Analyzing Gene Relationships for Down Syndrome with Labeled

Analyzing Gene Relationships for Down Syndrome with Labeled Transition Graphs Neha Rungta, Hyrum Carroll, Eric Mercer, Mark Clement, and Quinn Snell Computer Science Department, Brigham Young University, Provo, UT Randall Roper Department of Biology and Indiana University for Regenerative Biology & Medicine, Indiana University-Purdue University, Indianapolis, IN, USA

Goal w How is the SHH gene related to chromosome 21 genes?
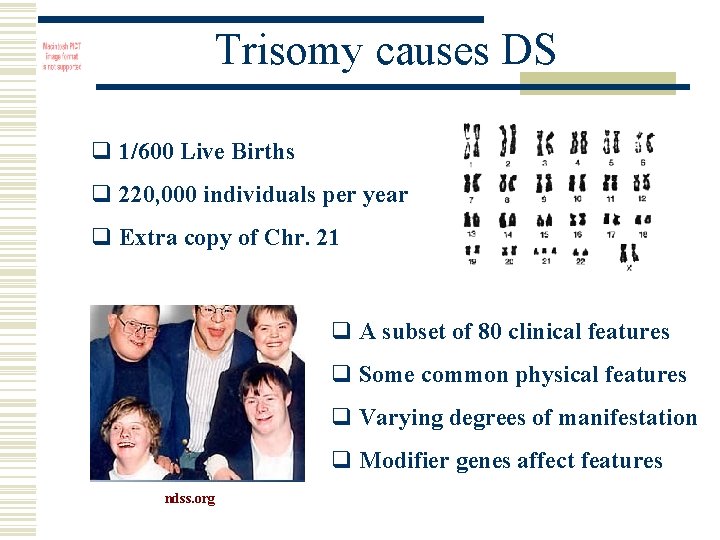
Trisomy causes DS q 1/600 Live Births q 220, 000 individuals per year q Extra copy of Chr. 21 q A subset of 80 clinical features q Some common physical features q Varying degrees of manifestation q Modifier genes affect features ndss. org
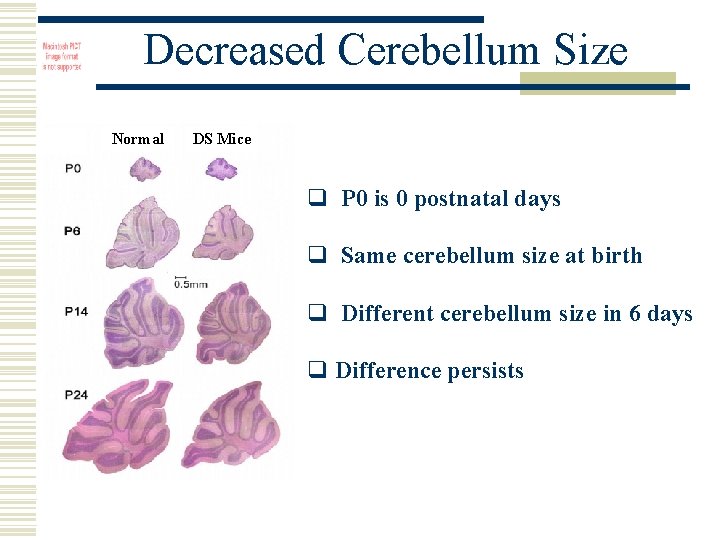
Decreased Cerebellum Size Normal DS Mice q P 0 is 0 postnatal days q Same cerebellum size at birth q Different cerebellum size in 6 days q Difference persists

SHH

Small Jaw

Modifier gene - SHH w SHH is a modifier gene w Affects DS characteristics w Is key factor in a lot of development w How the SHH gene is linked to Chr. 21? w Analyze relationships

Hedgehog signal Pathway Image from genome. jp

State of art pathway analysis w Cyctoscape w Reactome w Bind w Visualize gene regulatory networks w Basic query mechanisms to probe structure

Formal Verification approaches w Heath et al. (`06), Kuwahara et al. (`06), Kwiatkowska et al. (`05), and Dill et al. (`05). w Models a single network in isolation w Predicts pathway behaviors w Models the different reactions and their rates

Our Approach w Abstracts a large amount of information w Connects different networks w Single labeled transition graph w Exhaustive search with a random BFS w Find relations between SHH and Chr. 21

Kegg Database w 1 st source of data w Largest and most comprehensible database w 175 pathways 12, 000 genes - human genome w Pictorial and XML representation

Pub. Med Abstracts w 2 nd source of data w Premier journal for bio-medical articles w Bunescu et al. extract gene relationships using natural language processing algorithm

Graph Construction w Take a gene regulatory pathway w Abstract away the reactions and compounds w Create intra-pathway gene connections w Create inter-pathway gene connections w Add Pub. Med relations
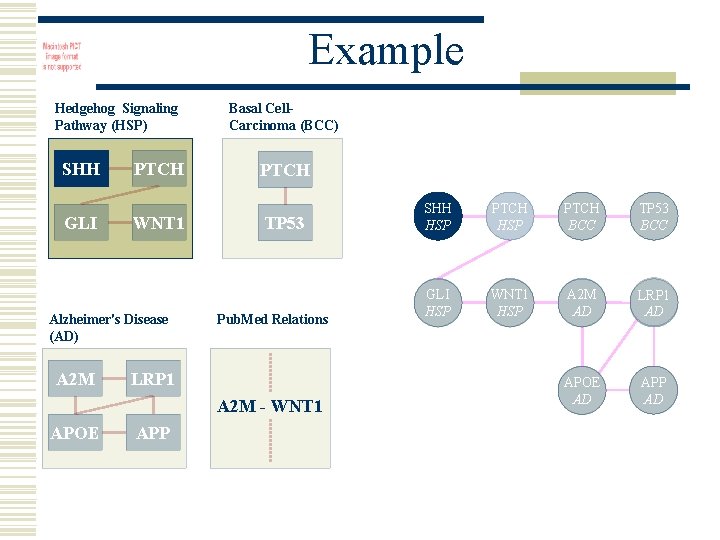
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
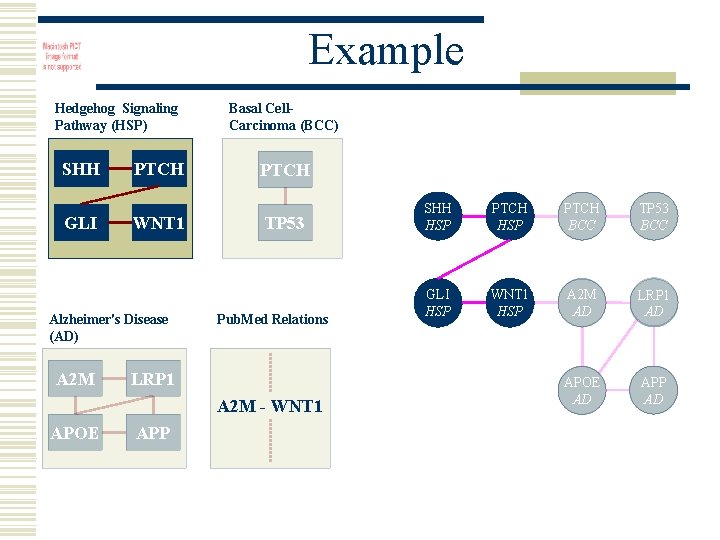
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
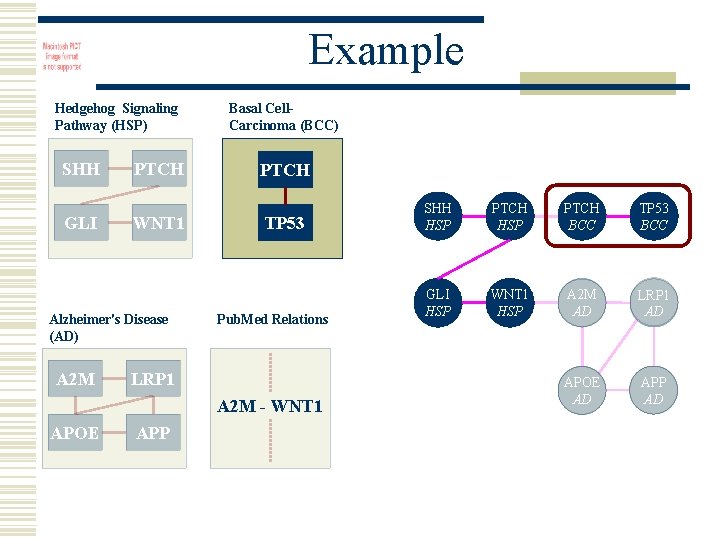
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
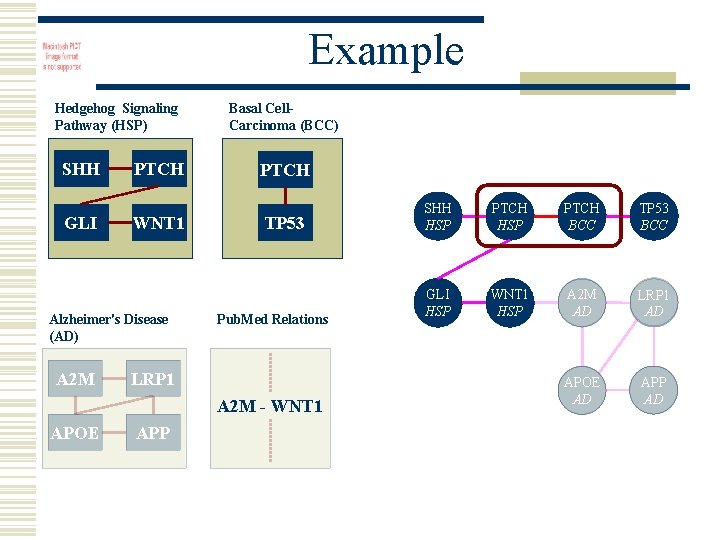
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
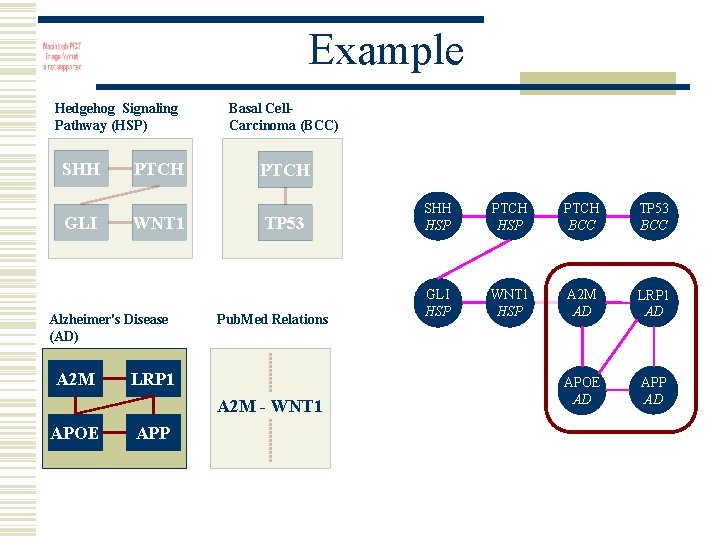
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
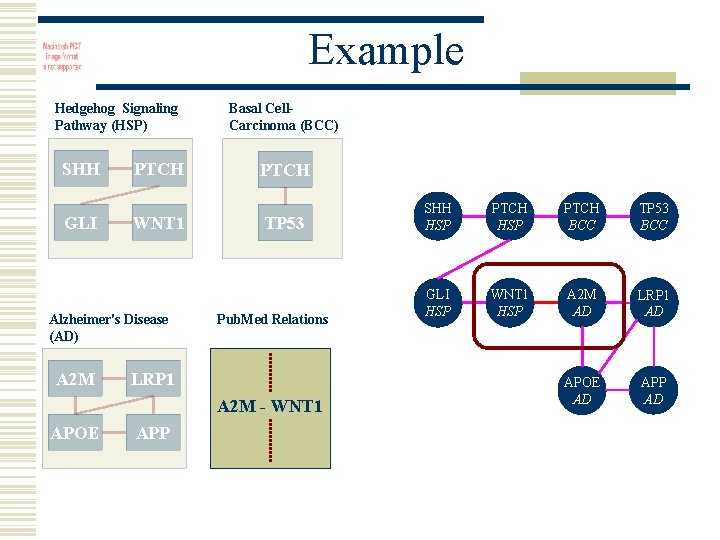
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
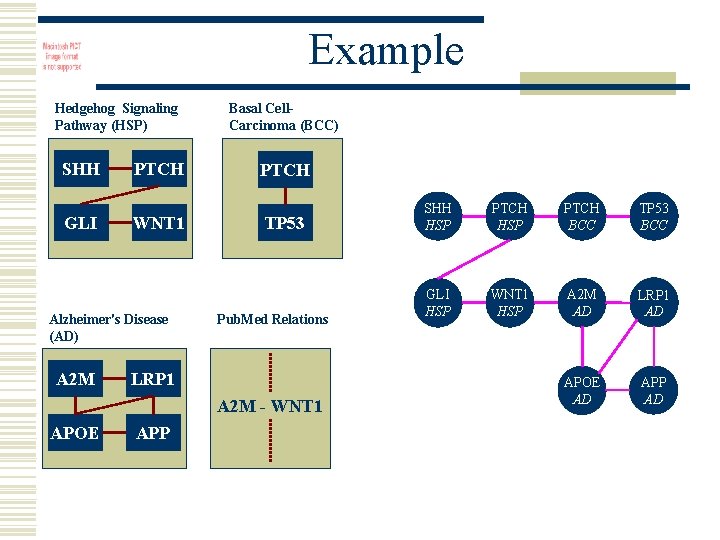
Example Hedgehog Signaling Pathway (HSP) SHH GLI PTCH WNT 1 Alzheimer's Disease (AD) A 2 M Basal Cell. Carcinoma (BCC) PTCH TP 53 Pub. Med Relations LRP 1 A 2 M - WNT 1 APOE APP SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD

Analysis w 12, 000 nodes w 1 million edges w 90% connections due to common genes w Large unique shortest paths w Random BFS allows sampling
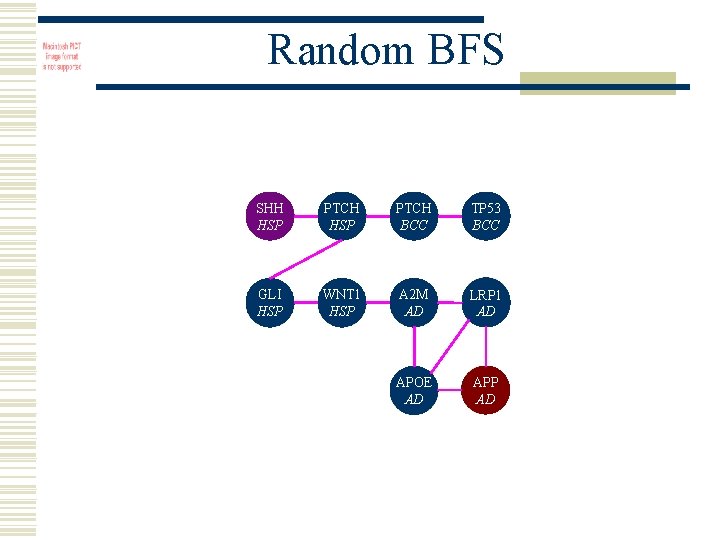
Random BFS SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD
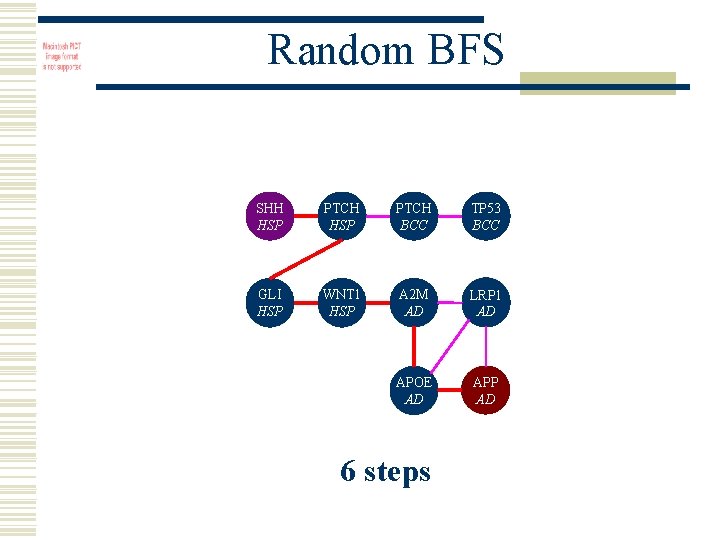
Random BFS SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD 6 steps
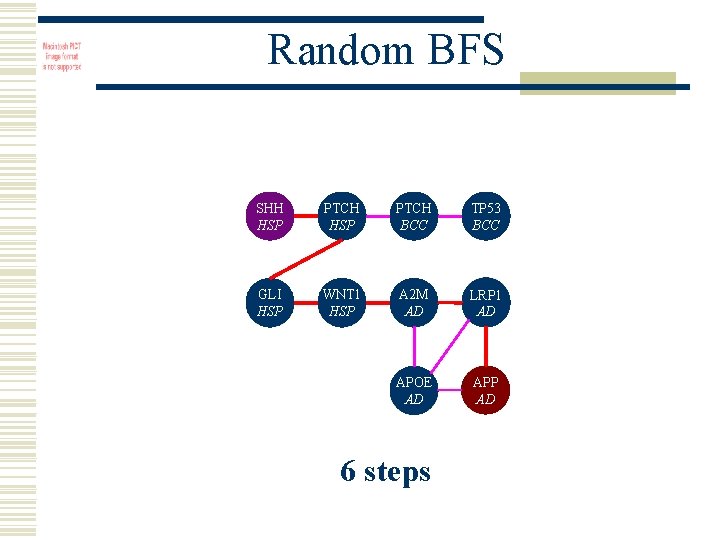
Random BFS SHH HSP PTCH BCC TP 53 BCC GLI HSP WNT 1 HSP A 2 M AD LRP 1 AD APOE AD APP AD 6 steps
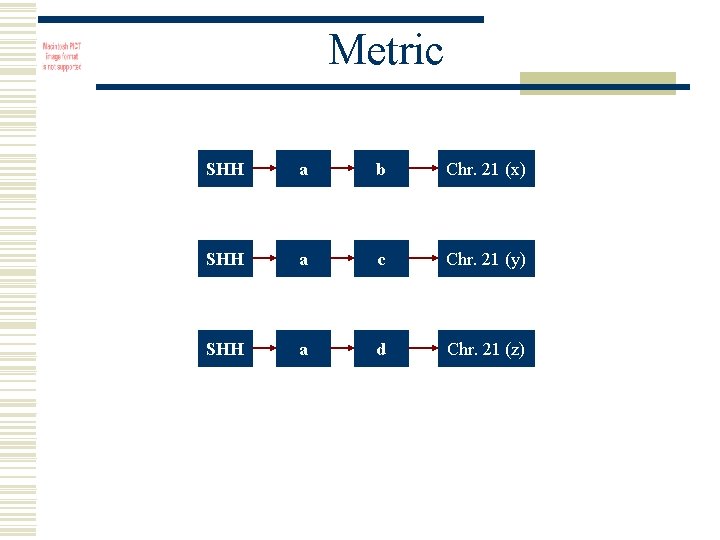
Metric SHH a b Chr. 21 (x) SHH a c Chr. 21 (y) SHH a d Chr. 21 (z)
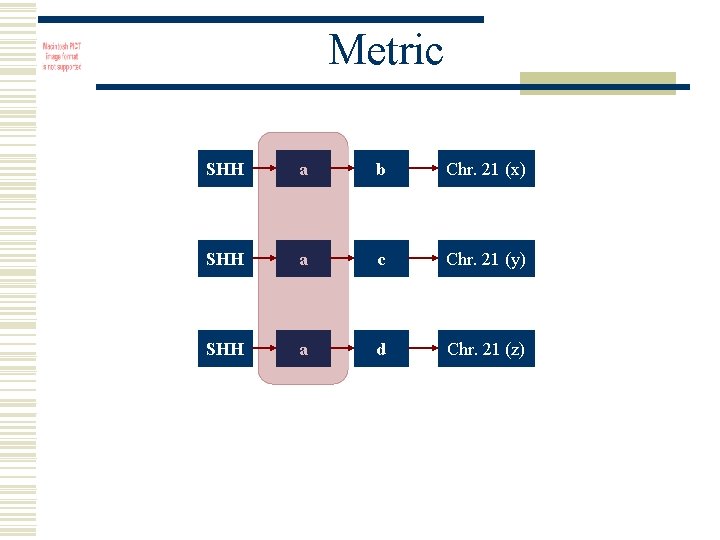
Metric SHH a b Chr. 21 (x) SHH a c Chr. 21 (y) SHH a d Chr. 21 (z)
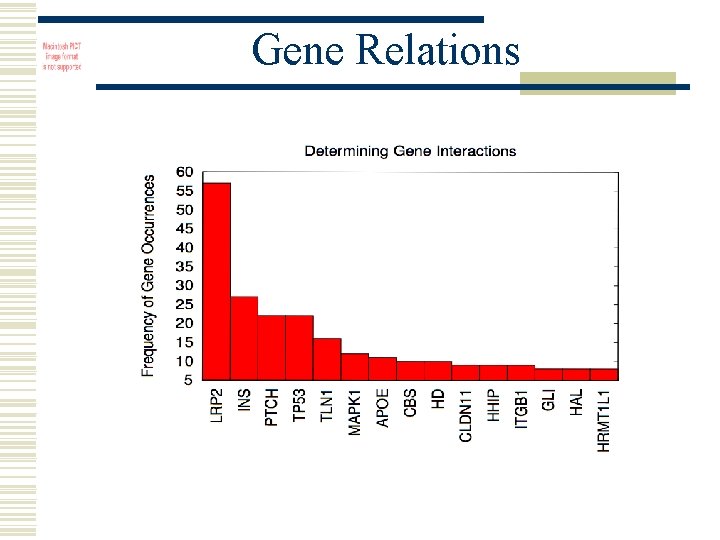
Gene Relations
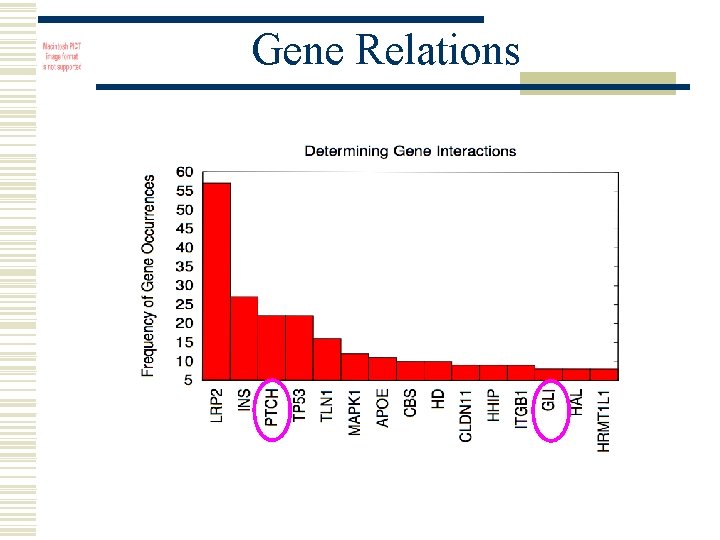
Gene Relations
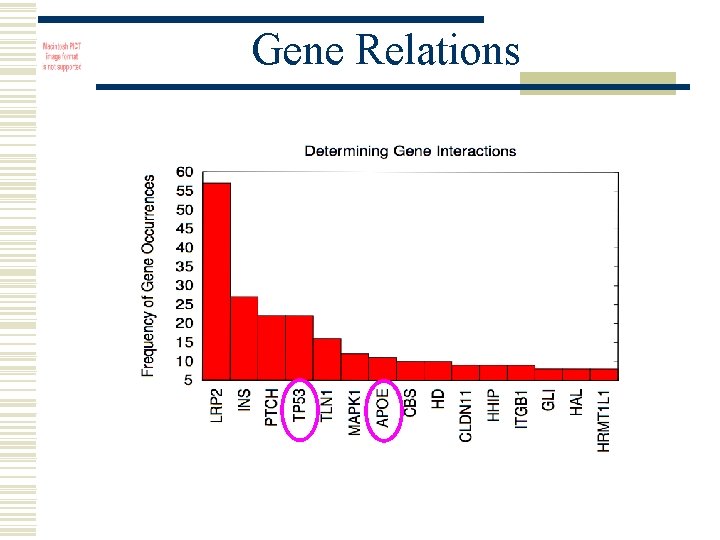
Gene Relations
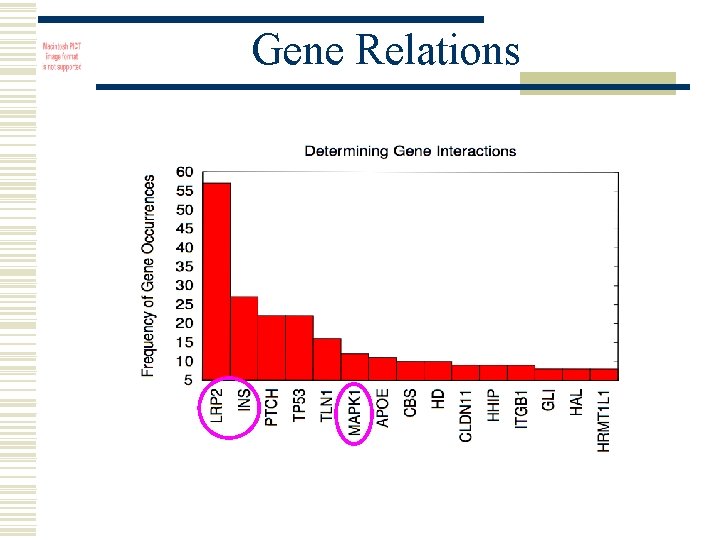
Gene Relations

Conclusions & Future Work w New technique to combine different gene regulatory networks with a transition graph w Discovers new relations w More refined analysis with transition graphs w Use mu-calculus to pose interesting questions

Questions ?
- Slides: 33