Analysis of the Unknown Gene Bradi 1 g

Analysis of the Unknown Gene Bradi 1 g 67840. 1 CHM Taylor Carmichael and Vicente Gris

Why Brachypodium distachyon? ● Functional model for genomic analyses ● Relatively small genome ○ Attractive for genomic analyses ● Small in stature and can be grown rapidly ● Similar to common crops ● However, the B. distachyon is still far too extensive to individually distinguish one gene https: //genome. jgi. doe. gov/portal/brach y/brachy. home. html

Purpose We sought to define and distinguish key characteristics about our unknown B. distachyon gene, Bradi 1 g 67840. 1. Using molecular genetics and recombinant DNA searches, we were able to acquire information about our gene’s expression pattern and make conclusions regarding its function.

Working Map Acquisition ● Six-frame translation for our raw nucleotide sequence ● The FGENESH software was used as an ab initio to predict the coding portions ● Two exon regions were observed Gene map of the unknown gene, Bradi 1 g 67840. 1

Homology Based Annotation ● Basic Local Alignment Tool software, or BLAST, was used to identify the expressed sequence tags (ESTs) that match the sequence of our unknown gene ○ A full length c. DNA sequence was matched to our unknown gene using the Nucleotide collection (nr/nt) database on the BLAST software BLASTx distribution of a highly conserved protein sequence, LRR

Bradi 1 g 67840. 1 Function Analysis ● BLASTx and BLASTn software suggested species that share a homologous sequence with our gene ○ Oryza sativa Function: ATP binding enabling. NBS-LRR resistance gene that functions by a gene-for-gene protein-ligand model between a specific resistance (R) gene and an associated pathogen avirulence (Avr) gene. Involved in a pathogen defense mechanism
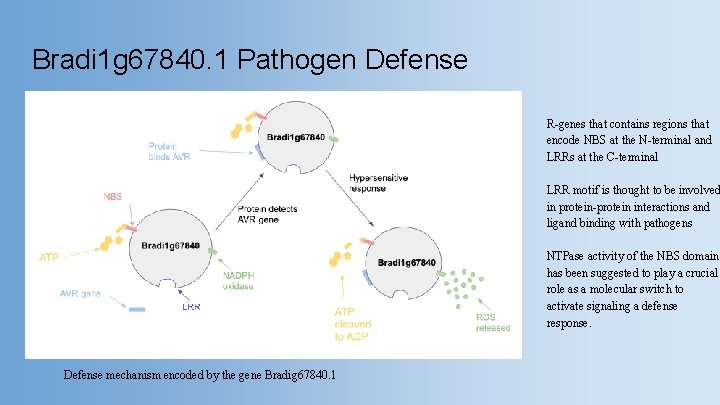
Bradi 1 g 67840. 1 Pathogen Defense R-genes that contains regions that encode NBS at the N-terminal and LRRs at the C-terminal LRR motif is thought to be involved in protein-protein interactions and ligand binding with pathogens NTPase activity of the NBS domain has been suggested to play a crucial role as a molecular switch to activate signaling a defense response. Defense mechanism encoded by the gene Bradig 67840. 1

Mutant Analysis ● The position and effect of high and moderate impact mutations were identified using the Phytozyme computer software ● High impact mutation: Na. N 184 ○ ○ Located on exon 2 Nonsense mutation changing the codon G to A Relative positions of five high and moderate impact mutations on the unknown gene, Bradi 1 g 67840. 1

PCR Analysis ● Primers were identified using Primer 3 software ● Primers located on either side of Na. N 184
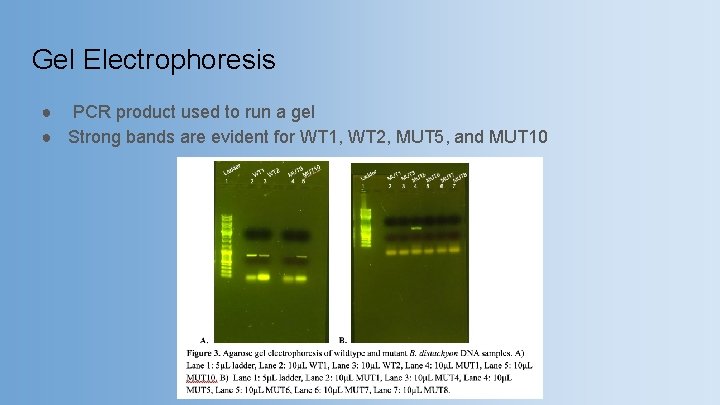
Gel Electrophoresis ● PCR product used to run a gel ● Strong bands are evident for WT 1, WT 2, MUT 5, and MUT 10
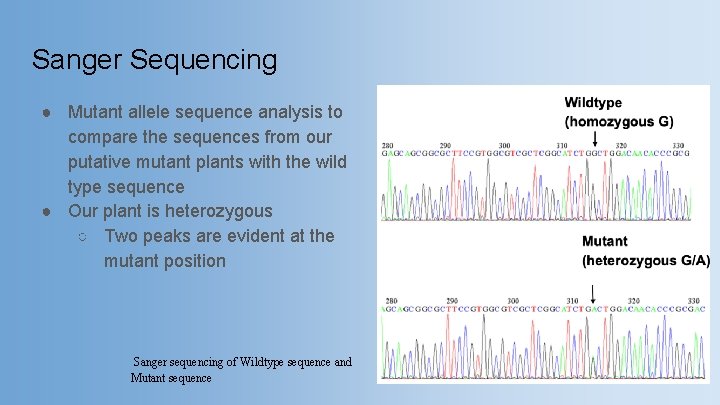
Sanger Sequencing ● Mutant allele sequence analysis to compare the sequences from our putative mutant plants with the wild type sequence ● Our plant is heterozygous ○ Two peaks are evident at the mutant position Sanger sequencing of Wildtype sequence and Mutant sequence
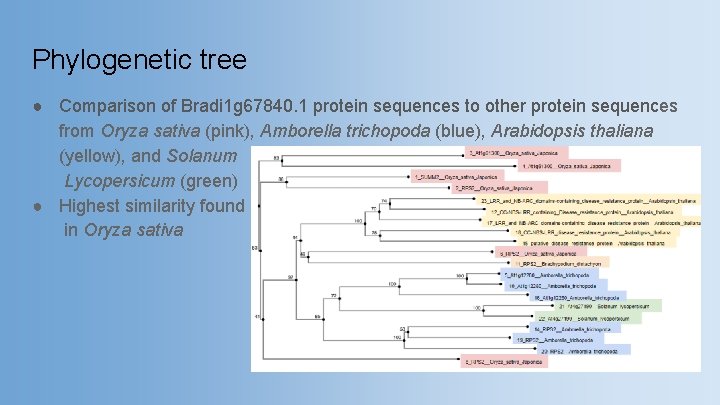
Phylogenetic tree ● Comparison of Bradi 1 g 67840. 1 protein sequences to other protein sequences from Oryza sativa (pink), Amborella trichopoda (blue), Arabidopsis thaliana (yellow), and Solanum Lycopersicum (green) ● Highest similarity found in Oryza sativa
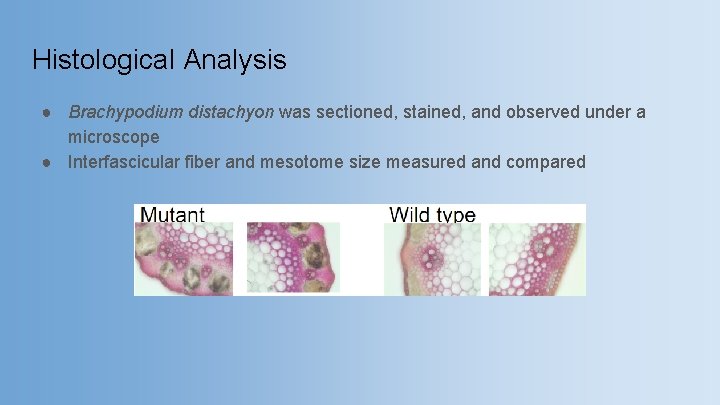
Histological Analysis ● Brachypodium distachyon was sectioned, stained, and observed under a microscope ● Interfascicular fiber and mesotome size measured and compared
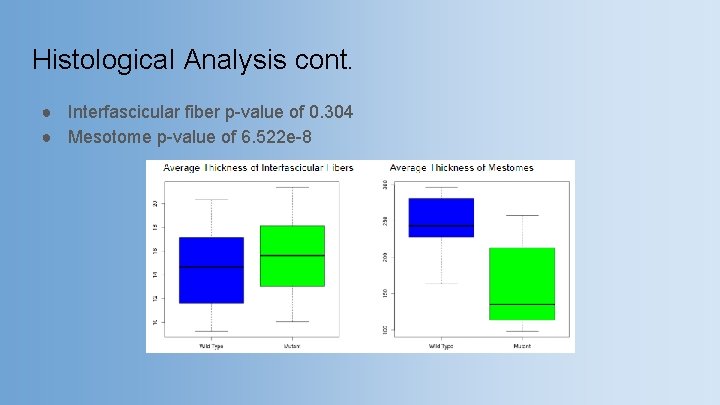
Histological Analysis cont. ● Interfascicular fiber p-value of 0. 304 ● Mesotome p-value of 6. 522 e-8
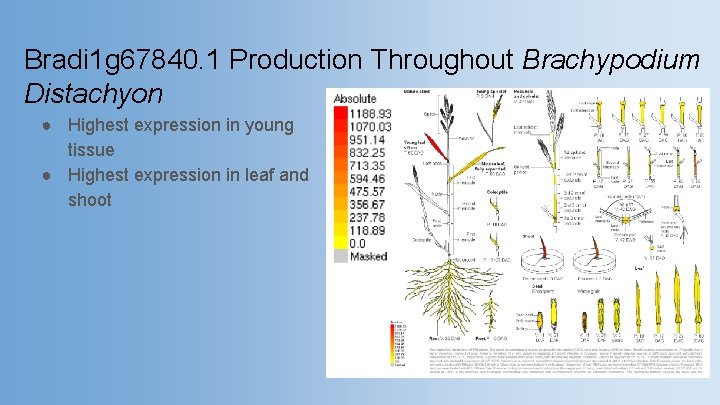
Bradi 1 g 67840. 1 Production Throughout Brachypodium Distachyon ● Highest expression in young tissue ● Highest expression in leaf and shoot
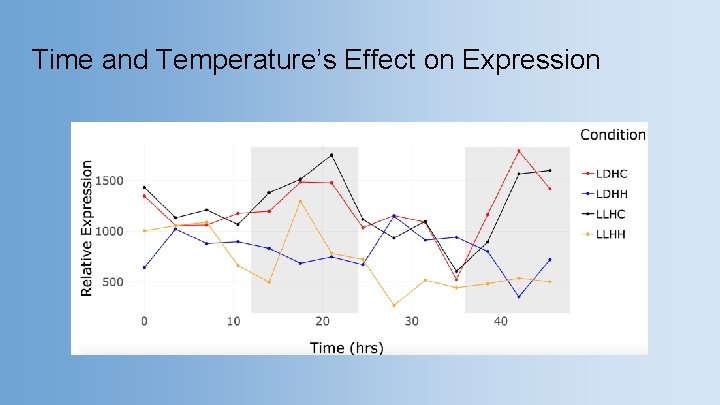
Time and Temperature’s Effect on Expression
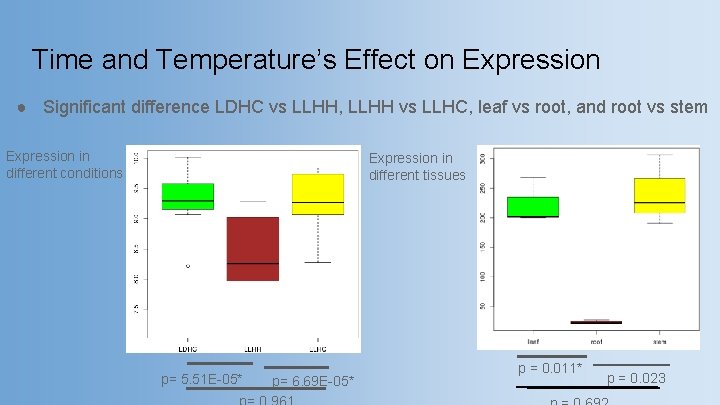
Time and Temperature’s Effect on Expression ● Significant difference LDHC vs LLHH, LLHH vs LLHC, leaf vs root, and root vs stem Expression in different conditions Expression in different tissues p= 5. 51 E-05* p= 6. 69 E-05* p = 0. 011* p = 0. 023
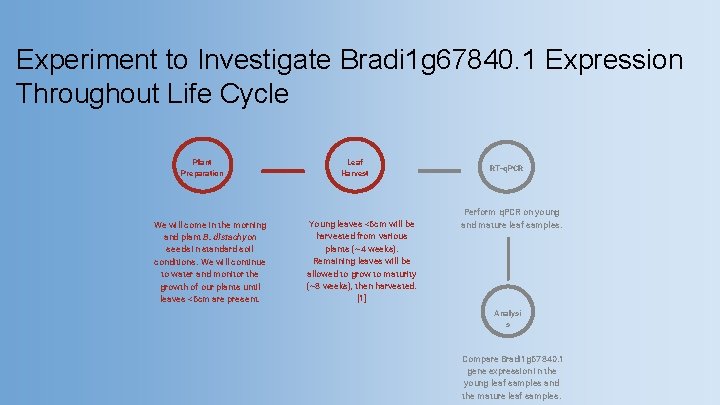
Experiment to Investigate Bradi 1 g 67840. 1 Expression Throughout Life Cycle Plant Preparation We will come in the morning and plant B. distachyon seeds in standard soil conditions. We will continue to water and monitor the growth of our plants until leaves <6 cm are present. Leaf Harvest Young leaves <6 cm will be harvested from various plants (~4 weeks). Remaining leaves will be allowed to grow to maturity (~8 weeks), then harvested. [1] RT-q. PCR Perform q. PCR on young and mature leaf samples. Analysi s Compare Bradi 1 g 67840. 1 gene expression in the young leaf samples and the mature leaf samples.

Primer Design for q. PCR ● Quant. Prime used to develop primer for q. PCR ● Forward primer on exon 1 and reverse primer on exon 2

Conclusions 1. Gene function 2. Gene phylogeny and conservation 3. Gene mutant and consequences 4. Gene expression conditions 5. Experiment for gene expression conditions

References 1. 2. 3. 4. Lucas S. J. Bastas K. , Budak H. (2014) Exploring the interaction between small RNAs and R genes during Brachypodium Response to Fusarium culmorum infection. Science Direct. 536(2): 254 -264 pp. Jia Y. , Mc. Adams S. A. , Bryan G. T. , Hershey H. P. , Valent B. (2000). Direct interaction of resistance gene and avirulence gene products confers rice blast resistance. The EMBO Journal. 19(5): 4004 -4014 pp. Tan S. , Wu S. (2012). Genome Wide Analysis of Nucleotide-Binding Site Disease Resistance Genes in Brachypodium distachyon. Hindawi Publishing Corporation. 1 -12 pp. Grigoriev IV, Nordberg H, Shabalov I, Aerts A, Cantor M, Goodstein D, Kuo A, Minovitsky S, Nikitin R, Ohm RA, Otillar R, Poliakov A, Ratnere I, Riley R, Smirnova T, Rokhsar D, Dubchak I. (2012). The Genome Portal of the Department of Energy Joint Genome Institute. Nucleic Acids Res. 26 -32 pp.
- Slides: 21