Analyzing Base Pair Differences Between Subjects with Significantly

Analyzing Base Pair Differences Between Subjects with Significantly Different T-Cell Rate Alex George J’aime Moehlman Amanda Wavrin Loyola Marymount University March 2, 2010

Outline • • • Introduction Materials/Methods Results Discussion References

Introduction to HIV-1 • HIV-1 relation to CD 4 T-Cells. • Significance of the Env gene – gp 120 • Studies suggest adaptation leads to pathogenicity of HIV-1.

Our Proposed Question • Is there a specific base pair difference between the subjects with a high rate of CD 4 T-Cell decline versus the subjects with an increase in CD 4 T-Cells?

1 st Hypothesis • H 1: There will be a specific nucleotide difference between each group that is responsible for CD 4 T-Cell increase or decline. • Two Groups: – Group 1 (Decline): Subjects 10, 11, 15 – Group 2 (Increase): Subjects 6, 12 13

Methods for Hypothesis 1 1. We selected all clones from all subjects from Visit 4. 2. Unrooted Tree of all Six Subjects. 3. Multiple Sequence Alignment within groups on Biology Workbench. 4. Compared similarities within Group 1 to similarities within Group 2.
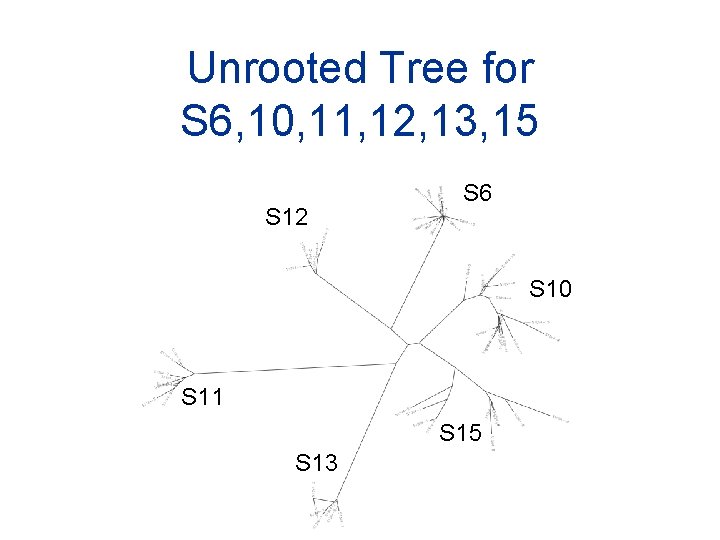
Unrooted Tree for S 6, 10, 11, 12, 13, 15 S 12 S 6 S 10 S 11 S 15 S 13

Example Comparison between S 10 and S 6 S 10, V 4 -1 S 6, V 4 -1

Results • The similar sequences were identical for both groups. • Our hypothesis is rejected: – H 0: There was no constant DNA sequence in Group 1 that differed from sequences in Group 2. • These results led us to formulate a second hypothesis.

2 nd Hypothesis • H 2: The range of difference in Group 1 will be larger than the range of difference for Group 2. • Two Groups: – Group 1 (Decline): Subjects 10, 11, 15 – Group 2 (Increase): Subjects 6, 12, 13

Methods for Hypothesis 2 1. Observed differences from original Unrooted Tree. 2. Use Clustdist tool in Biology Workbench to generate distance matrices for each group. 3. Multiplied Min. and Max. by 285 to calculate raw number of differences.
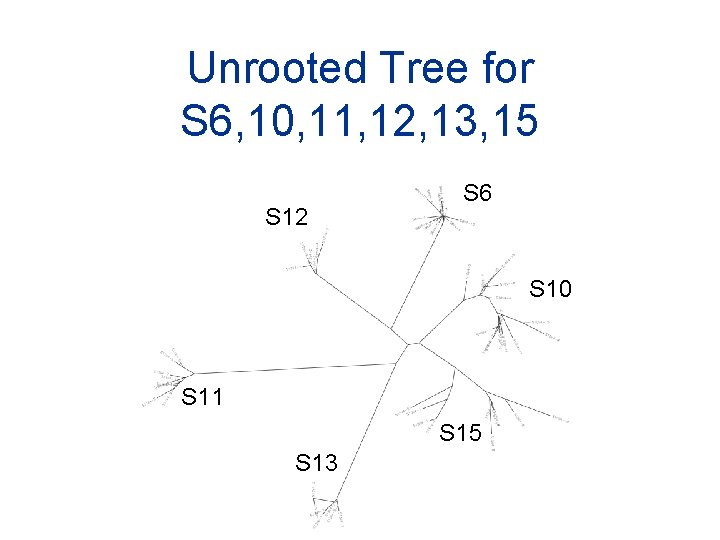
Unrooted Tree for S 6, 10, 11, 12, 13, 15 S 12 S 6 S 10 S 11 S 15 S 13
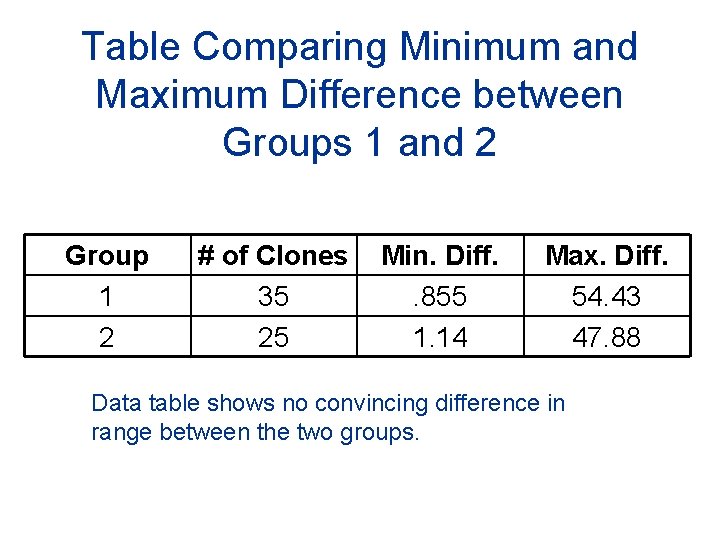
Table Comparing Minimum and Maximum Difference between Groups 1 and 2 Group 1 2 # of Clones 35 25 Min. Diff. . 855 1. 14 Max. Diff. 54. 43 47. 88 Data table shows no convincing difference in range between the two groups.

Results • The range difference of Group 1 was larger than Group 2, but is not convincing. • In the V 3 region, the similarities within Group 1 were identical to the similarities within Group 2.

Discussion • The V 3 region does not have the most adaptive events. • The base pair differences could lie in a different region of the gene. • Future Research: – Analyze other regions of the gene in a similar manner to find differences in base pairs.

References – Williamson, Scott. 2003. Adaptation in the env Gene of HIV-1 and Evolutionary Theories of Disease Progression. Molecular Biology Evolution. 20(8): 1318– 1325 – Markham RB, Wang WC, Weisstein AE, Wang Z, Munoz A, Templeton A, Margolick J, Vlahov D, Quinn T, Farzadegan H, and Yu XF. Patterns of HIV-1 evolution in individuals with differing rates of CD 4 T cell decline. Proc Natl Acad Sci U S A 1998 Oct 13; 95(21) 12568 -73. pmid: 9770526.
- Slides: 16