An Introduction to Modeling Biochemical Signal Transduction Jim

An Introduction to Modeling Biochemical Signal Transduction Jim Faeder Department of Computational and Systems Biology University of Pittsburgh School of Medicine 2014 CMACS Winter Workshop Lehman College
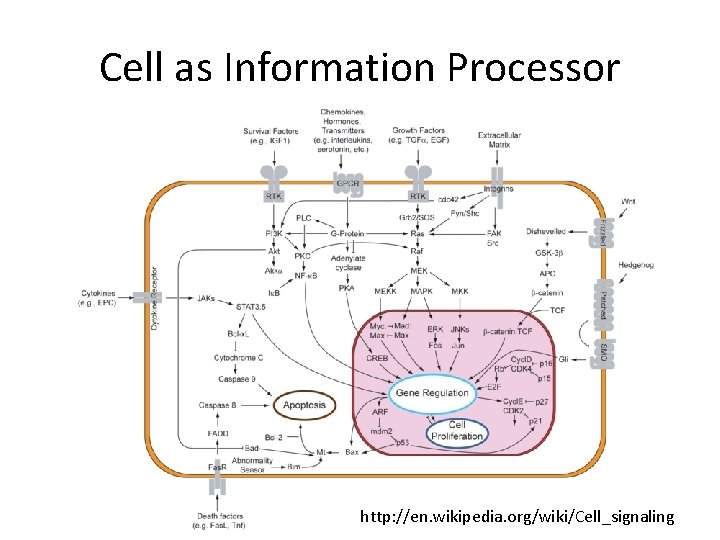
Cell as Information Processor http: //en. wikipedia. org/wiki/Cell_signaling
The cellular brain Original film from David Rogers (Vanderbuilt University) http: //www. biochemweb. org/fenteany/research/cell_migration/neutrophil. html
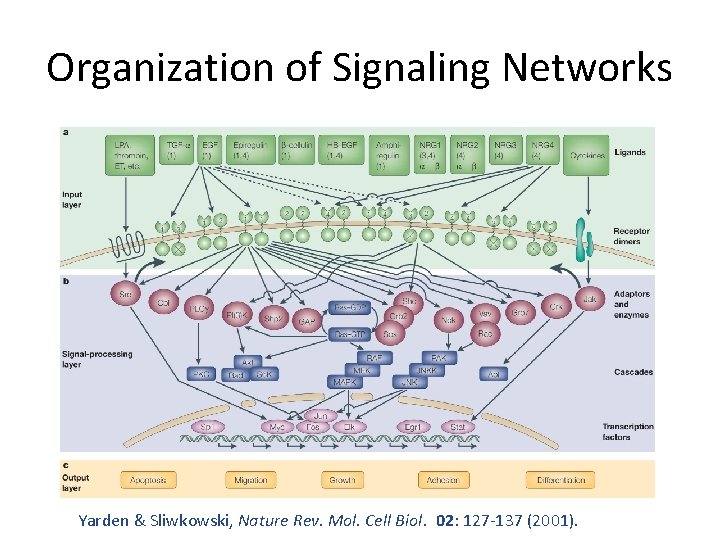
Organization of Signaling Networks Yarden & Sliwkowski, Nature Rev. Mol. Cell Biol. 02: 127 -137 (2001).
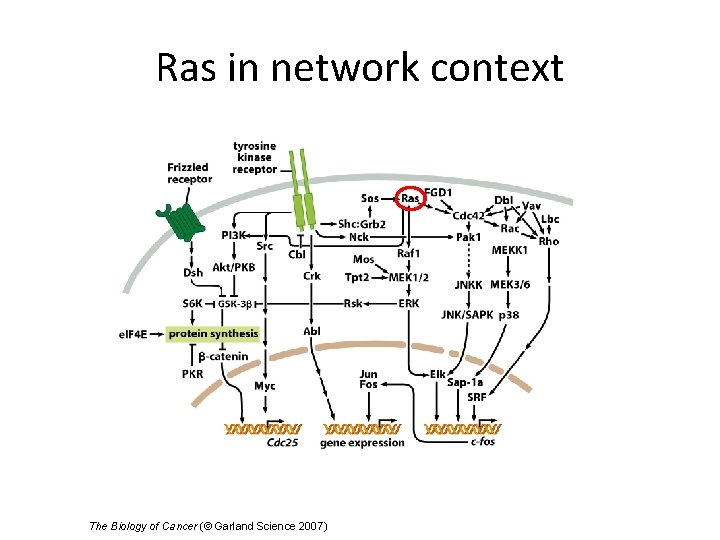
Ras in network context The Biology of Cancer (© Garland Science 2007)
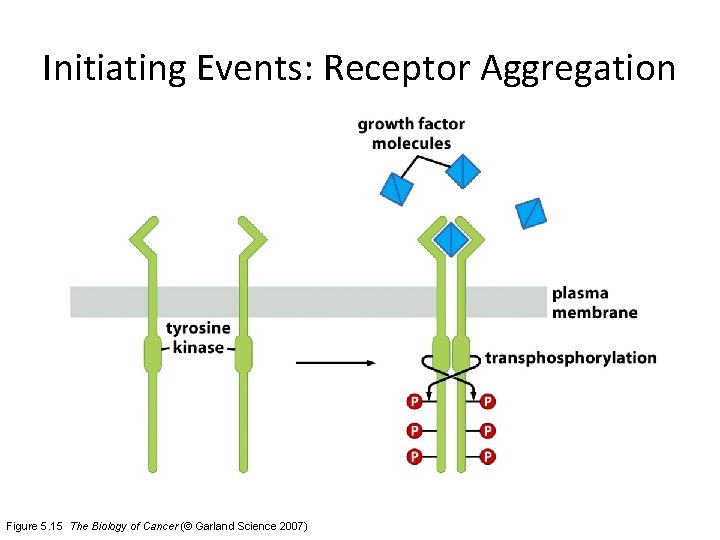
Initiating Events: Receptor Aggregation Figure 5. 15 The Biology of Cancer (© Garland Science 2007)
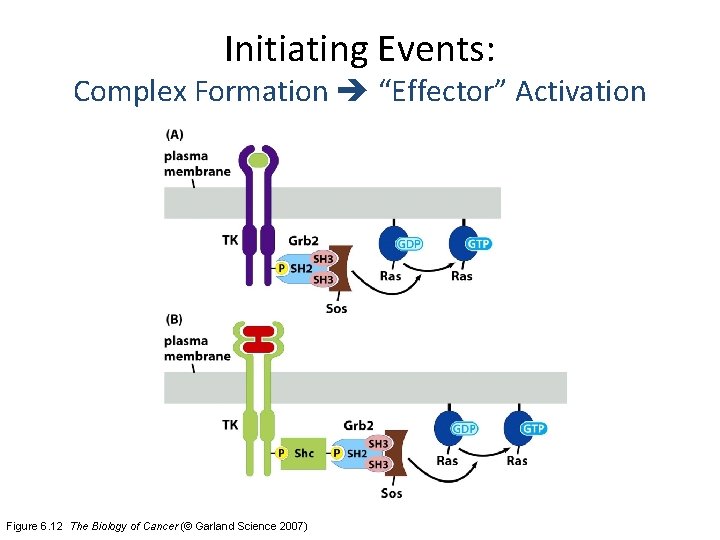
Initiating Events: Complex Formation “Effector” Activation Figure 6. 12 The Biology of Cancer (© Garland Science 2007)
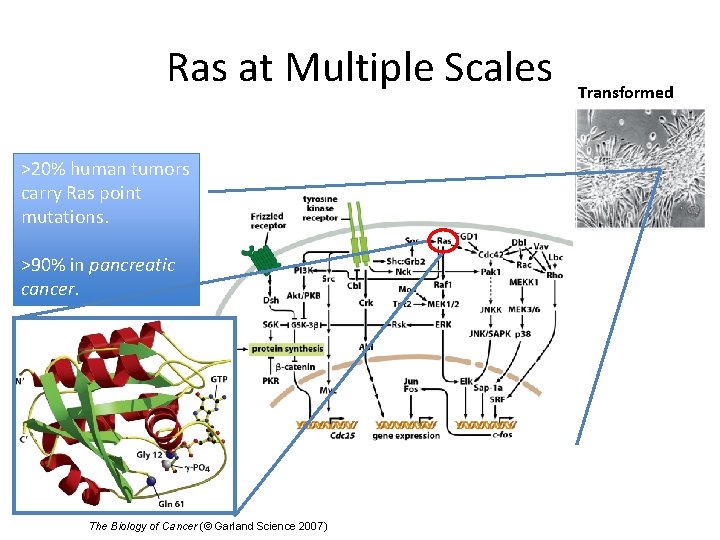
Ras at Multiple Scales >20% human tumors carry Ras point mutations. >90% in pancreatic cancer. The Biology of Cancer (© Garland Science 2007) Transformed

Video of Ras Activation Ras structure and function
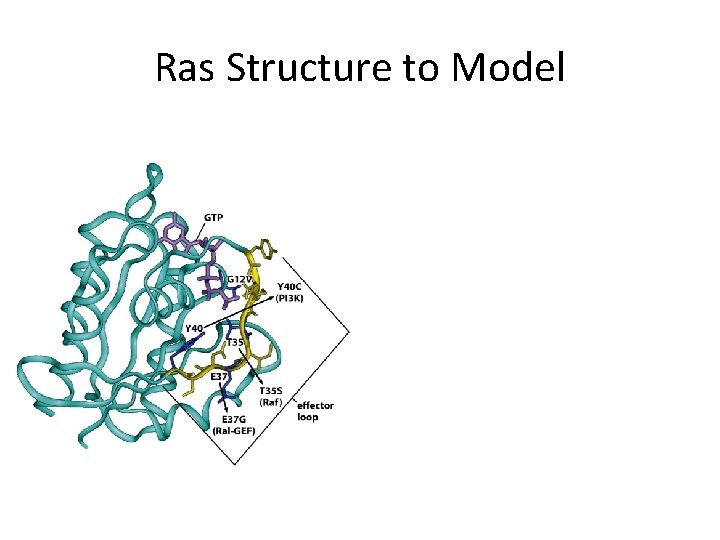
Ras Structure to Model
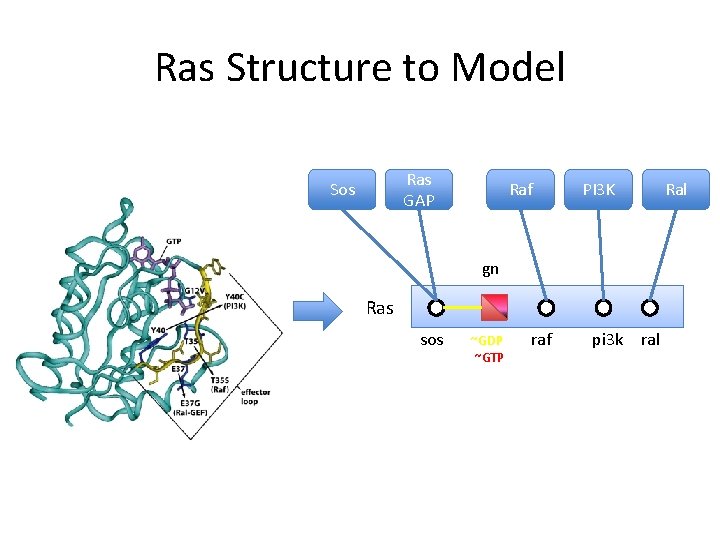
Ras Structure to Model Ras GAP Sos Raf PI 3 K Ral gn Ras sos ~GDP ~GTP raf pi 3 k ral
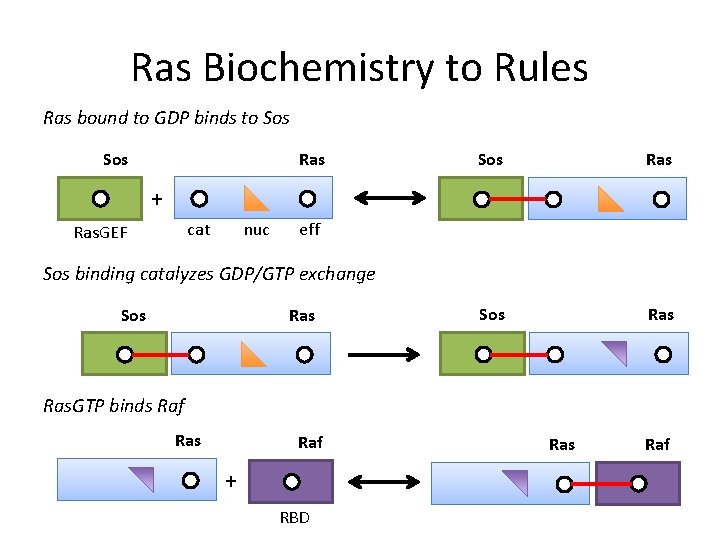
Ras Biochemistry to Rules Ras bound to GDP binds to Sos Ras + cat Ras. GEF nuc eff Sos binding catalyzes GDP/GTP exchange Sos Ras. GTP binds Raf Ras Raf + RBD Ras Raf
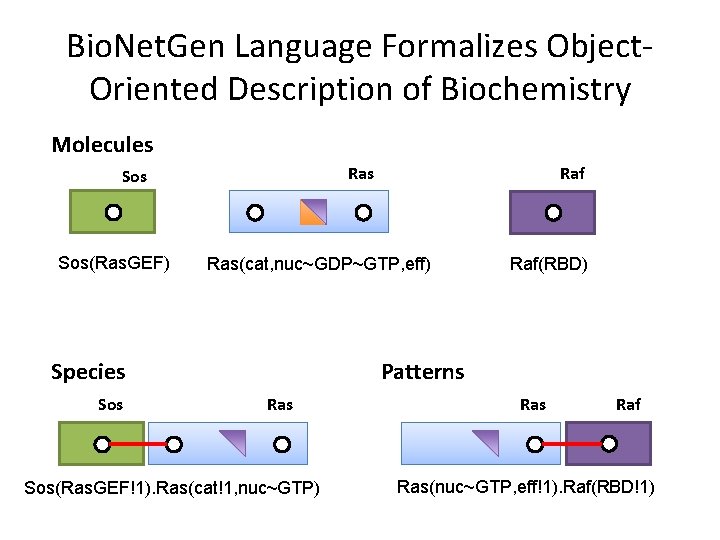
Bio. Net. Gen Language Formalizes Object. Oriented Description of Biochemistry Molecules Ras Sos(Ras. GEF) Ras(cat, nuc~GDP~GTP, eff) Species Sos Raf(RBD) Patterns Ras Sos(Ras. GEF!1). Ras(cat!1, nuc~GTP) Ras Raf Ras(nuc~GTP, eff!1). Raf(RBD!1)
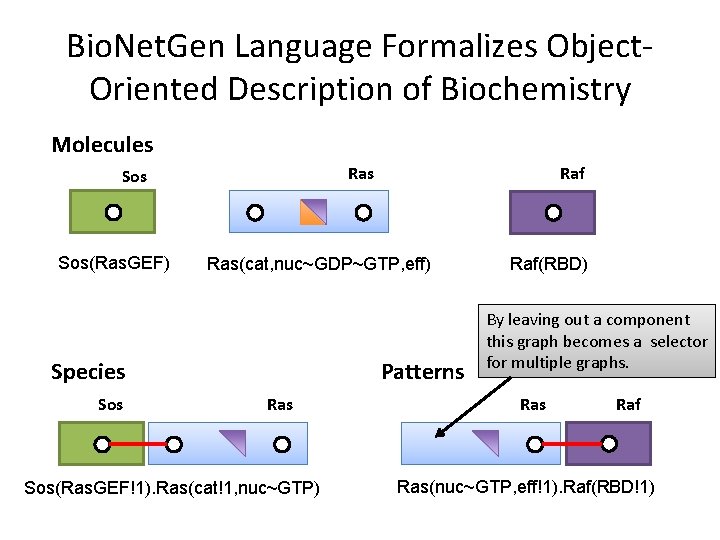
Bio. Net. Gen Language Formalizes Object. Oriented Description of Biochemistry Molecules Ras Sos(Ras. GEF) Ras(cat, nuc~GDP~GTP, eff) Species Sos Raf Patterns Ras Sos(Ras. GEF!1). Ras(cat!1, nuc~GTP) Raf(RBD) By leaving out a component this graph becomes a selector for multiple graphs. Ras Raf Ras(nuc~GTP, eff!1). Raf(RBD!1)
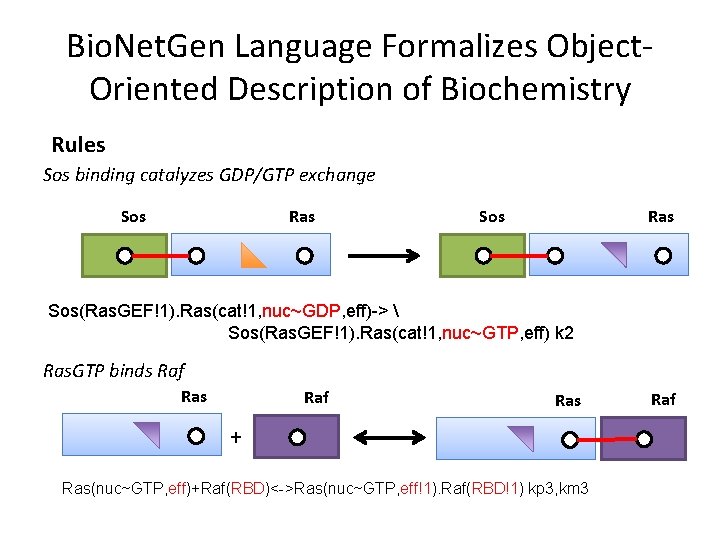
Bio. Net. Gen Language Formalizes Object. Oriented Description of Biochemistry Rules Sos binding catalyzes GDP/GTP exchange Sos Ras Sos(Ras. GEF!1). Ras(cat!1, nuc~GDP, eff)-> Sos(Ras. GEF!1). Ras(cat!1, nuc~GTP, eff) k 2 Ras. GTP binds Raf Ras + Ras(nuc~GTP, eff)+Raf(RBD)<->Ras(nuc~GTP, eff!1). Raf(RBD!1) kp 3, km 3 Raf
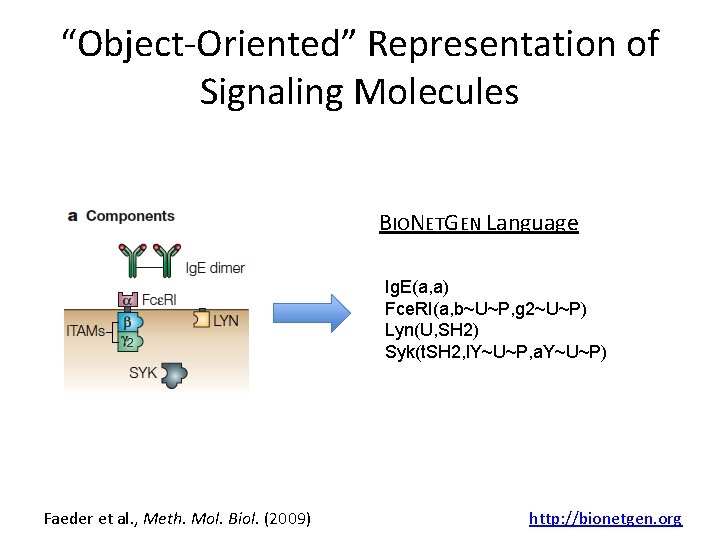
“Object-Oriented” Representation of Signaling Molecules BIONETGEN Language Ig. E(a, a) Fce. RI(a, b~U~P, g 2~U~P) Lyn(U, SH 2) Syk(t. SH 2, l. Y~U~P, a. Y~U~P) Faeder et al. , Meth. Mol. Biol. (2009) http: //bionetgen. org
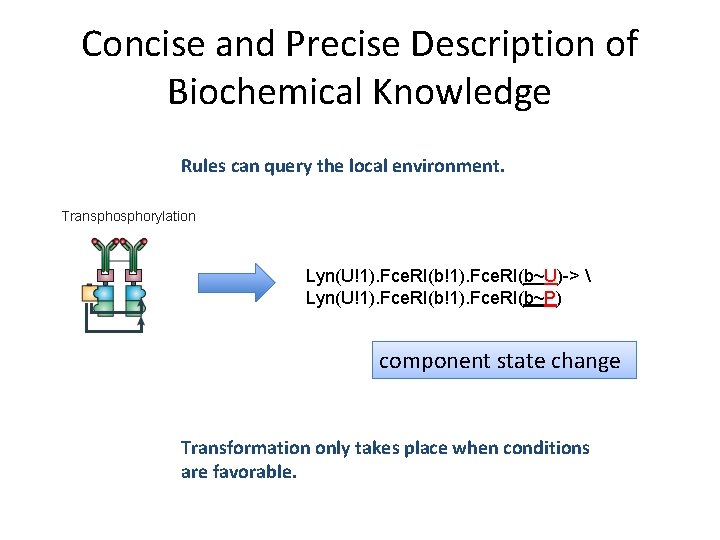
Concise and Precise Description of Biochemical Knowledge Rules can query the local environment. Transphorylation Lyn(U!1). Fce. RI(b~U)-> Lyn(U!1). Fce. RI(b~P) component state change Transformation only takes place when conditions are favorable.
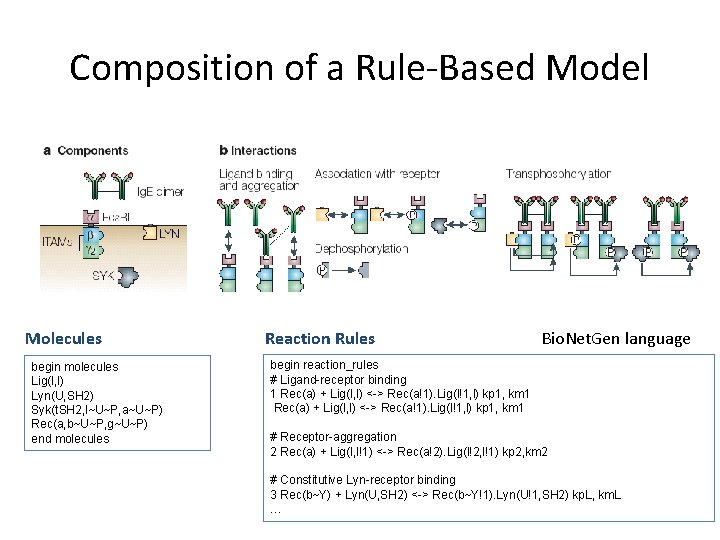
Composition of a Rule-Based Model Molecules begin molecules Lig(l, l) Lyn(U, SH 2) Syk(t. SH 2, l~U~P, a~U~P) Rec(a, b~U~P, g~U~P) end molecules Reaction Rules Bio. Net. Gen language begin reaction_rules # Ligand-receptor binding 1 Rec(a) + Lig(l, l) <-> Rec(a!1). Lig(l!1, l) kp 1, km 1 # Receptor-aggregation 2 Rec(a) + Lig(l, l!1) <-> Rec(a!2). Lig(l!2, l!1) kp 2, km 2 # Constitutive Lyn-receptor binding 3 Rec(b~Y) + Lyn(U, SH 2) <-> Rec(b~Y!1). Lyn(U!1, SH 2) kp. L, km. L …
Modeling cell signaling AIM: Model the biochemical machinery by which cells process information (and respond to it). How do we simulate dynamics of signaling networks? Representation Simulation
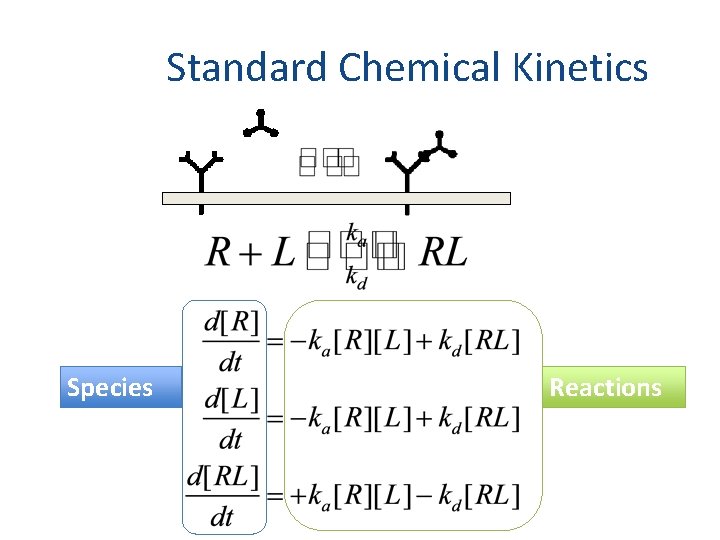
Standard Chemical Kinetics Species Reactions

Reaction Network Model of Signaling EGF EGFR SHC GRB 2 SOS Kholodenko et al. , J. Biol. Chem. 274, 30169 (1999)

Reaction Network Model of Signaling 22 species 25 reactions Kholodenko et al. , J. Biol. Chem. 274, 30169 (1999)

General formulation of chemical kinetics (continuum limit) x is vector of species concentrations S is the “stoichiometry matrix”, Sij= number of molecules of species i consumed by reaction j. v is the “reaction flux vector”, vj is the rate of reaction j. For an elementary reaction,
Modeling cell signaling How does set of Molecules and Rules get transformed into a Reaction Network of Species and Reactions? Reaction Network Representation Simulation
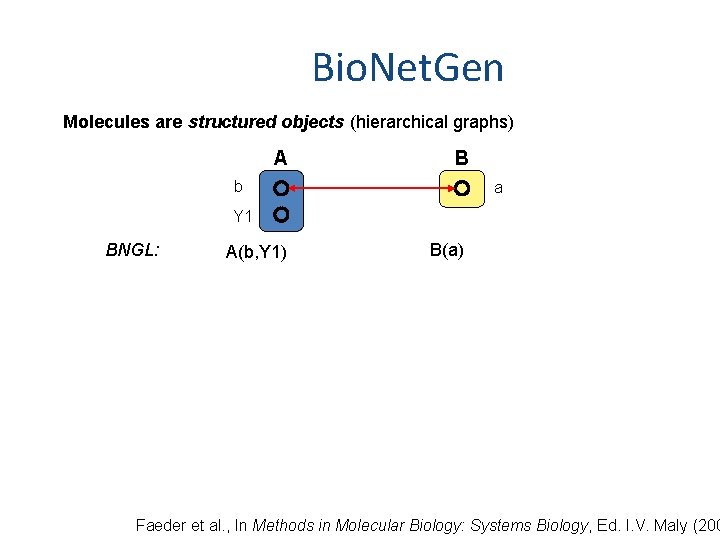
Bio. Net. Gen Molecules are structured objects (hierarchical graphs) A B b a Y 1 BNGL: A(b, Y 1) B(a) Faeder et al. , In Methods in Molecular Biology: Systems Biology, Ed. I. V. Maly (200
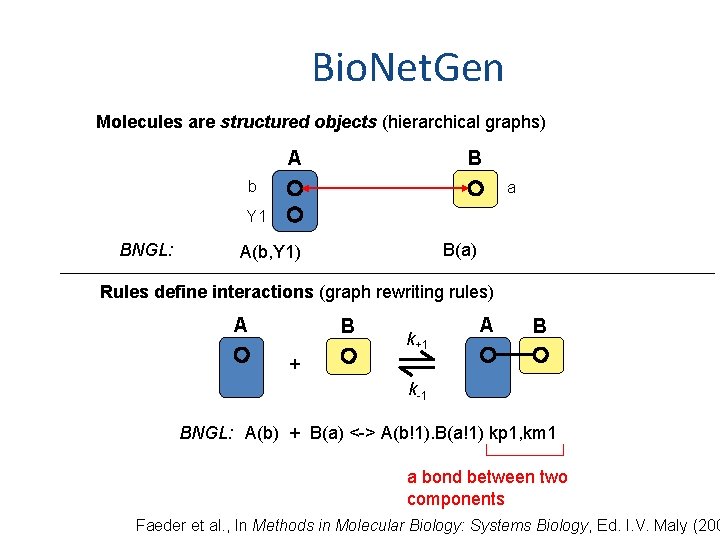
Bio. Net. Gen Molecules are structured objects (hierarchical graphs) A B b a Y 1 BNGL: B(a) A(b, Y 1) Rules define interactions (graph rewriting rules) A B k+1 A B + k-1 BNGL: A(b) + B(a) <-> A(b!1). B(a!1) kp 1, km 1 a bond between two components Faeder et al. , In Methods in Molecular Biology: Systems Biology, Ed. I. V. Maly (200
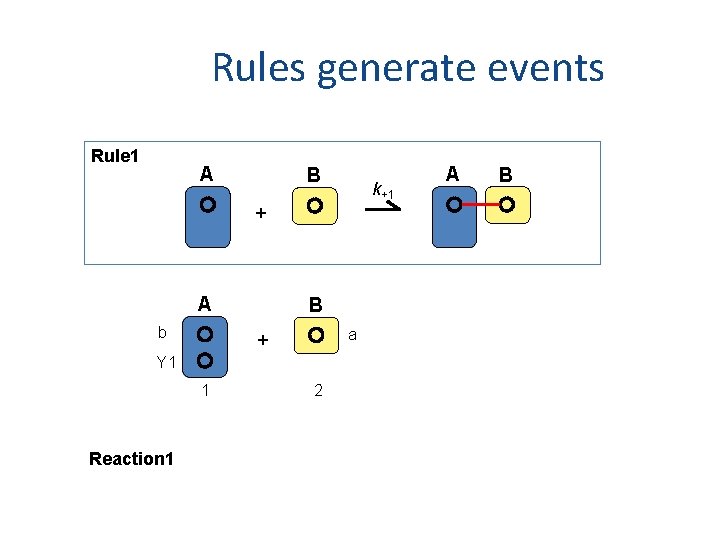
Rules generate events Rule 1 A B k+1 + A b B a + Y 1 1 Reaction 1 2 A B
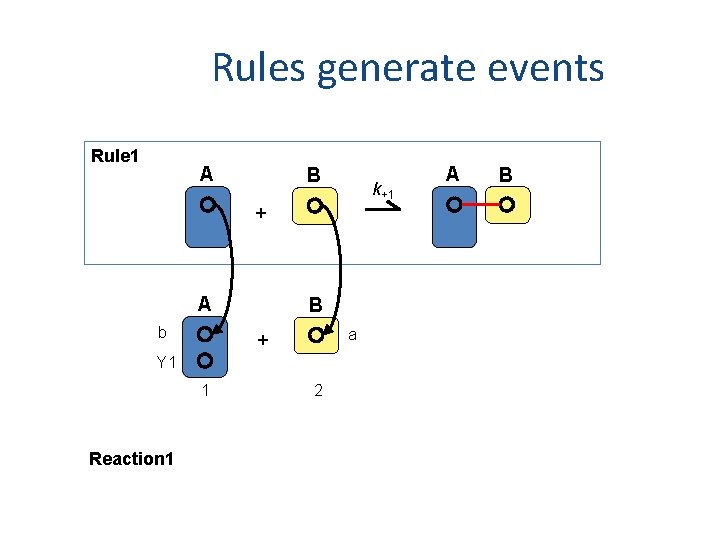
Rules generate events Rule 1 A B k+1 + A b B a + Y 1 1 Reaction 1 2 A B
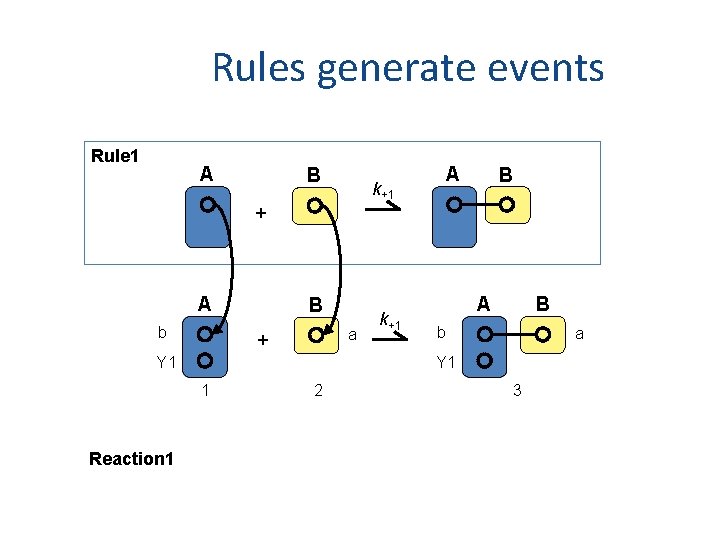
Rules generate events Rule 1 A B A k+1 B + A b B a + Y 1 B b a Y 1 1 Reaction 1 k+1 A 2 3

Rules may specify contextual requirements Rule 2 must be bound context A A p 1 b b Y 1 BNGL: Y 1 P A(b!+, Y 1~U) -> A(b!+, Y 1~P) p 1 A Reaction 2 context not changed by rule B b a Y 1 3

Rules may specify contextual requirements Rule 2 must be bound context A A p 1 b b Y 1 BNGL: Y 1 P A(b!+, Y 1~U) -> A(b!+, Y 1~P) p 1 A Reaction 2 context not changed by rule B b a Y 1 3

Rules may specify contextual requirements Rule 2 must be bound context A A p 1 b b Y 1 BNGL: Y 1 P A(b!+, Y 1~U) -> A(b!+, Y 1~P) p 1 A Reaction 2 context not changed by rule A B b a Y 1 p 1 b Y 1 3 B a P 4
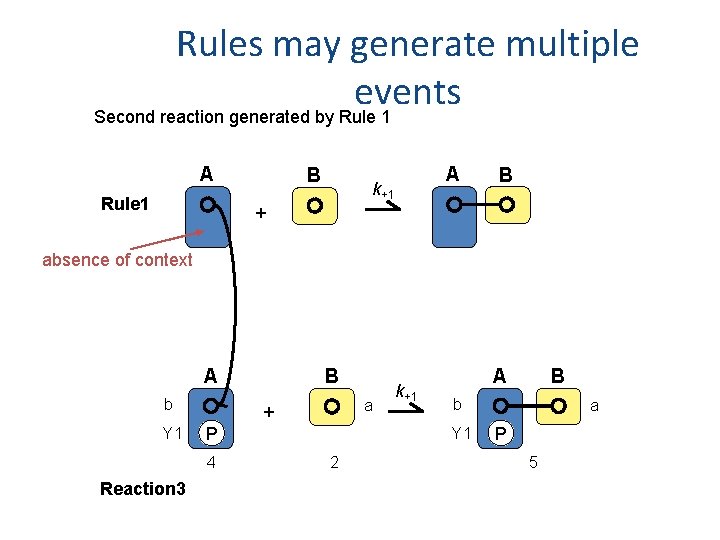
Rules may generate multiple events Second reaction generated by Rule 1 A Rule 1 B A k+1 B + absence of context A b Y 1 a + P 4 Reaction 3 B k+1 A b Y 1 2 B a P 5
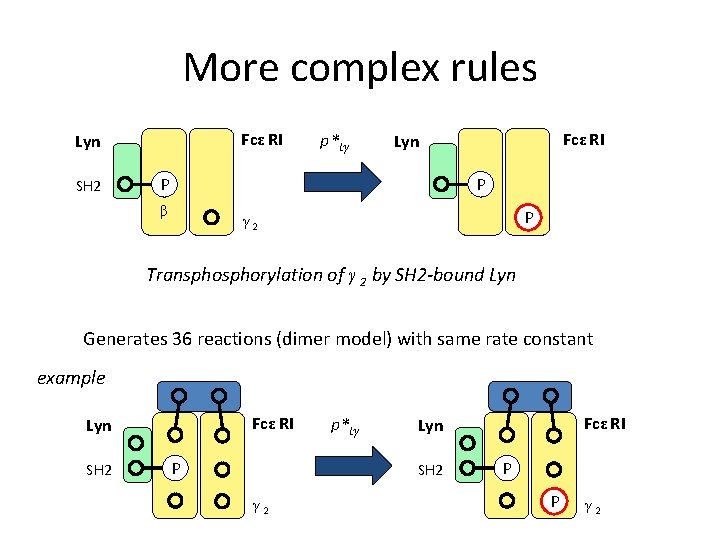
More complex rules Fcε RI Lyn SH 2 p*Lγ Fcε RI Lyn P P β P γ 2 Transphorylation of γ 2 by SH 2 -bound Lyn Generates 36 reactions (dimer model) with same rate constant example Fcε RI Lyn SH 2 P p*Lγ SH 2 γ 2 Fcε RI Lyn P P γ 2
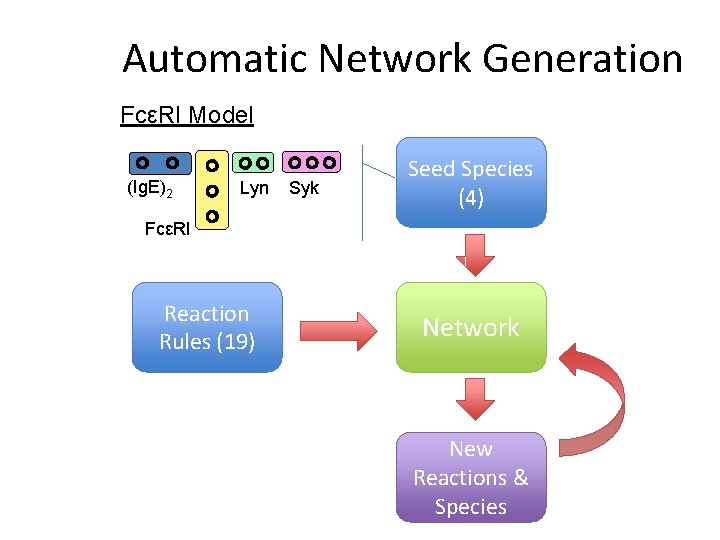
Automatic Network Generation FcεRI Model (Ig. E)2 Lyn Syk Seed Species (4) FcεRI Reaction Rules (19) Network New Reactions & Species
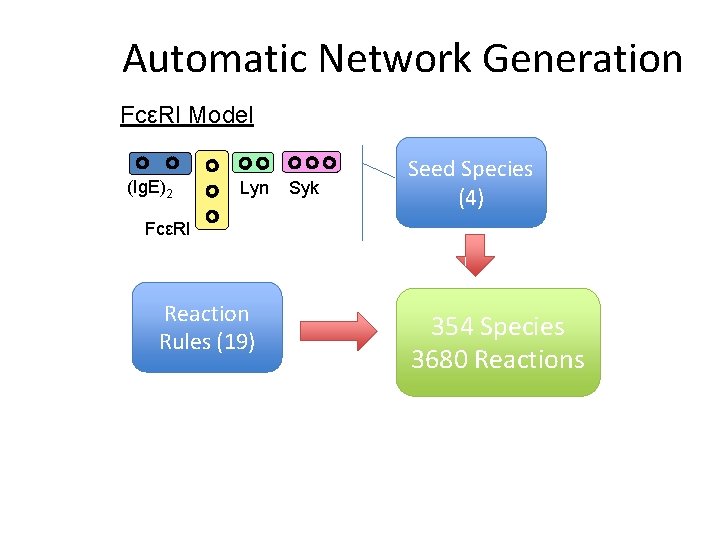
Automatic Network Generation FcεRI Model (Ig. E)2 Lyn Syk Seed Species (4) FcεRI Reaction Rules (19) 354 Species 3680 Reactions
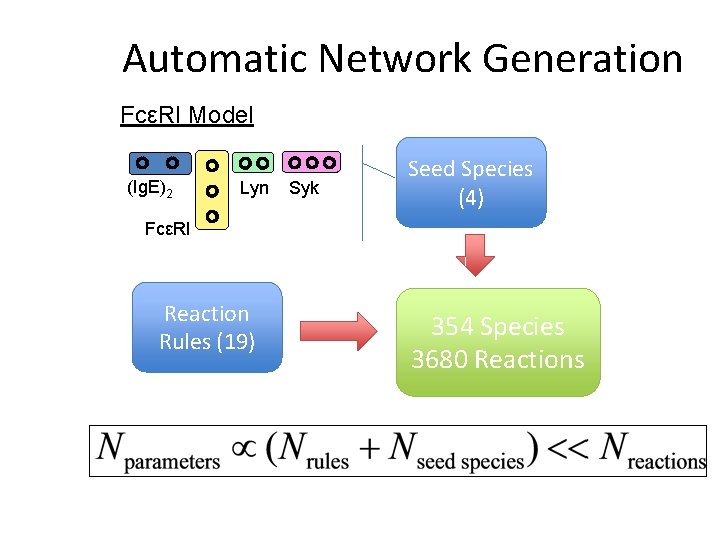
Automatic Network Generation FcεRI Model (Ig. E)2 Lyn Syk Seed Species (4) FcεRI Reaction Rules (19) 354 Species 3680 Reactions
- Slides: 37