An Introduction to Ontology for Evolutionary Biology Barry

An Introduction to Ontology for Evolutionary Biology Barry Smith 1

Who am I? NCBO: National Center for Biomedical Ontology (NIH Roadmap Center) − Stanford Medical Informatics − University of San Francisco Medical Center − The Mayo Clinic − University at Buffalo (PI of Dissemination and Ontology Best Practices) 2

NCBO will offer • Technology for uploading, browsing, and using biomedical ontologies • Methods to make the online “publication” of ontologies more like that of journal articles • Tools to enable the biomedical community to put ontologies to work on a daily basis 3
http: //bioportal. bioontology. org 4

Hierarchy-to-root view 5

Who am I? Co-PI Protein Ontology Advisory Boards of Ontology for Biomedical Investigations Cleveland Clinic Semantic Database in Cardiothoracic Surgery Gene Ontology Scientific Advisory Board Advancing Clinico-Genomic Trials on Cancer (ACGT) 6

W-LOV World’s Longest Ontology Video Introduction to Biomedical Ontologies This 8 -lecture course provides a basic introduction to ontology, with special reference to applications in the field of biomedical research. It is designed to be of interest to both philosophers and those with a background in the life sciences. 1. What is an ontology and what is it useful for? 2. Basic Formal Ontology: An upper-level ontology for scientific research 3. Open Biomedical Ontologies (OBO) and the Web Ontology Language (OWL) 4. The OBO Relation Ontology 5. An ontological introduction to biomedicine: Defining organism, function and disease 6. The Gene Ontology (GO), the Foundational Model of Anatomy (FMA) and the Infectious Disease Ontology (IDO) 7. The OBO Foundry: A suite of biomedical ontologies to support reasoning and data integration 8. Further applications http: //ontology. buffalo. edu/smith/Ontology_Course. html 7

MKVSDRRKFEKANFDEFESALNNKNDLVHCPSITLFES How to do biology across the genome? IPTEVRSFYEDEKSGLIKVVKFRTGAMDRKRSFEKVVIS VMVGKNVKKFLTFVEDEPDFQGGPISKYLIPKKINLMVY TLFQVHTLKFNRKDYDTLSLFYLNRGYYNELSFRVLER CHEIASARPNDSSTMRTFTDFVSGAPIVRSLQKSTIRKY GYNLAPYMFLLLHVDELSIFSAYQASLPGEKKVDTERL KRDLCPRKPIEIKYFSQICNDMMNKKDRLGDILHIILRAC ALNFGAGPRGGAGDEEDRSITNEEPIIPSVDEHGLKVC KLRSPNTPRRLRKTLDAVKALLVSSCACTARDLDIFDD NNGVAMWKWIKILYHEVAQETTLKDSYRITLVPSSDGI SLLAFAGPQRNVYVDDTTRRIQLYTDYNKNGSSEPRLK TLDGLTSDYVFYFVTVLRQMQICALGNSYDAFNHDPW MDVVGFEDPNQVTNRDISRIVLYSYMFLNTAKGCLVEY ATFRQYMRELPKNAPQKLNFREMRQGLIALGRHCVGS RFETDLYESATSELMANHSVQTGRNIYGVDFSLTSVSG 8 TTATLLQERASERWIQWLGLESDYHCSFSSTRNAEDV
MKVSDRRKFEKANFDEFESALNNKNDLVHCPSITLFESIPTEVRSFYEDEKSGLIKVVKFRTGAMDR KRSFEKVVISVMVGKNVKKFLTFVEDEPDFQGGPIPSKYLIPKKINLMVYTLFQVHTLKFNRKDYDTL SLFYLNRGYYNELSFRVLERCHEIASARPNDSSTMRTFTDFVSGAPIVRSLQKSTIRKYGYNLAPYM FLLLHVDELSIFSAYQASLPGEKKVDTERLKRDLCPRKPIEIKYFSQICNDMMNKKDRLGDILHIILRA CALNFGAGPRGGAGDEEDRSITNEEPIIPSVDEHGLKVCKLRSPNTPRRLRKTLDAVKALLVSSCAC TARDLDIFDDNNGVAMWKWIKILYHEVAQETTLKDSYRITLVPSSDGISLLAFAGPQRNVYVDDTTR RIQLYTDYNKNGSSEPRLKTLDGLTSDYVFYFVTVLRQMQICALGNSYDAFNHDPWMDVVGFEDP NQVTNRDISRIVLYSYMFLNTAKGCLVEYATFRQYMRELPKNAPQKLNFREMRQGLIALGRHCVGS RFETDLYESATSELMANHSVQTGRNIYGVDSFSLTSVSGTTATLLQERASERWIQWLGLESDYHCS FSSTRNAEDVVAGEAASSNHHQKISRVTRKRPREPKSTNDILVAGQKLFGSSFEFRDLHQLRLCYEI YMADTPSVAVQAPPGYGKTELFHLPLIALASKGDVEYVSFLFVPYTVLLANCMIRLGRRGCLNVAPV RNFIEEGYDGVTDLYVGIYDDLASTNFTDRIAAWENIVECTFRTNNVKLGYLIVDEFHNFETEVYRQS QFGGITNLDFDAFEKAIFLSGTAPEAVADAALQRIGLTGLAKKSMDINELKRSEDLSRGLSSYPTRMF NLIKEKSEVPLGHVHKIRKKVESQPEEALKLLLALFESEPESKAIVVASTTNEVEELACSWRKYFRVV WIHGKLGAAEKVSRTKEFVTDGSMQVLIGTKLVTEGIDIKQLMMVIMLDNRLNIIELIQGVGRLRDGG LCYLLSRKNSWAARNRKGELPPKEGCITEQVREFYGLESKKGKKGQHVGCCGSRTDLSADTVELIE RMDRLAEKQATASMSIVALPSSFQESNSSDRYRKYCSSDEDSNTCIHGSANASTNAITTAST NVRTNATTNASTNASTNATTNSSTNATTTASTNVRTSATTTASINVRTSAT TTESTNSSTNATTTESTNSNTSATTTASINVRTSATTTESTNSSTSATTTASINVR TSATTTKSINSSTNATTTESTNSNTNATTTESTNSSTNATTTESTNSNTSAATTES TNSNTSATTTESTNASAKEDANKDGNAEDNRFHPVTDINKESYKRKGSQMVLLERKKLKAQFPNTS ENMNVLQFLGFRSDEIKHLFLYGIDIYFCPEGVFTQYGLCKGCQKMFELCVCWAGQKVSYRRIAWE ALAVERMLRNDEEYKEYLEDIEPYHGDPVGYLKYFSVKRREIYSQIQRNYAWYLAITRRRETISVLDS TRGKQGSQVFRMSGRQIKELYFKVWSNLRESKTEVLQYFLNWDEKKCQEEWEAKDDTVVVEALE KGGVFQRLRSMTSAGLQGPQYVKLQFSRHHRQLRSRYELSLGMHLRDQIALGVTPSKVPHWTAFL SMLIGLFYNKTFRQKLEYLLEQISEVWLLPHWLDLANVEVLAADDTRVPLYMLMVAVHKELDSDDVP DGRFDILLCRDSSREVGE 9

The GO idea: through annotation of da what cellular component? what molecular function? what biological process? 10

three types of data what cellular component? what molecular function? what biological process? 11

The GO Idea Gly. Prot Mouse. Ecotope sphingolipid transporter activity Diabet. In. Gene Glu. Chem

The GO Idea Gly. Prot Mouse. Ecotope Holliday junction helicase complex Diabet. In. Gene Glu. Chem

The GO Idea Gly. Prot Mouse. Ecotope sphingolipid transporter activity Diabet. In. Gene Glu. Chem

Benefits of GO 1. rooted in experimental biology 2. links people to data and to literature 3. links data to data (comparability) • across species (human, mouse, yeast, fly. . . ) • across granularities (molecule, cell, organ, organism, population) 4. links medicine to biological science 5. serves cumulation of scientific knowledge in algorithmically tractable form

How to extend the GO methodology to other areas of the life sciences? OBO (Open Biomedical Ontologies) created 2001 in Ashburner and Lewis a shared portal for (so far) 60 ontologies http: //obo. sourceforge. net with a common OBO flatfile format 16 16
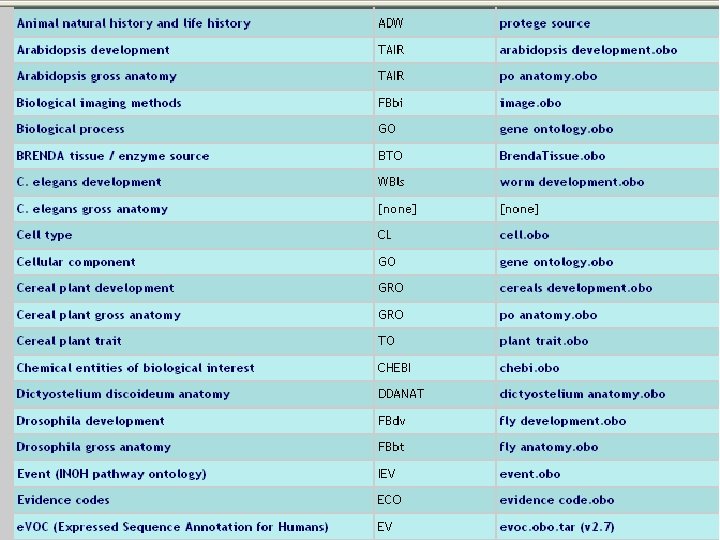
17

In 2004 reform efforts initiated linking GO to other ontologies and data sources via formal relations GO id: CL: 0000062 name: osteoblast def: "A bone-forming cell which secretes an extracellular matrix. Hydroxyapatite crystals are then deposited into the matrix to form bone. " is_a: CL: 0000055 relationship: develops_from CL: 0000008 relationship: develops_from CL: 0000375 Osteoblast differentiation: Processes whereby an osteoprogenitor cell or a cranial neural crest cell acquires the specialized features of an osteoblast, a bone -forming cell which secretes extracellular matrix. + Cell type = New Definition 18
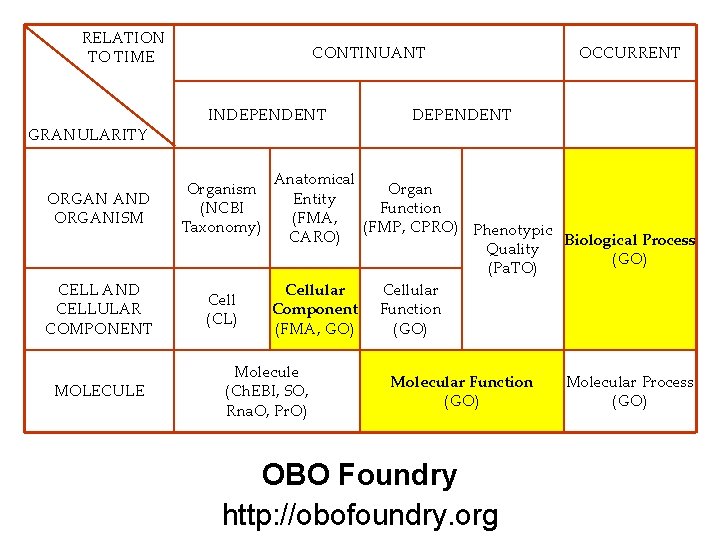
RELATION TO TIME GRANULARITY CONTINUANT INDEPENDENT ORGAN AND ORGANISM Organism (NCBI Taxonomy) CELL AND CELLULAR COMPONENT Cell (CL) MOLECULE OCCURRENT DEPENDENT Anatomical Organ Entity Function (FMA, (FMP, CPRO) Phenotypic CARO) Biological Process Quality (GO) (Pa. TO) Cellular Component Function (FMA, GO) (GO) Molecule (Ch. EBI, SO, Rna. O, Pr. O) Molecular Function (GO) OBO Foundry http: //obofoundry. org Molecular Process (GO) 19
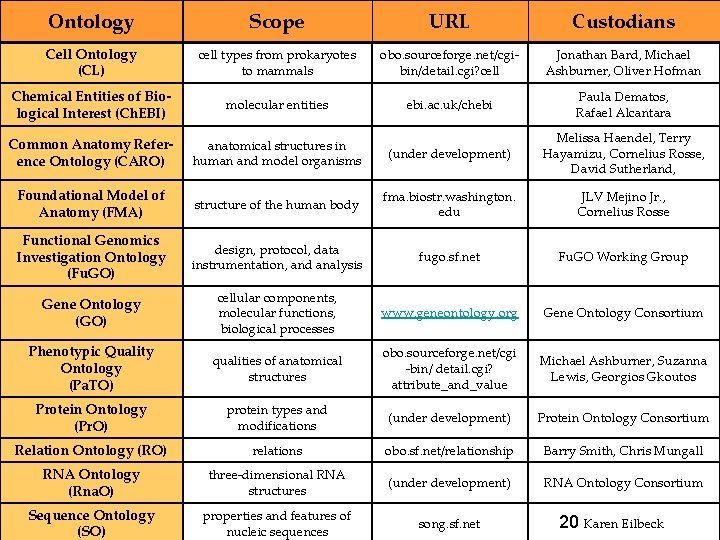
Ontology Scope URL Custodians Cell Ontology (CL) cell types from prokaryotes to mammals obo. sourceforge. net/cgibin/detail. cgi? cell Jonathan Bard, Michael Ashburner, Oliver Hofman Chemical Entities of Biological Interest (Ch. EBI) molecular entities ebi. ac. uk/chebi Paula Dematos, Rafael Alcantara Common Anatomy Reference Ontology (CARO) anatomical structures in human and model organisms (under development) Melissa Haendel, Terry Hayamizu, Cornelius Rosse, David Sutherland, Foundational Model of Anatomy (FMA) structure of the human body fma. biostr. washington. edu JLV Mejino Jr. , Cornelius Rosse Functional Genomics Investigation Ontology (Fu. GO) design, protocol, data instrumentation, and analysis fugo. sf. net Fu. GO Working Group Gene Ontology (GO) cellular components, molecular functions, biological processes www. geneontology. org Gene Ontology Consortium Phenotypic Quality Ontology (Pa. TO) qualities of anatomical structures obo. sourceforge. net/cgi -bin/ detail. cgi? attribute_and_value Michael Ashburner, Suzanna Lewis, Georgios Gkoutos Protein Ontology (Pr. O) protein types and modifications (under development) Protein Ontology Consortium Relation Ontology (RO) relations obo. sf. net/relationship Barry Smith, Chris Mungall RNA Ontology (Rna. O) three-dimensional RNA structures (under development) RNA Ontology Consortium Sequence Ontology (SO) properties and features of nucleic sequences song. sf. net 20 Karen Eilbeck

Goal: create the ontology resources for evolutionary biology 21

The ontologies in the OBO Foundry are scientific ontologies 22

Administrative/database ontologies Highly task-dependent – reusability and compatibility not (always) important Entities may be brought into existence by the ontology itself (convention. . . ) If there is no field for gender in our database, then persons do not have gender Can be secret, local, temporary Are comparable to software artifacts 23

Scientific ontologies are comparable to scientific theories must be open, based on consensus must be compatible with neighboring scientific ontologies and with results of scientifc research must be stable, evolve gracefully in tandem with the advance of knowledge must be evidence-based (testable) 24

Foundry ontologies are scientific ontologies Every representational unit in the ontology must be such that the developers believe it to refer to some entity on the basis of the best current scientific evidence Important role of instances that we can observe in the laboratory 25

Ontologies are like science texts – they are representations of what is aka universals, kinds, types, categories, general in reality species, genera, . . . aka universals, kinds, types, categories, species, genera, . . . 26

A central distinction universal vs. instance (catalog vs. inventory) (science text vs. diary) (human being vs. Arnold Schwarzenegger) 27

For scientific ontologies it is generalizations (universals) that are important For databases it is (normally) instances that are important = particulars in reality: mouse #000001 tail #00004 video image #23300014, etc. 28

Ontologies are representations of what is general in reality aka universals, kinds, types, categories, species, genera, . . . aka universals, kinds, types, categories, species, genera, . . . instances in reality are linked to universals via the instance_of relation 29

The distinction between universals and instances allows us to provide clear formal definitions of the relations which connect ontology terms A is_a B =def. A is narrower in meaning than B cancer documentation is_a cancer 30

The distinction between universals and instances allows us to provide clear logical definitions of the relations which connect ontology terms A is_a B =def. every instance of A is an instance of B 31

part_of A part_of B =def. every instance of A is an instance-level part of some instance of B Mary’s heart instance-level part of Mary cell nucleus part_of cell 32
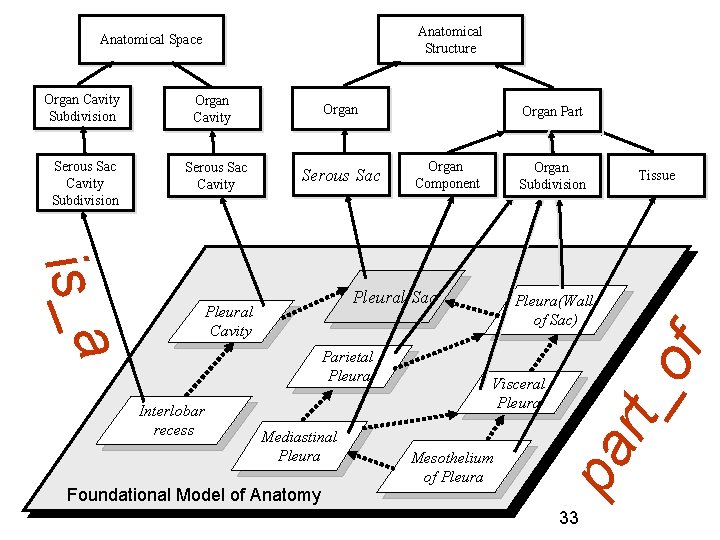
Anatomical Structure Organ Serous Sac Cavity Subdivision Serous Sac Cavity Serous Sac Organ Component Organ Subdivision is_a Pleural Sac Pleural Cavity Parietal Pleura Interlobar recess Organ Part Mediastinal Pleura Foundational Model of Anatomy FMA Tissue Pleura(Wall of Sac) of Organ Cavity Visceral Pleura Mesothelium of Pleura rt_ Organ Cavity Subdivision pa Anatomical Space 33

Kinds of relations <universal, universal>: is_a, part_of, . . . <instance, universal>: this cell instance_of the universal cell <instance, instance>: Mary’s heart part_of Mary 34

Foundry principle for definitions Definitions should be of the following form an A =def. a B which Cs where B is the is_a parent of A and C is some differentia Definitions are rooted in the is_a hierarchy 35

OBO Relation Ontology 1. 0 Foundational is_a part_of Spatial located_in contained_in adjacent_to Temporal transformation_of derives_from preceded_by Participation has_participant has_agent “Relations in Biomedical Ontologies”, Genome Biology, April 2005 36
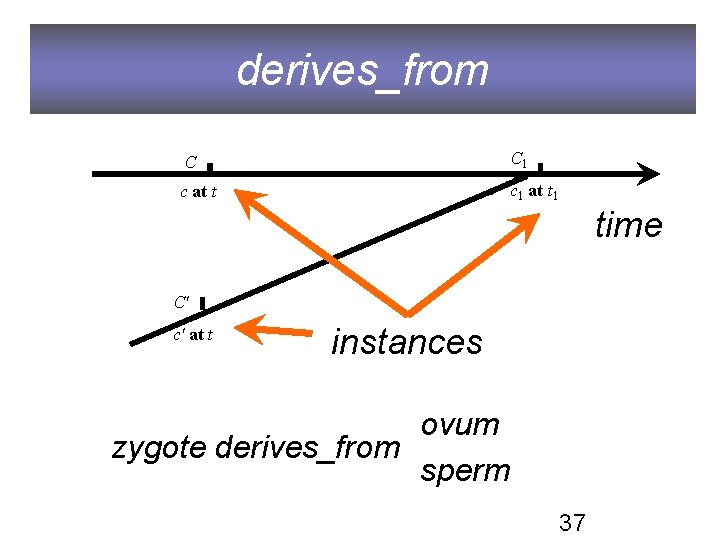
derives_from C C 1 c at t c 1 at t 1 time C' c' at t instances ovum zygote derives_from sperm 37
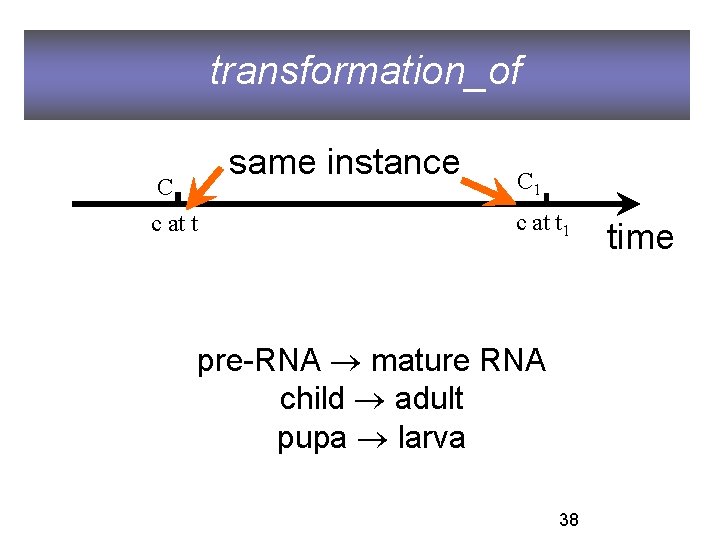
transformation_of same instance C c at t C 1 c at t 1 pre-RNA mature RNA child adult pupa larva 38 time

transformation_of C 2 transformation_of C 1 =def. any instance of C 2 was at some earlier time an instance of C 1 fetus transformation_of embryo larva transformation_of pupa adult transformation_of child 39
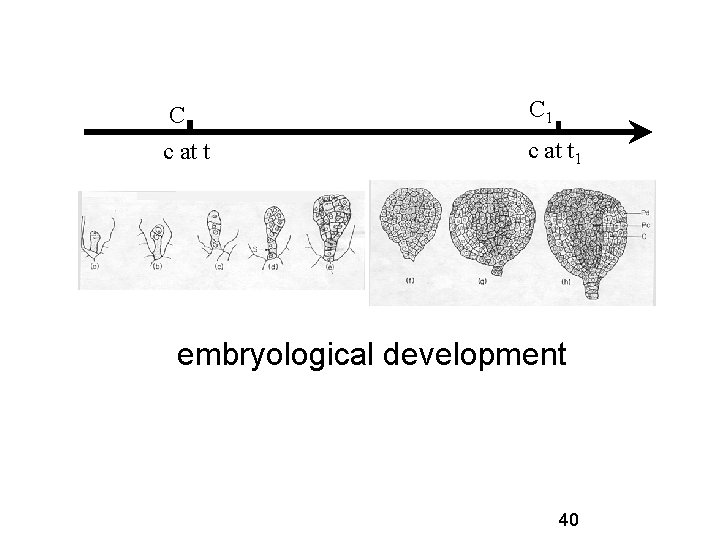
C C 1 c at t 1 embryological development 40
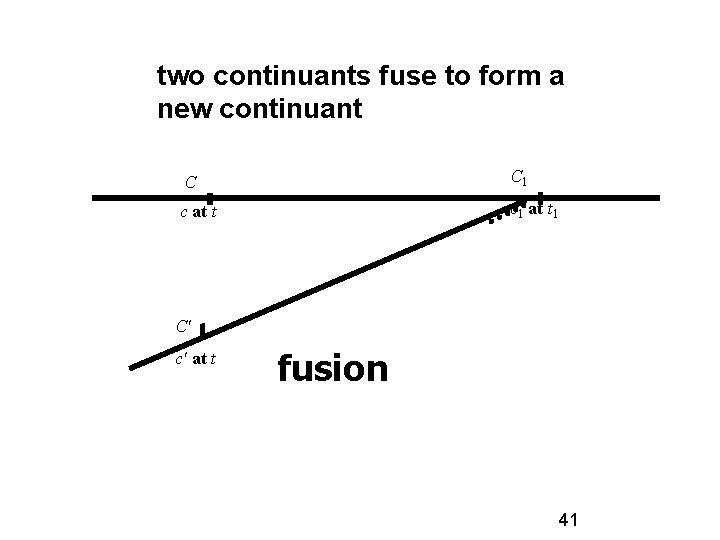
two continuants fuse to form a new continuant C C 1 c at t c 1 at t 1 C' c' at t fusion 41
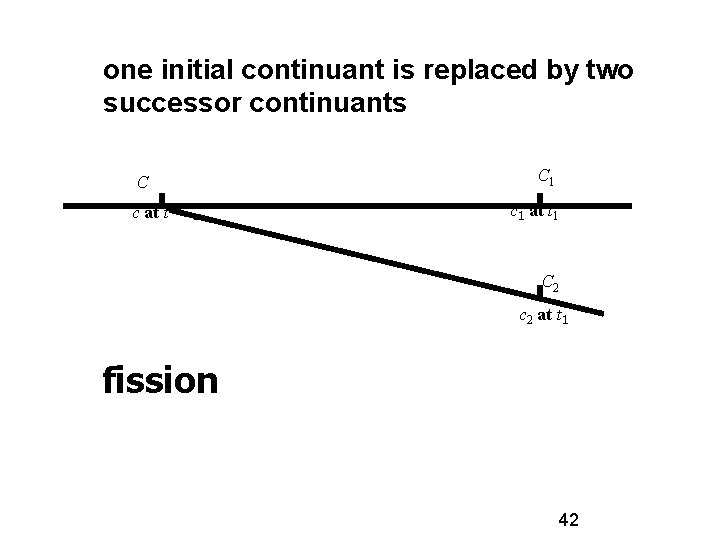
one initial continuant is replaced by two successor continuants C c at t C 1 c 1 at t 1 C 2 c 2 at t 1 fission 42
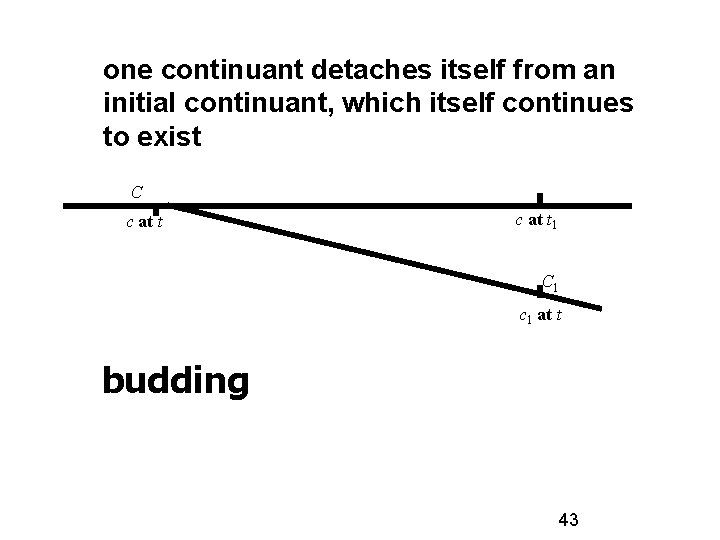
one continuant detaches itself from an initial continuant, which itself continues to exist C c at t 1 C 1 c 1 at t budding 43
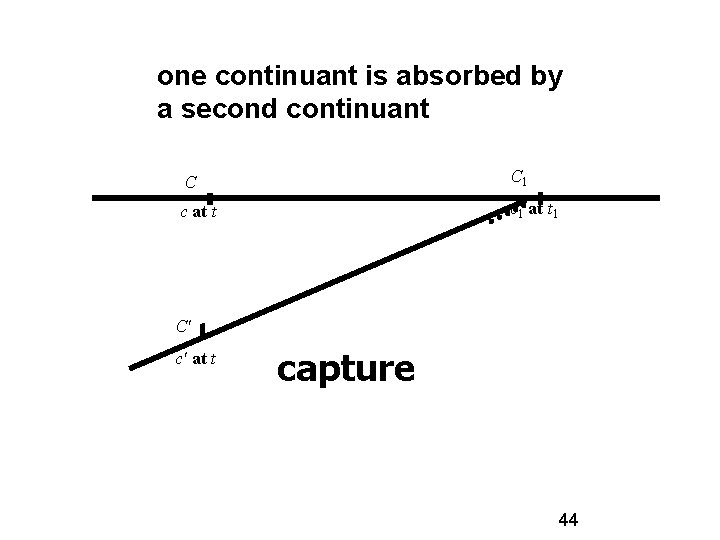
one continuant is absorbed by a second continuant C C 1 c at t c 1 at t 1 C' c' at t capture 44

New ‘regulates' relations in GO def: "A relation between a process and a process. A regulates B if the unfolding of A affects the frequency, rate or extent of B. A is called the regulating process, B the regulated process“ A regulates B =def. A is a process type and B is a process type and every instance of A is such that its unfolding affects the frequency, rate or extent of some instance of B. 45

Relations proposed for RO 2. 0 inheres_in has_input has_function has_quality realization_of directly_descends_from (CARO) homologous_to (CARO) 46

An ontology is a representation of universals We learn about universals in reality from looking at the results of scientific experiments as expressed in the form of scientific theories – which describe, not what is particular in reality, but what is general 47
A photographic image is a representation of an instance 48

A photographic image is a representation of an instance We learn about instances in reality by performing scientific experiments on the basis of scientific hypotheses and describing the results in general terms provided (ideally) by ontologies 49

Mature OBO Foundry ontologies Cell Ontology (CL) Foundational Model of Anatomy (FMA) Gene Ontology (GO) Phenotypic Quality Ontology (PATO) Relation Ontology (RO) Sequence Ontology (SO) 50

Foundry ontologies being built ab initio Common Anatomy Reference Ontology (CARO) – and various organism specific anatomy ontologies Ontology for Biomedical Investigations (OBI) Protein Ontology (PRO) RNA Ontology (Rna. O) Subcellular Anatomy Ontology (SAO) 51

Ontologies in planning phase Environment Ontology (Env. O) Infectious Disease Ontology (IDO) Biobank/Biorepository Ontology Food Ontology Allergy Ontology Vaccine Ontology Still needed: Organism Taxonomy 52

RELATION TO TIME GRANULARITY CONTINUANT INDEPENDENT ORGAN AND ORGANISM Organism (NCBI Taxonomy) CELL AND CELLULAR COMPONENT Cell (CL) MOLECULE OCCURRENT DEPENDENT Anatomical Organ Entity Function (FMA, (FMP, CPRO) Phenotypic CARO) Biological Process Quality (GO) (Pa. TO) Cellular Component Function (FMA, GO) (GO) Molecule (Ch. EBI, SO, Rna. O, Pr. O) Molecular Function (GO) Molecular Process (GO) continuants vs. occurrents independent vs. dependent entities 53
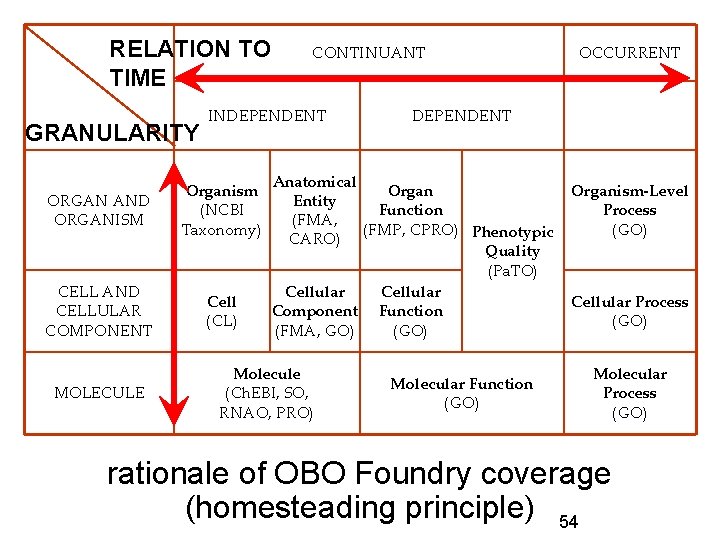
RELATION TO TIME GRANULARITY INDEPENDENT ORGAN AND ORGANISM Organism (NCBI Taxonomy) CELL AND CELLULAR COMPONENT Cell (CL) MOLECULE CONTINUANT DEPENDENT Anatomical Organ Entity Function (FMA, (FMP, CPRO) Phenotypic CARO) Quality (Pa. TO) Cellular Component Function (FMA, GO) (GO) Molecule (Ch. EBI, SO, RNAO, PRO) OCCURRENT Molecular Function (GO) Organism-Level Process (GO) Cellular Process (GO) Molecular Process (GO) rationale of OBO Foundry coverage (homesteading principle) 54
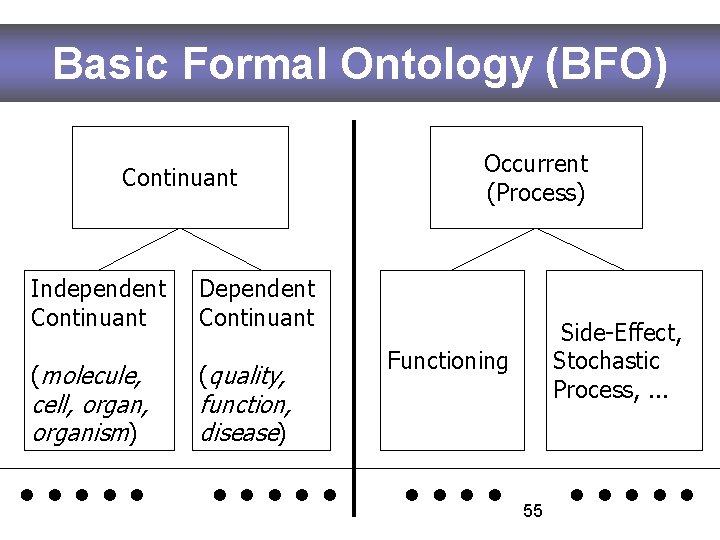
Basic Formal Ontology (BFO) Continuant Independent Continuant Dependent Continuant (molecule, (quality, cell, organism) Occurrent (Process) Side-Effect, Stochastic Process, . . . Functioning function, disease) . . 55
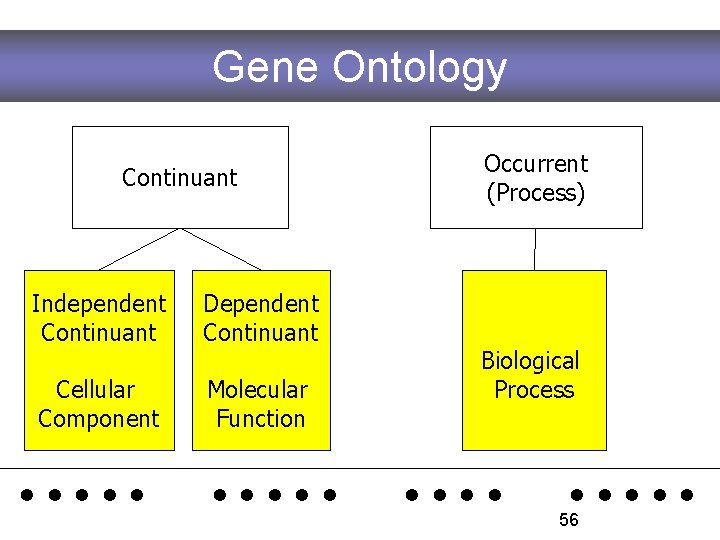
Gene Ontology Continuant Independent Continuant Dependent Continuant Cellular Component Molecular Function Occurrent (Process) Biological Process . . 56
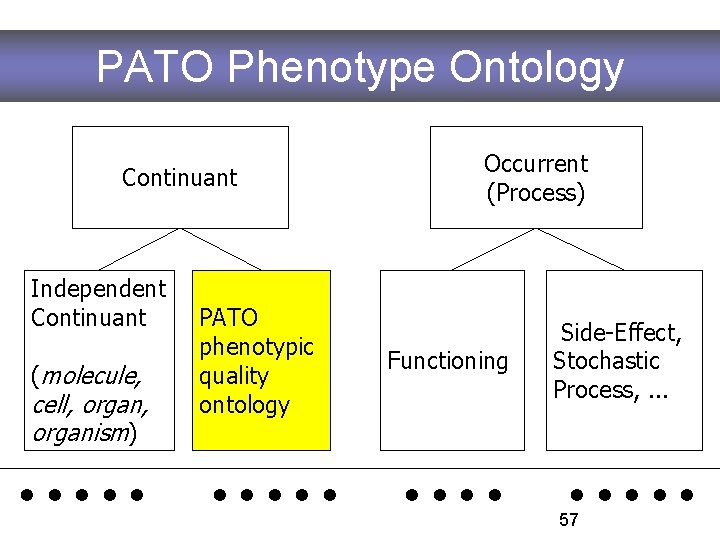
PATO Phenotype Ontology Continuant Independent Continuant (molecule, cell, organism) PATO phenotypic quality ontology Occurrent (Process) Functioning Side-Effect, Stochastic Process, . . 57

An example of a PATO quality The particular redness of the left eye of a single individual fly – An instance of a quality universal The color ‘red’ – A quality universal Note: the eye does not instantiate ‘red’ PATO represents quality universals: color, temperature, texture, shape … 58

Qualities are dependent entities Qualities require (depend on) bearers, which are independent continuants Example: – A shape requires a physical object as its bearer – If the physical object ceases to exist (e. g. it decomposes), then the shape ceases to exist 59
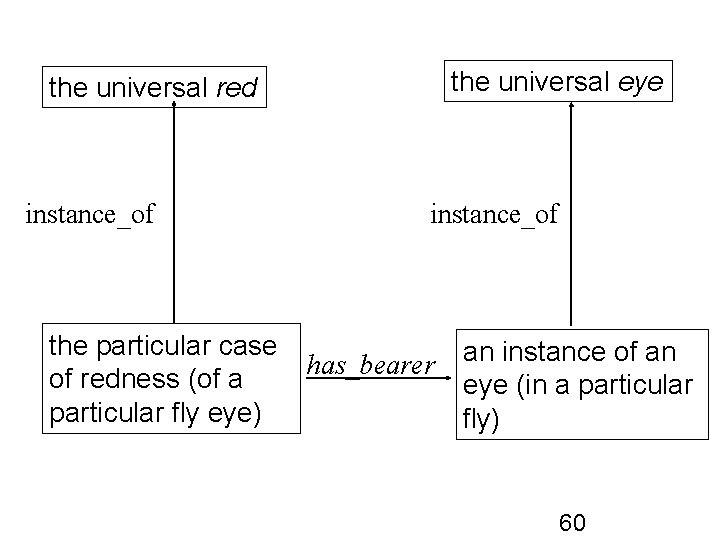
the universal eye the universal red instance_of the particular case has_bearer of redness (of a particular fly eye) an instance of an eye (in a particular fly) 60

What a quality is NOT Qualities are not measurements – Instances of qualities exist independently of their measurements – Qualities can have zero or more measurements These are not the names of qualities: – – percentage process abnormal high Open problem: how relate qualities such as length to measurement values? 61

How to do anatomy ontology Functional: cardiovascular system, nervous system Spatial: head, trunk, limb Developmental: endoderm, germ ring, lens placode Structural: tissue, organ, cell Stage: developmental staging series 62

CARO – Common Anatomy Reference Ontology for the first time provides guidelines for model organism researchers who wish to achieve comparability of annotations based on anatomy and development 63

64
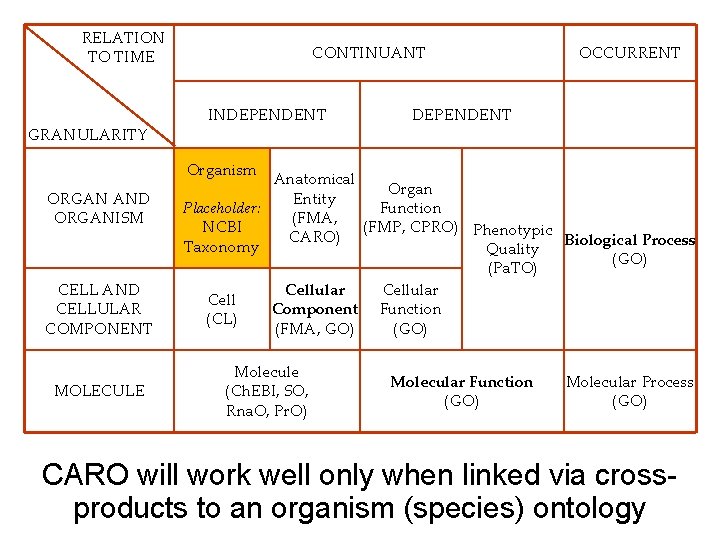
RELATION TO TIME GRANULARITY CONTINUANT INDEPENDENT OCCURRENT DEPENDENT Organism ORGAN AND ORGANISM CELL AND CELLULAR COMPONENT MOLECULE Anatomical Organ Entity Placeholder: Function (FMA, NCBI (FMP, CPRO) Phenotypic CARO) Biological Process Taxonomy Quality (GO) (Pa. TO) Cellular Cell Component Function (CL) (FMA, GO) (GO) Molecule (Ch. EBI, SO, Rna. O, Pr. O) Molecular Function (GO) Molecular Process (GO) CARO will work well only when linked via crossproducts to an organism (species) ontology 65

All OBO Foundry ontologies work in the same way – we have data (biosample, haplotype, clinical data, survey data, . . . ) – we need to make this data available for semantic search and algorithmic processing – we create a consensus-based ontology for annotating the data – we use cross-products to compose more complex terms and assertions 69
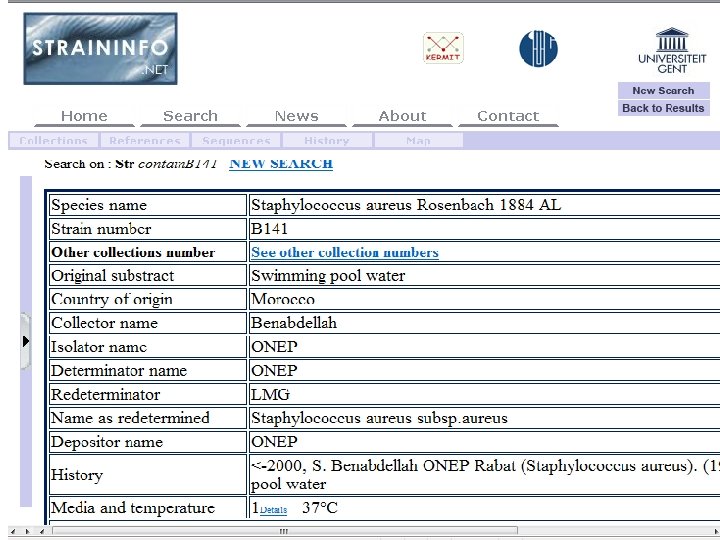
70
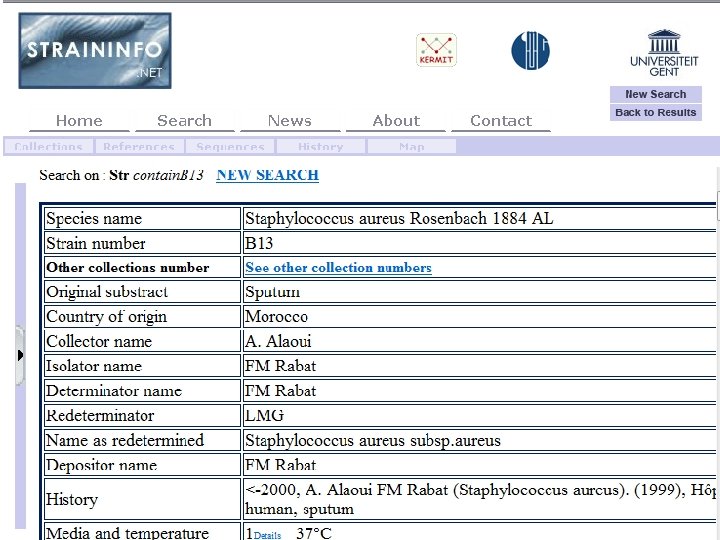
71
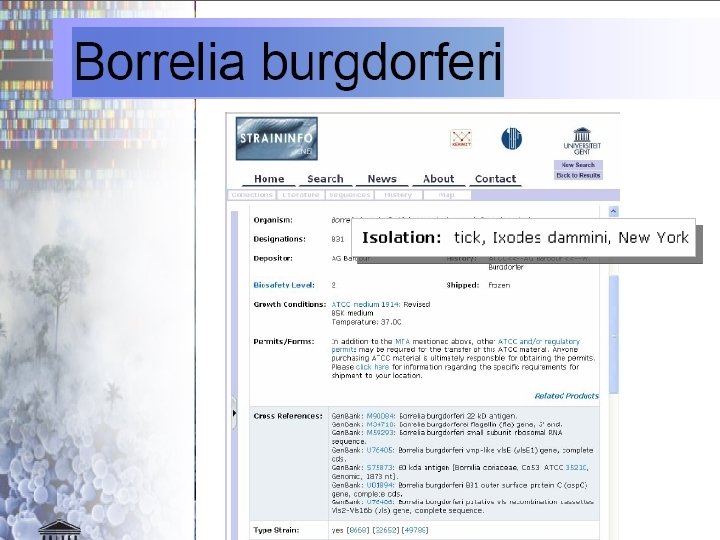
72

The Environment Ontology 73

The Hole Story


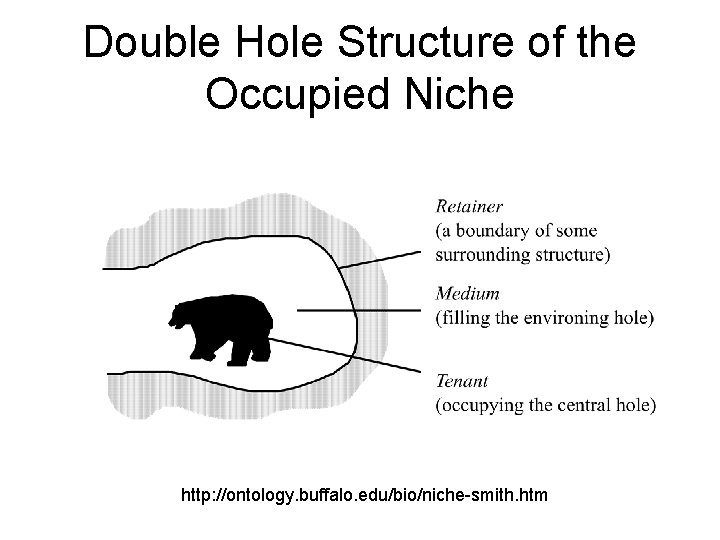
Double Hole Structure of the Occupied Niche http: //ontology. buffalo. edu/bio/niche-smith. htm

Tenant, medium and retainer the medium of the bear’s niche is a circumscribed body of air medium might be body of water, cytosol, nasal mucosa, epithelium, endocardium, synovial tissue. . .
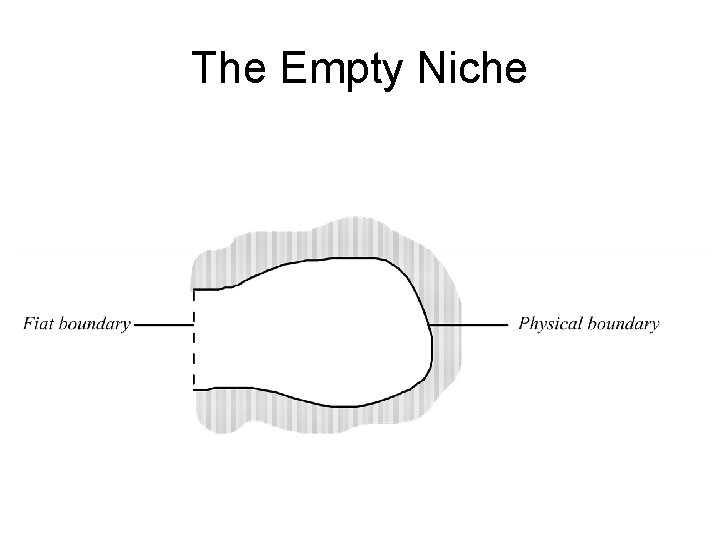
The Empty Niche
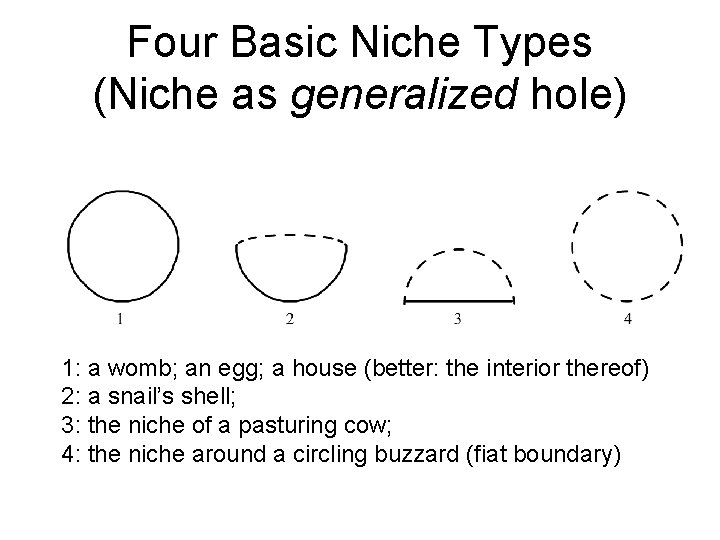
Four Basic Niche Types (Niche as generalized hole) 1: a womb; an egg; a house (better: the interior thereof) 2: a snail’s shell; 3: the niche of a pasturing cow; 4: the niche around a circling buzzard (fiat boundary)

Elton – niche as role the ‘niche’ of an animal means its place in the biotic environment, its relations to food and enemies. [. . . ] When an ecologist says ‘there goes a badger’ he should include in his thoughts some definite idea of the animal’s place in the community to which it belongs, just as if he had said ‘there goes the vicar’ (Elton 1927, pp. 63 f. )

G. E. Hutchinson: niche as volume in a functionally defined space the niche = an n-dimensional hypervolume whose dimensions correspond to resource gradients over which species are distributed
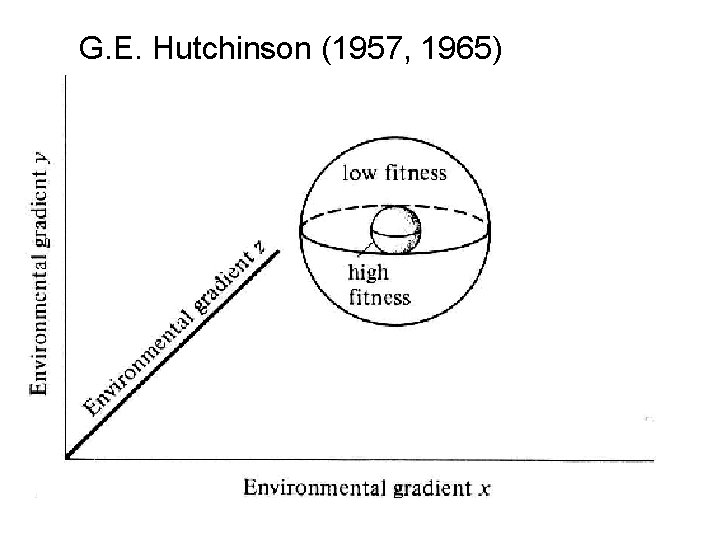
G. E. Hutchinson (1957, 1965)

Hypervolume niche = a location in an attribute space defined by a specific constellation of environmental variables such as degree of slope, exposure to sunlight, soil fertility, foliage density, salinity. . .

The Environment Ontology Genomic Standards Consortium National Environment Research Council (UK) Barcode of Life Project Encyclopedia of Life Project 86

Env. O combines the spatial and Hutchinsonian perspectives to create a consensus controlled vocabulary for representing macroscopic (geographical) mesoscopic (behavioral) microscopic (cellular, molecular …) environments 87

Applications of Env. O in biology 88
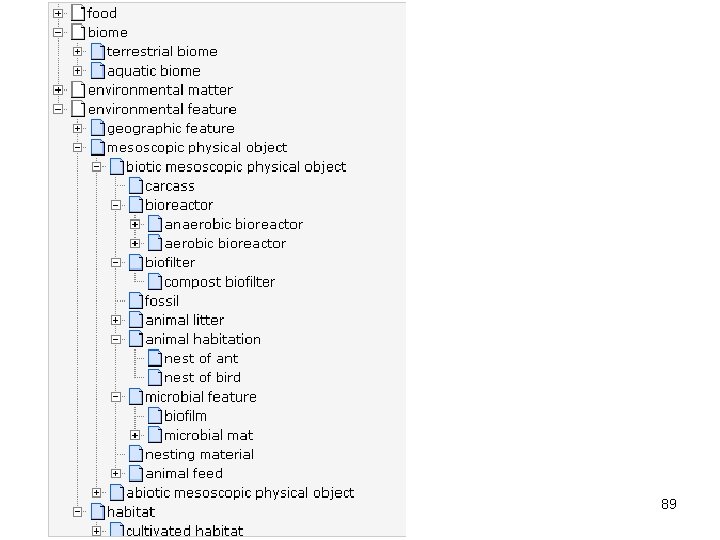
89
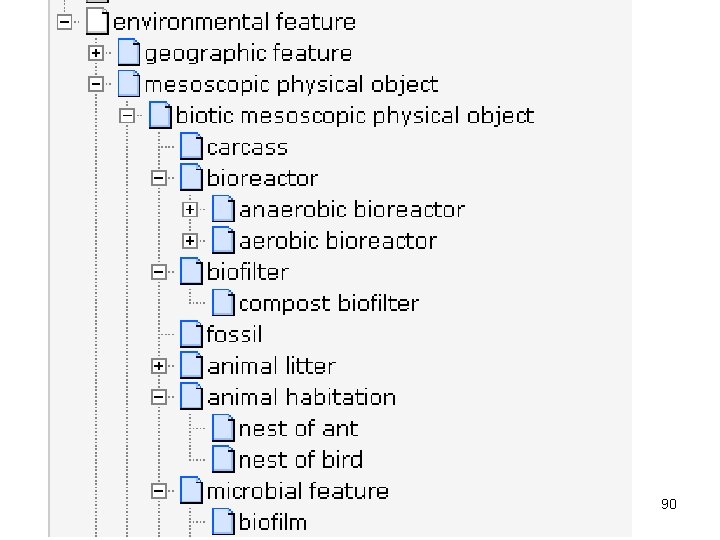
90

Environment = totality of circumstances external to a living organism or group of organisms – p. H – evapotranspiration – turbidity – available light – predominant vegetation – predatory pressure – nutrient limitation –… 91

How Env. O currently works for information retrieval Retrieve all experiments on organisms obtained from: – deep-sea thermal vents – arctic ice cores – rainforest canopy – alpine melt zone Retrieve all data on organisms sampled from: – hot and dry environments – cold and wet environments – a height above 5, 000 meters Retrieve all the omic data from soil organisms subject to: – moderate heavy metal contamination 92

Scale: From microbiological to geographic Data on locations of organisms/samples, sources of museum artifacts. . . Environments have spatial locations Data on organism interactions, e. g. on bacterial infection – how the interior of one organism or organism part serves as environment for another organism

The Environment Ontology OBO Foundry Genomic Standards Consortium National Environment Research Council (UK) Barcode of Life Project Encyclopedia of Life Project 94
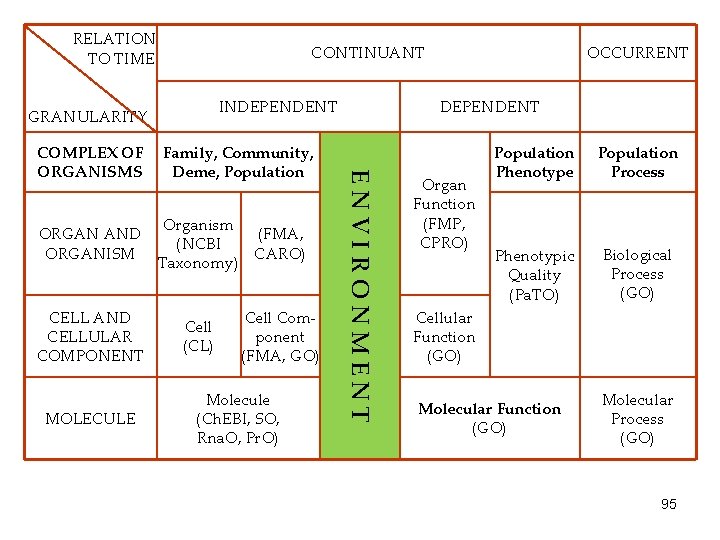
RELATION TO TIME CONTINUANT INDEPENDENT GRANULARITY ORGAN AND ORGANISM CELL AND CELLULAR COMPONENT MOLECULE Family, Community, Deme, Population Organism (FMA, (NCBI CARO) Taxonomy) Cell (CL) Cell Component (FMA, GO) Molecule (Ch. EBI, SO, Rna. O, Pr. O) DEPENDENT ENVIRONMENT COMPLEX OF ORGANISMS OCCURRENT Organ Function (FMP, CPRO) Population Phenotype Population Process Phenotypic Quality (Pa. TO) Biological Process (GO) Cellular Function (GO) Molecular Process (GO) 95
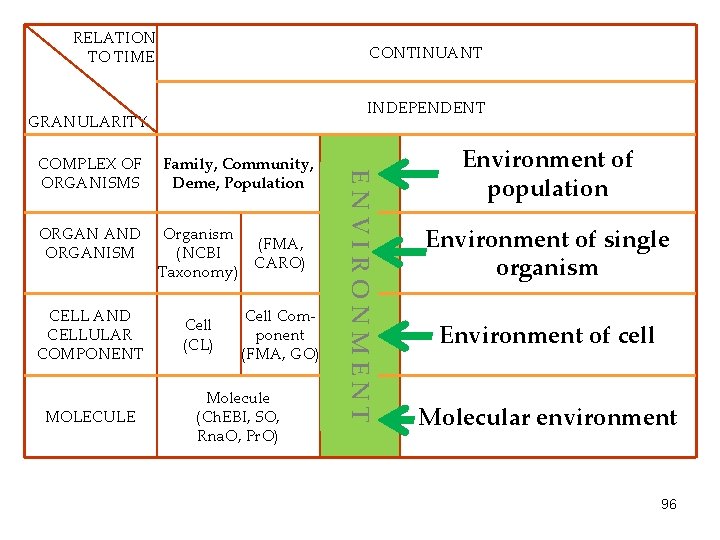
RELATION TO TIME CONTINUANT INDEPENDENT GRANULARITY ORGAN AND ORGANISM CELL AND CELLULAR COMPONENT MOLECULE Family, Community, Deme, Population Organism (FMA, (NCBI CARO) Taxonomy) Cell (CL) Cell Component (FMA, GO) Molecule (Ch. EBI, SO, Rna. O, Pr. O) ENVIRONMENT COMPLEX OF ORGANISMS Environment of population Environment of single organism Environment of cell Molecular environment 96
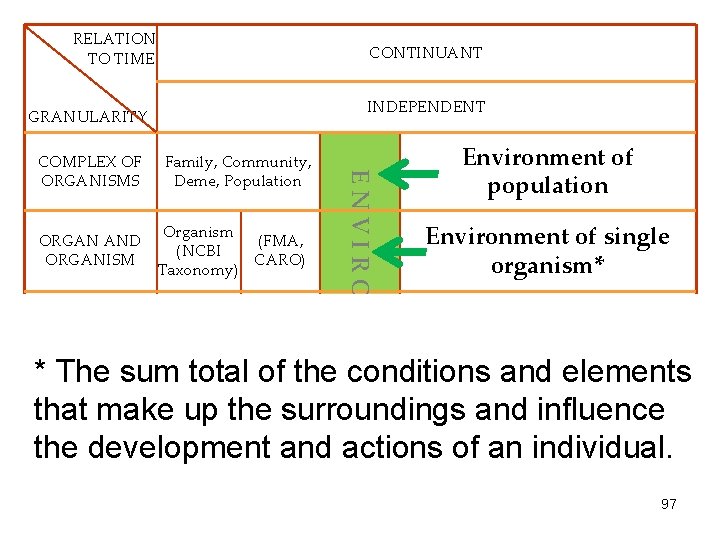
RELATION TO TIME CONTINUANT INDEPENDENT GRANULARITY ORGAN AND ORGANISM CELL AND CELLULAR COMPONENT Family, Community, Deme, Population Organism (FMA, (NCBI CARO) Taxonomy) Cell (CL) Cell Component (FMA, GO) ENVIRONMENT COMPLEX OF ORGANISMS Environment of population Environment of single organism* Environment of cell * The sum total of the conditions and elements Molecule (Ch. EBI, SO, MOLECULE that make up the surroundings and influence Molecular environment Rna. O, Pr. O) the development and actions of an individual. 97
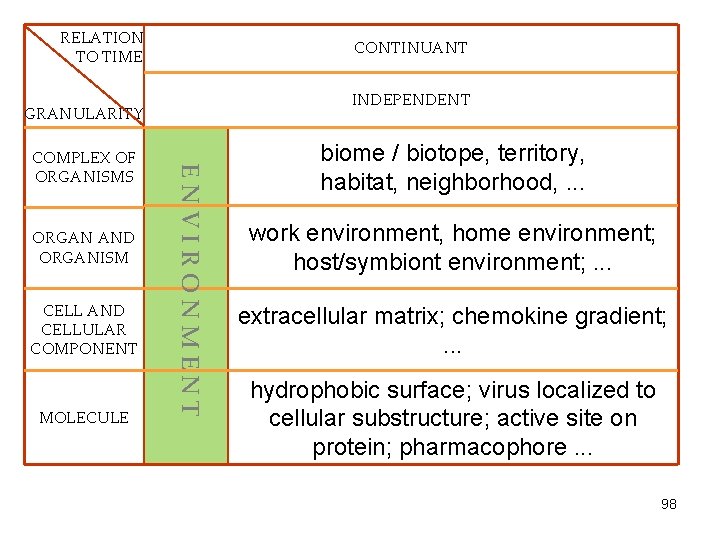
RELATION TO TIME CONTINUANT INDEPENDENT GRANULARITY CELL AND CELLULAR COMPONENT MOLECULE ENVIRONMENT ORGAN AND ORGANISM COMPLEX OF ORGANISMS biome / biotope, territory, habitat, neighborhood, . . . work environment, home environment; host/symbiont environment; . . . extracellular matrix; chemokine gradient; . . . hydrophobic surface; virus localized to cellular substructure; active site on protein; pharmacophore. . . 98

OBO Foundry principle of orthogonality designed to foster division of labor methodology for coordination designed to support cross-linkage between orthogonal ontologies

The methodology of cross-products compound terms in ontologies to be defined as cross-products of simpler terms: elevated blood glucose cross-product of PATO: increased concentration with FMA: blood and Che. BI: glucose. factoring out of ontologies into disciplinespecific modules (orthogonality) 100

The methodology of cross-products enforcing use of OBO’s Relation Ontology in linking terms drawn from Foundry ontologies creates a systematic approach to termformation makes the results algorithmically processible in virtue of the logical definitions provided by the RO ensures that the ontologies in the Foundry are networked together 101

Questions for an Evolution Ontology Granularity: evolution of proteins, RNA, of organisms … (PRO-Evo, RNAO, …) Organism / niche (ENVO) Derivation / homology Evolution and development (CARO / RO) Co-evolution (single organism vs. multiple organism) (GO, IDO)

How to build an ontology work with scientists with data annotation needs to create an initial top-level classification find ~50 most commonly used terms corresponding to universals in reality; address links to other ontologies arrange these terms into an informal is_a hierarchy according to the universality principle A is_a B every instance of A is an instance of B draw on the main BFO divisions and relations from RO, filling in missing terms needed to complete the hierarchy recruit domain scientists with data annotation needs to help populate the lower levels of the hierarchy and foster data integration 103

Principle of Low Hanging Fruit Include even absolutely trivial assertions (assertions you know to be universally true) cellular development process is_a cellular process cell death is_a death pneumococcal bacterium is_a bacterium Computers need to be led by the hand 104

which of these terms already exist in OBO Foundry ontologies? gene allelic variation gene pool genotype phenotype population trait speciation homology mutation inheritance organism extinction
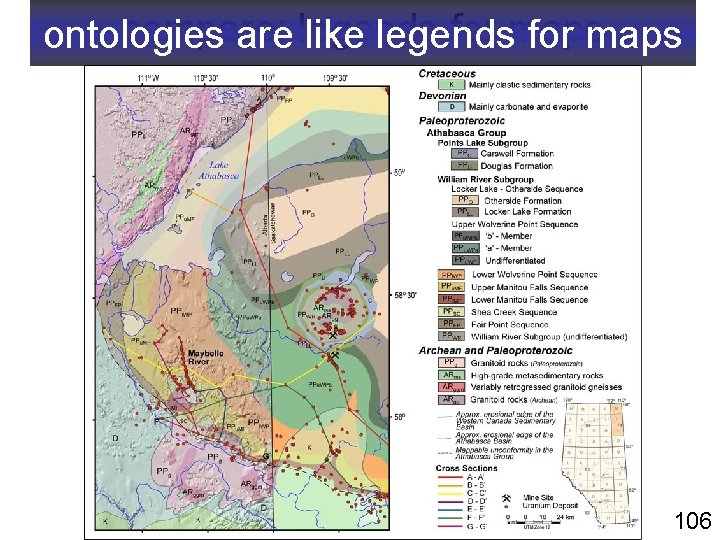
compare: legends for maps ontologies are like legends for maps 106
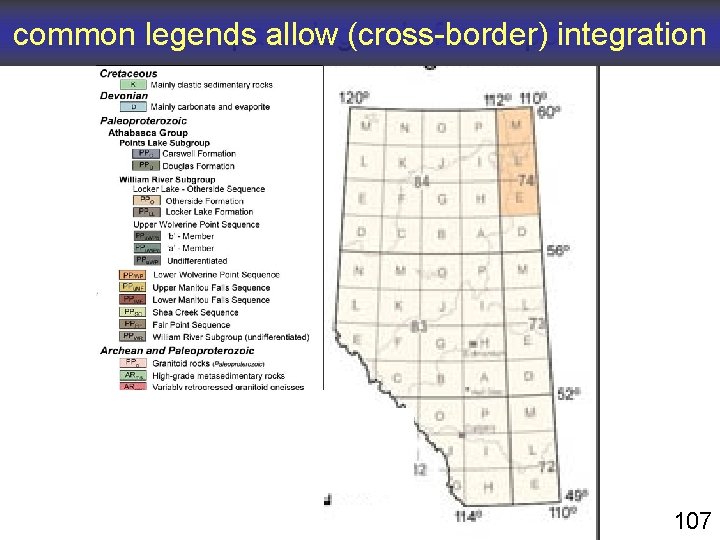
common legends allow (cross-border) integration compare: legends for maps 107

common legends help human beings use and understand complex representations of reality help human beings create useful complex representations of reality help computers process complex representations of reality help glue data together 108

Why do we need rules/standards for good ontology? Ontologies must be intelligible both to humans (for annotation and curation) and to machines (for reasoning and error-checking): the lack of rules for classification leads to human error and blocks automatic reasoning and error-checking Intuitive rules facilitate training of curators and annotators Common rules allow alignment with other ontologies 109
- Slides: 106