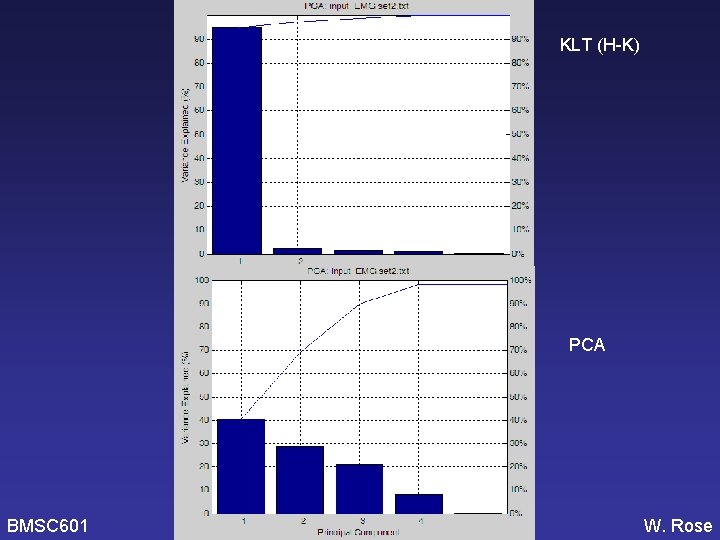
KLT (H-K) PCA BMSC 601 W. Rose
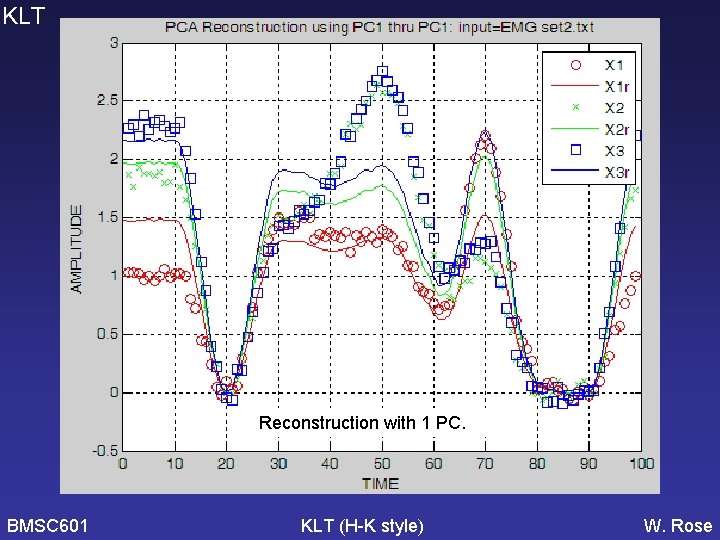
KLT Reconstruction with 1 PC. BMSC 601 KLT (H-K style) W. Rose
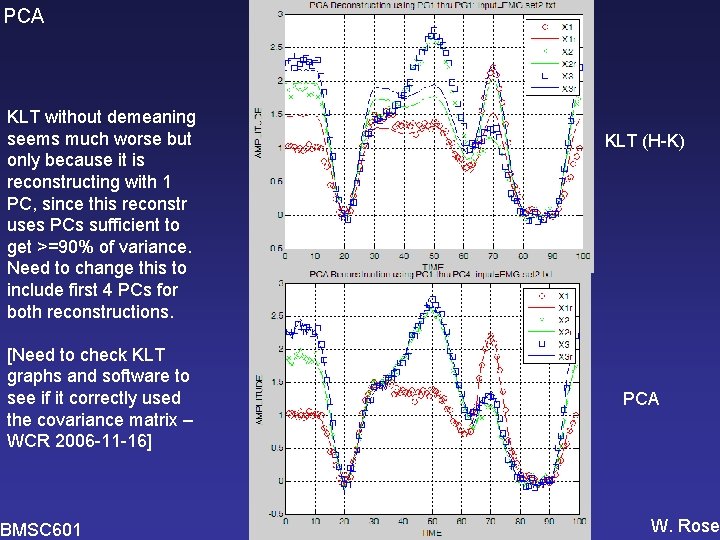
PCA KLT without demeaning seems much worse but only because it is reconstructing with 1 PC, since this reconstr uses PCs sufficient to get >=90% of variance. Need to change this to include first 4 PCs for both reconstructions. [Need to check KLT graphs and software to see if it correctly used the covariance matrix – WCR 2006 -11 -16] BMSC 601 KLT (H-K) PCA W. Rose

PCA New simulated data set: EMG_set 3. Pulses used are same as before. Weights are greater for pulses with greater variances. (All 4 pulses have min=0, max=1, but some are wider so have greater power. ) Will this allow PCA to find PCs that correspond better to the original pulses? s 1=(4, 8) s 2=(2, 6) s 3=(3, 7) s 4 a=(1, 5) (m=40) s 1=(4, 8) s 2=(2, 6) s 3=(3, 7) s 4 b=(0, 4) (m=40) BMSC 601 W. Rose
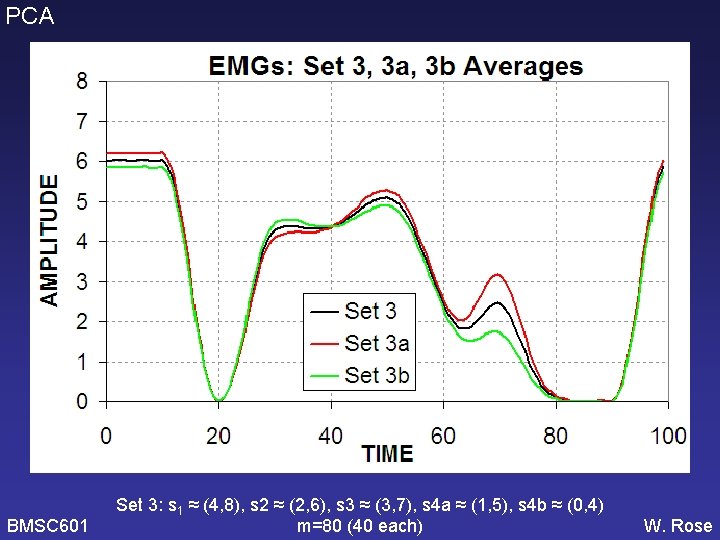
PCA BMSC 601 Set 3: s 1 ≈ (4, 8), s 2 ≈ (2, 6), s 3 ≈ (3, 7), s 4 a ≈ (1, 5), s 4 b ≈ (0, 4) m=80 (40 each) W. Rose
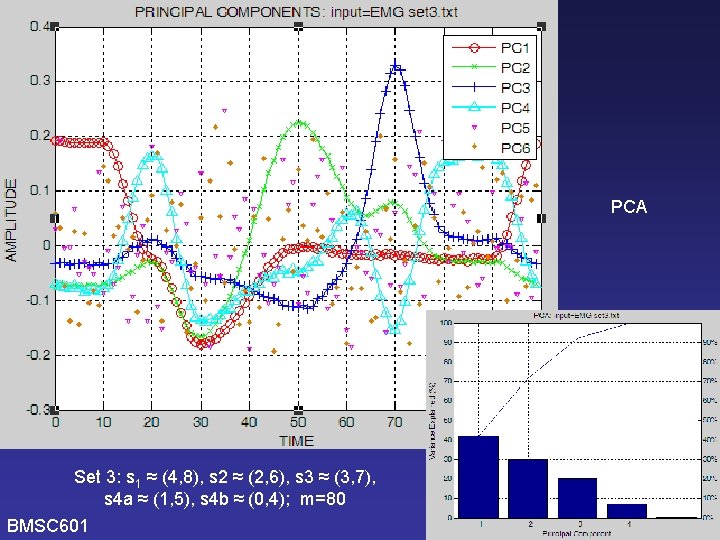
PCA Set 3: s 1 ≈ (4, 8), s 2 ≈ (2, 6), s 3 ≈ (3, 7), s 4 a ≈ (1, 5), s 4 b ≈ (0, 4); m=80 BMSC 601 W. Rose
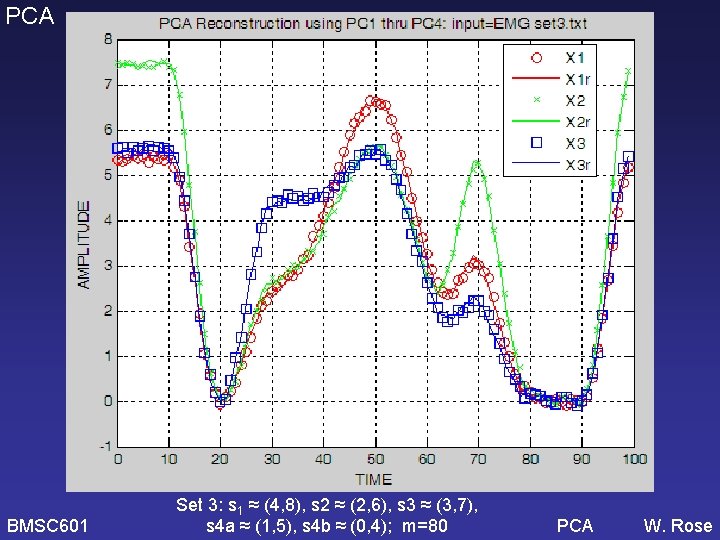
PCA BMSC 601 Set 3: s 1 ≈ (4, 8), s 2 ≈ (2, 6), s 3 ≈ (3, 7), s 4 a ≈ (1, 5), s 4 b ≈ (0, 4); m=80 PCA W. Rose

PCA BMSC 601 Do it again using Hubley. Kozey’s version of KLT (i. e. without de-meaning) [Need to check KLT graphs and software to see if it correctly used the covariance matrix – WCR 2006 -11 -16] W. Rose
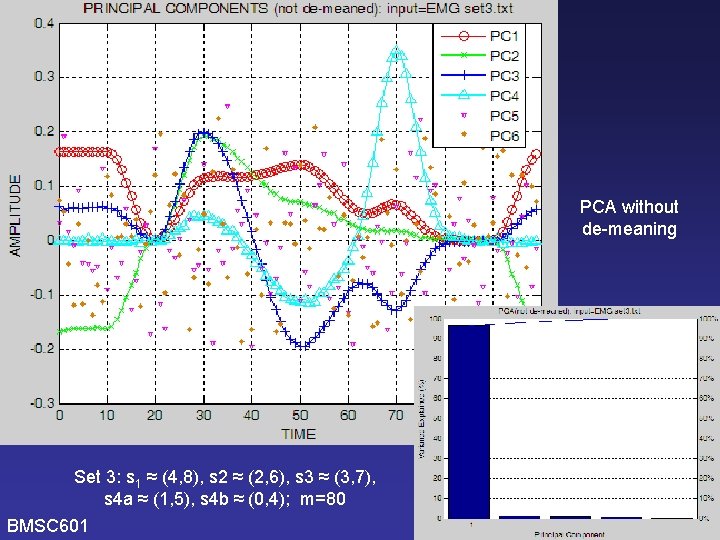
PCA without de-meaning Set 3: s 1 ≈ (4, 8), s 2 ≈ (2, 6), s 3 ≈ (3, 7), s 4 a ≈ (1, 5), s 4 b ≈ (0, 4); m=80 BMSC 601 W. Rose
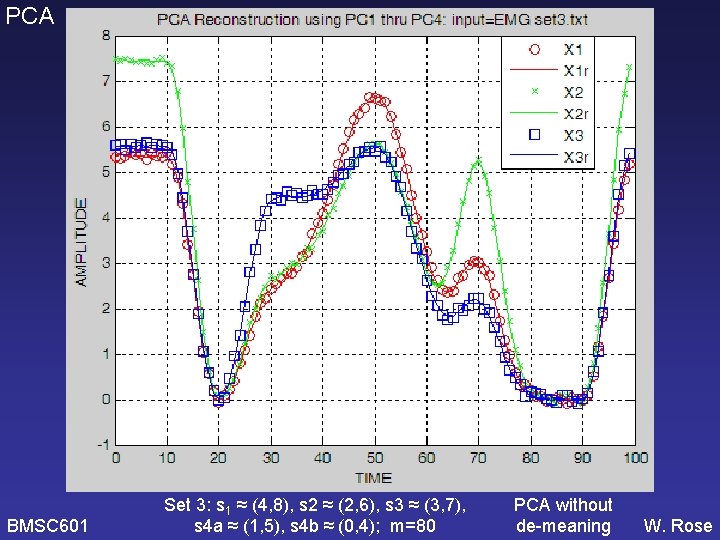
PCA BMSC 601 Set 3: s 1 ≈ (4, 8), s 2 ≈ (2, 6), s 3 ≈ (3, 7), s 4 a ≈ (1, 5), s 4 b ≈ (0, 4); m=80 PCA without de-meaning W. Rose

PCA is analogous to finding a set of orthogonal basis functions which can be added with various weights to reconstruct measured data. When the principal components account for similar amounts (%ages) of total variance, there’s an indeterminacy problem. Example of this problem: find principal axes for COP data that is best fit by a circle. The solution is not unique and the specific vectors chosen will be determined by “random noise” in the data, not by the underlying structure of the model. The basis vectors (1, 0); (0, 1) can account for the variance, but the vectors (1, 1); (-1, 1) will be just as good, and so will the vectors (1, 2); (2, -1). WCR 2010/12
- Slides: 52