An extended SNOMED CT concept model for observations

An extended SNOMED CT concept model for observations in molecular genetics James R. Campbell MDac Geoffrey Talmon MDad Allison Cushman-Vokoun MD Ph. Da Daniel Karlsson Ph. Dbc W. Scott Campbell Ph. Dad a. Departments of Internal Medicine and Pathology University of Nebraska Medical Center b. Department of Biomedical Engineering; Linkoping University; Linkoping, Sweden CObservables Project Group and di. PALM SIG; IHTSDO

Outline Ø Terminology challenges created by Personalized Medicine Ø What is an ‘Observable entity? Where do they fit in the SNOMED CT concept model? …the ONC terminology model? Ø LOINC – SNOMED CT harmonization model for observable entities Ø Anatomic and molecular pathology project at UNMC …Application of the harmonized model to genomics and molecular pathology? Ø Where do we go from here?

Revolution in healthcare Ø Sequencing of human genome has led to flood of new observational data appearing in scientific literature and now…clinical records Ø Majority of this clinical data is unstructured and not useful for decision support in the EHR or clinical research Ø Understanding of genetic and molecular basis of human disease now having impact on therapy Ø Challenge for precision medicine - to have sufficiently detailed molecular genetics observations and diagnostic data for clinical care and research

Limitations of ONC terminologies for observables in molecular pathology Research community Ø NCBI: Gene Ontology; HGNC; OMIM; Orphanet; Protein Ontology; Cell ontology Clinical community Ø SNOMED CT: – No concept model for subcellular anatomy (genes) or molecular structure (proteins) – No concept model for Observable entity or molecular basis of disease in Clinical findings – Little content Ø LOINC 2. 54: – 1275 PCR; 1406 MOLGEN; 116 FISH; 1502 CELL MARKERS – LOINC defining attributes inadequate to fully specify what is being resulted – Provides only tag-level interoperability of molecular data Ø No meaningful bridge joining genetic research findings with clinical concept models

Observable entity? ØIHTSDO “Observables are considered to be partial observation results, where there is a defined part of the observation missing. ” “Concepts in this hierarchy can be thought of as representing a question or procedure which can produce an answer or a result” “Observable + Evaluation (test) result = Finding” ØRegenstrief institute “…a set of universal names and ID codes for identifying laboratory and clinical test results” “…master file of standard ‘test’ names and codes that will cover most of the entries in these files of operational laboratory systems” “…organization of LOINC is divided first into four categories, ‘lab’, ‘clinical’ ‘attachments’ and ‘surveys’ ”

LOINC – SNOMED CT Harmonization of Observable entities Ø 2008 agreement between Regenstrief (RI) and IHTSDO https: //loinc. org/collaboration/ihtsdo/agreement. pdf – Extend and harmonize a shared concept model for 363787002|Observable entity| – Map and instantiate LOINC parts in SNOMED CT content – Jointly publish expression data set defining (lab) LOINC concepts within the harmonized concept model – Technology preview alpha January 2016 Ø Assertion: What is needed to support clinical decision making and research in molecular genetics is a unified domain ontology for Observable entities spanning clinical and research content including genomics

MP: Domain ontology Terminology Use Cases ØRetrieve all colon cancer tissue specimens which had BRAF gene testing (IHC or sequencing) ØRetrieve all breast cancer cases that tested ER-, PR- and Her 2/NeuØIdentify all colon cancer cases with high degree microsatellite instability by PCR or absence of mismatch repair protein by IHC ØIdentify all cases with Stage IIIa non-small cell lung cancer that had EGFR mutation testing by any method

UNMC: Project for structured encoding of AP/MP cancer reports Ø Objective: Detailed structured reporting of all anatomic and molecular pathology observations for all CAP synoptic cancer worksheets (82 types of malignancies) Ø Proposal: Analyze detailed semantics of CAP worksheets; apply harmonized concept model to develop terminology requirements; deploy as real-time structured reporting from COPATH system interfaced to tissue biobank and EPIC Ø Tooling: Nebraska Lexicon© extension namespace; SNOWOWL authoring platform; SNOMED CT International + US Extension + Technology preview; ELK 0. 4. 1 DL classifier Ø Penciled into IHTSDO workplan for 2017

Fully Encoded Checksheet
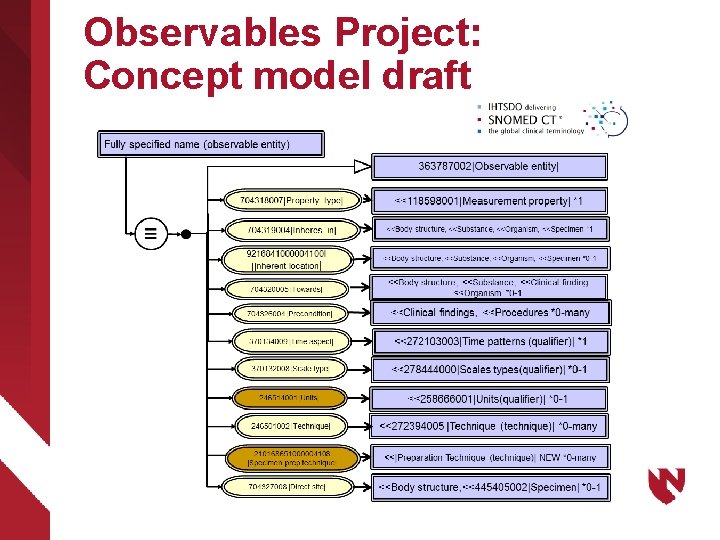
Observables Project: Concept model draft

SNOMED extensions for MP: Genes and proteins Ø Body structures>>Cell structures>>Nucleotide sequences and Genes Ø Substances>>Proteins Ø Qualifiers>>Techniques>> AP and MP methods Ø Qualifiers>>Measurement Properties>>AP and MP properties Ø Observable entity>>AP and MP observables Ø Clinical findings>>Anatomic and molecular genetic observation results and disorders
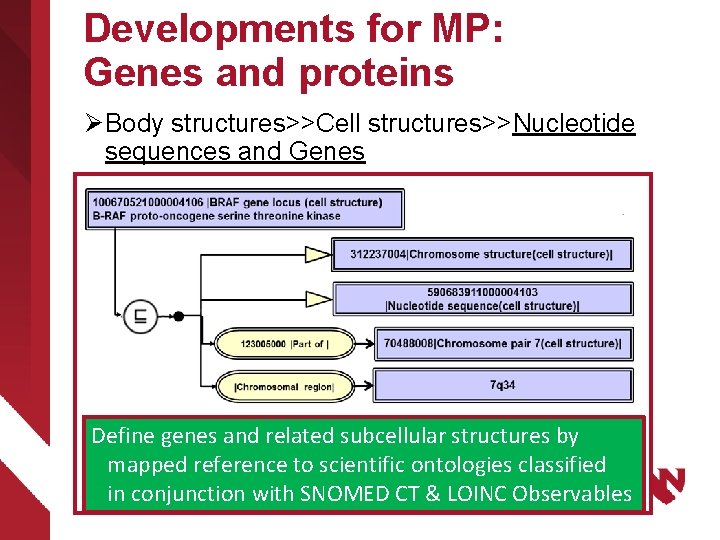
Developments for MP: Genes and proteins Ø Body structures>>Cell structures>>Nucleotide sequences and Genes Ø Substances>>Proteins Ø Qualifiers>>Techniques>> AP and MP methods Ø Qualifiers>>Measurement Properties>>AP and MP properties Ø Observable entity>>AP and MP observables Ø Clinical findings>>Anatomic and molecular genetic observation results and disorders Define genes and related subcellular structures by mapped reference to scientific ontologies classified in conjunction with SNOMED CT & LOINC Observables
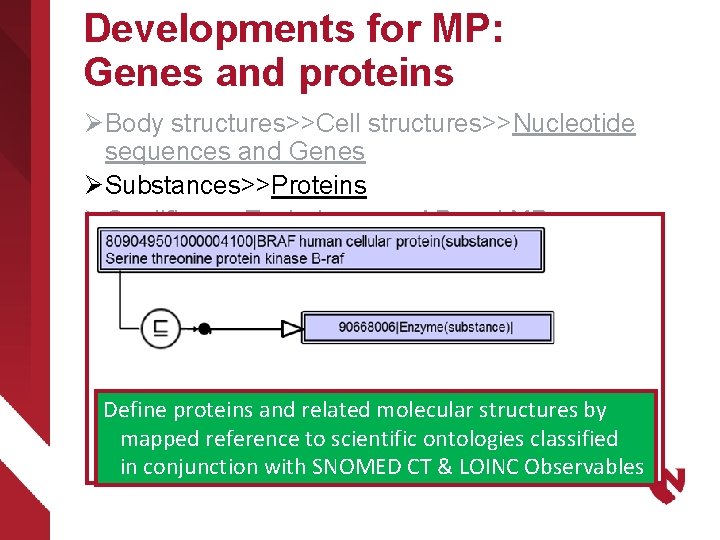
Developments for MP: Genes and proteins Ø Body structures>>Cell structures>>Nucleotide sequences and Genes Ø Substances>>Proteins Ø Qualifiers>>Techniques>> AP and MP methods Ø Qualifiers>>Measurement Properties>>AP and MP properties Ø Observable entity>>AP and MP observables Ø Clinical findings>>Anatomic and molecular genetic observation results and disorders Define proteins and related molecular structures by mapped reference to scientific ontologies classified in conjunction with SNOMED CT & LOINC Observables
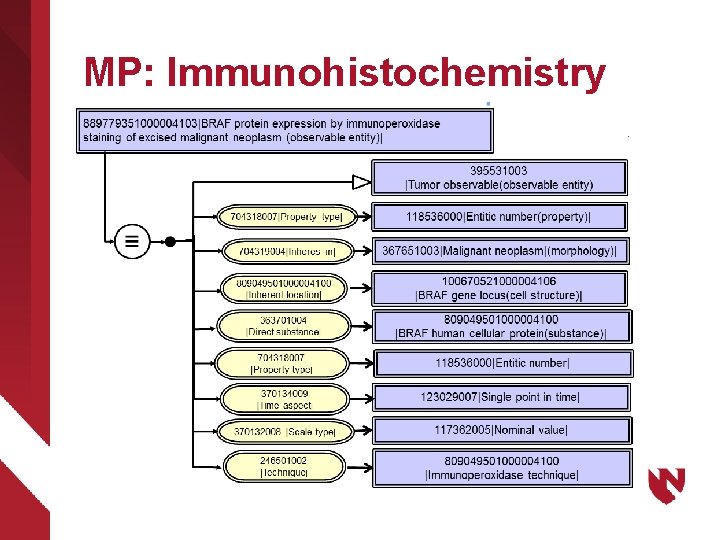
MP: Immunohistochemistry
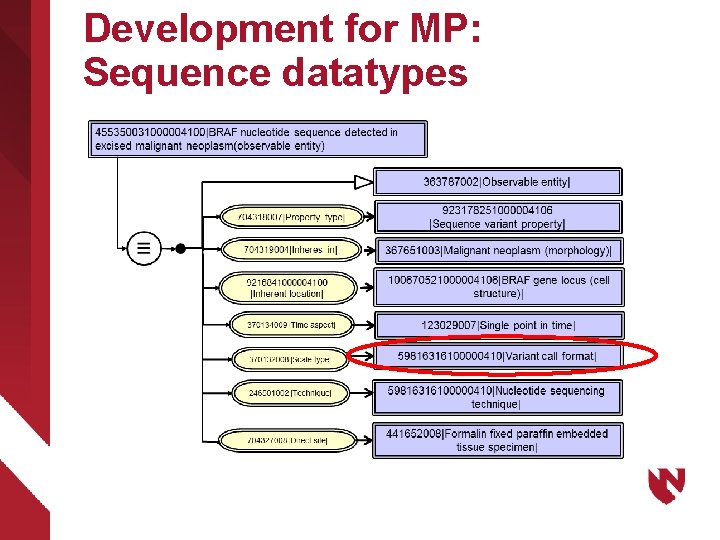
Development for MP: Sequence datatypes
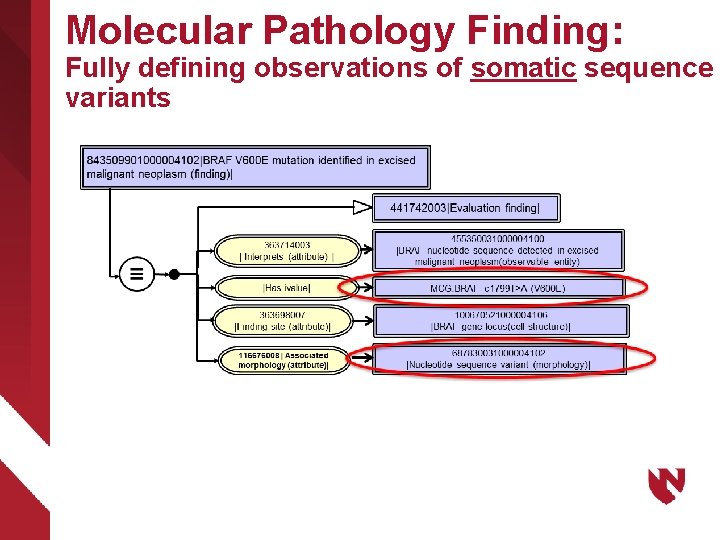
Molecular Pathology Finding: Fully defining observations of somatic sequence variants
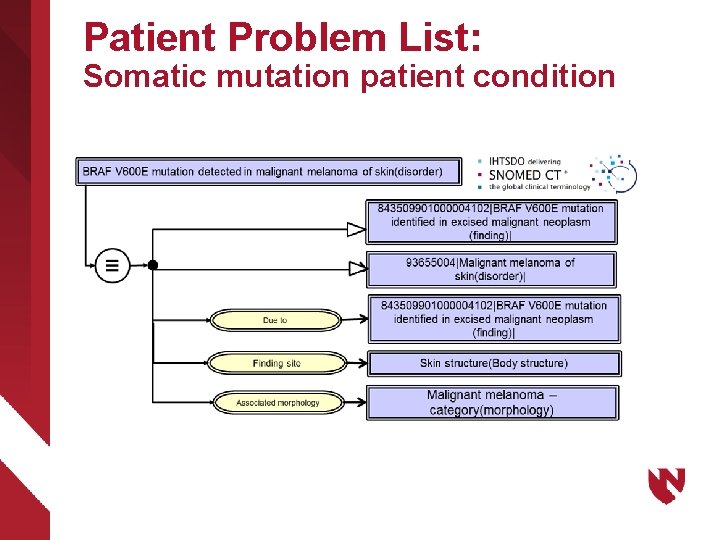
Patient Problem List: Somatic mutation patient condition

Congenital (germline) mutations “Familial adenomatous polyposis” Cancer predilection syndrome found to originate with multiple genetic etiologies: ØFAP 1: heterozygous germline mutation of the APC gene ØGardner syndrome: clinical variant of FAP 1; includes desmoid tumors and other hamartomas; mutation of APC gene ØFAP 2: mutation of MUTYH ØFAP 3: mutation of MTHL 1 ØFAP 4: mutation of MSH 3
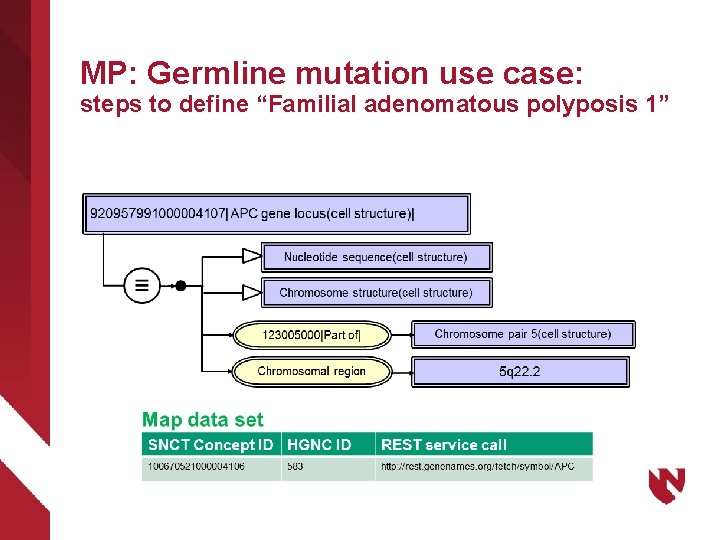
MP: Germline mutation use case: steps to define “Familial adenomatous polyposis 1”
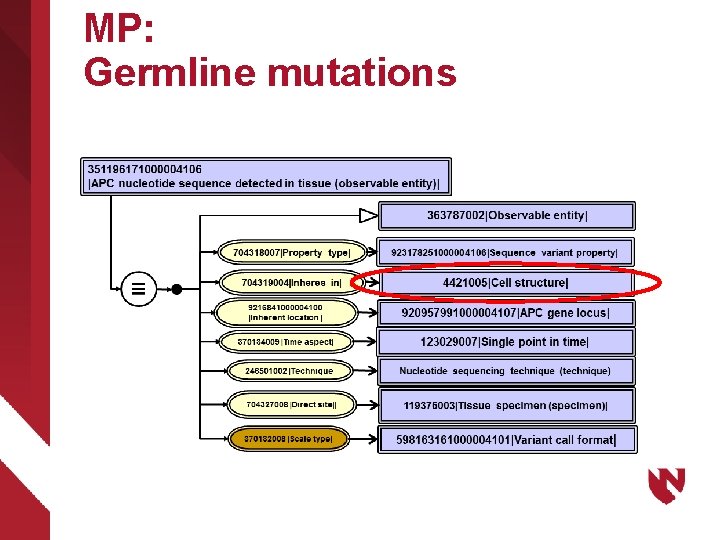
MP: Germline mutations
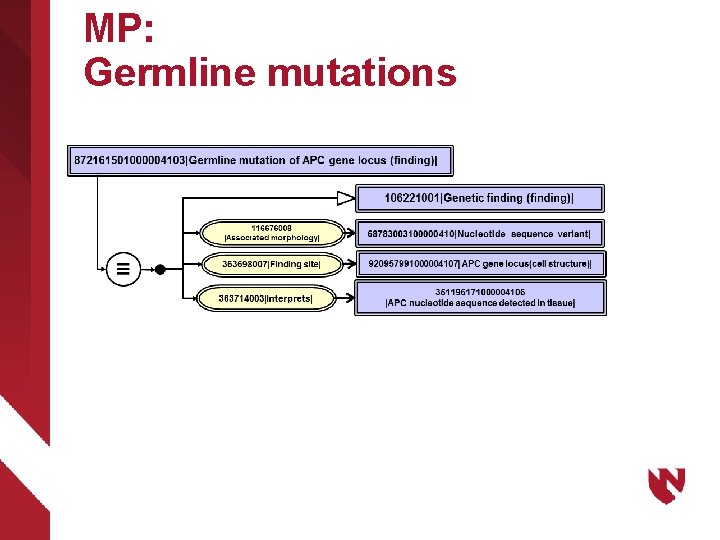
MP: Germline mutations
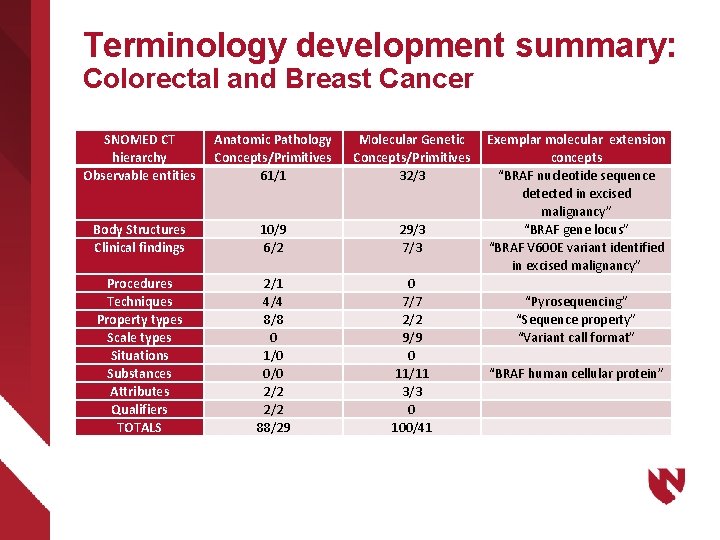
Terminology development summary: Colorectal and Breast Cancer SNOMED CT hierarchy Observable entities Anatomic Pathology Concepts/Primitives 61/1 Molecular Genetic Concepts/Primitives 32/3 Body Structures Clinical findings 10/9 6/2 29/3 7/3 Procedures Techniques Property types Scale types Situations Substances Attributes Qualifiers TOTALS 2/1 4/4 8/8 0 1/0 0/0 2/2 88/29 0 7/7 2/2 9/9 0 11/11 3/3 0 100/41 Exemplar molecular extension concepts “BRAF nucleotide sequence detected in excised malignancy” “BRAF gene locus” “BRAF V 600 E variant identified in excised malignancy” “Pyrosequencing” “Sequence property” “Variant call format” “BRAF human cellular protein”

Deployment of MP data ØHL 7 interface to tissue biobank for indexing of tissue for research community ØETL of data to i 2 b 2 data warehouse ØPublication of laboratory/pathology observables ontology with NLM and RI ØInterface development and extension to structured pathology data for EHR
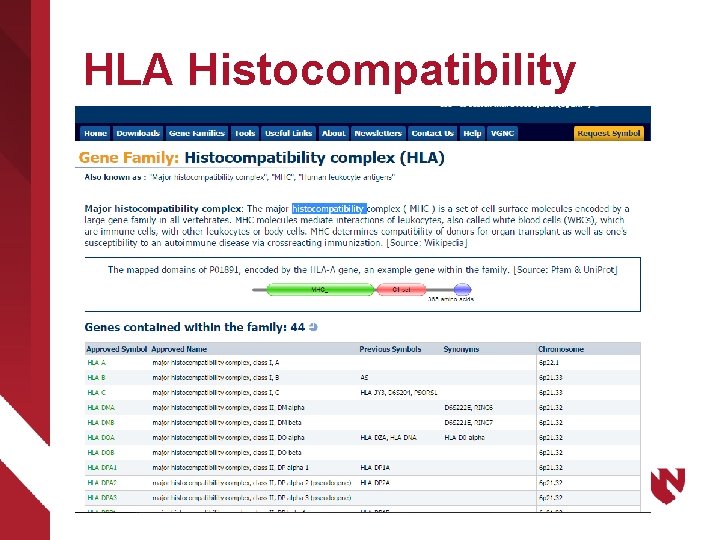
HLA Histocompatibility

Gene and associated protein ØHLA-A gene locus(cell structure) – ISA Nucleotide sequence – ISA Chromosome structure – Part of Chromosome pair 6 – Chromosome region 6 p 22. 1 Ø 49495002|HLA-A antigen(substance)|

- Slides: 28