AN EVALUATION OF THE OECD QSAR TOOLBOX PROFILERS

AN EVALUATION OF THE OECD QSAR TOOLBOX PROFILERS FOR IDENTIFYING DNA REACTIVE AND GENOTOXIC CHEMICALS Terry Schultz Professor Emeritus, The University of Tennessee-Knoxville & Secretariat, OECD, Paris

Outlook • The Toolbox • 1 st Exercise: Goal, Methods & Results • 2 nd Exercise: Goal, Methods & Results • 3 rd Exercise: Goal, Methods & Results • Conclusions 2

The OECD QSAR Toolbox The seminal feature of the Toolbox is its ability to quickly evaluate all members of a category for common toxicological behaviour or consistent trends within important regulatory endpoint data. This is done by first profiling the chemical by type, class or defined hazard identifiers and then selecting a category or subcategory. This feature often links the chemicals in the category to a single mechanism or mode of action. 3

Advantages of the Profile/Category Approach 1) it shifted emphasis to intrinsic chemical activity, 2) it allows for entire categories of chemicals to be assessed when only a few members are tested, 3) it allows for filling data gaps using read-across and trend analysis, and not just (Q)SAR models, and 4) it enables defensible hazard assessment through mechanistic comparisons without testing. 4

1 st Mc. Kim Workshop on Reducing Data Redundancy in Cancer Assessment The participants agreed that chemical which are QSAR predicted to be DNA reactive (+) and are also AMES test positive (+) would be chemicals of “very high concern” for the Rodent Cancer Assay. But you can have direct or indirect DNA-reactivity and AMES positive w/o S 9 and w/ S 9. So the first exercise was to look for members of the direct DNA-reactivity and AMES positive w/o S 9 category. 5
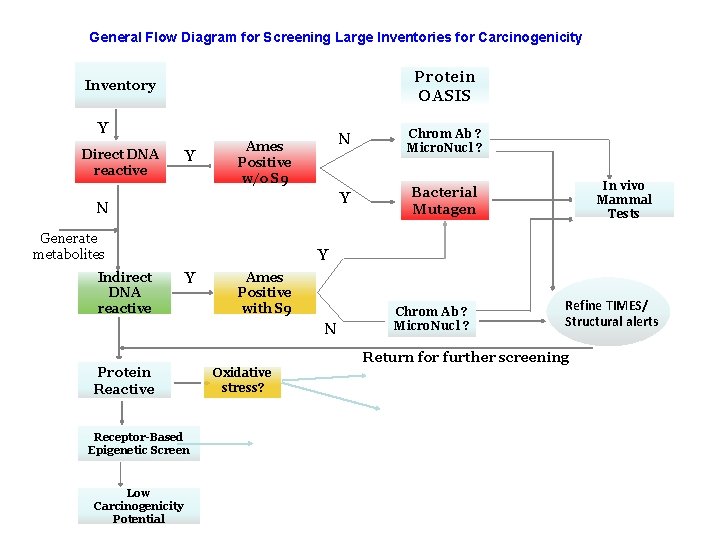
General Flow Diagram for Screening Large Inventories for Carcinogenicity Protein OASIS Inventory Y Direct DNA reactive Y Ames Positive w/o S 9 N Generate metabolites Indirect DNA reactive N Chrom Ab ? Micro. Nucl ? Y Bacterial Mutagen In vivo Mammal Tests Y Y Ames Positive with S 9 N Protein Reactive Chrom Ab ? Micro. Nucl ? Refine TIMES/ Structural alerts Return for further screening Oxidative stress? Receptor-Based Epigenetic Screen Low Carcinogenicity Potential 6
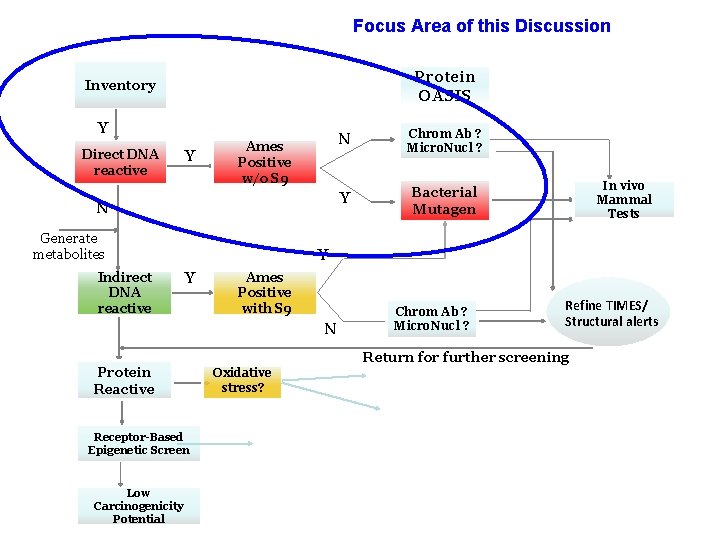
Focus Area of this Discussion Protein OASIS Inventory Y Direct DNA reactive Y Ames Positive w/o S 9 N Generate metabolites Indirect DNA reactive N Chrom Ab ? Micro. Nucl ? Y Bacterial Mutagen In vivo Mammal Tests Y Y Ames Positive with S 9 N Protein Reactive Chrom Ab ? Micro. Nucl ? Refine TIMES/ Structural alerts Return for further screening Oxidative stress? Receptor-Based Epigenetic Screen Low Carcinogenicity Potential 7
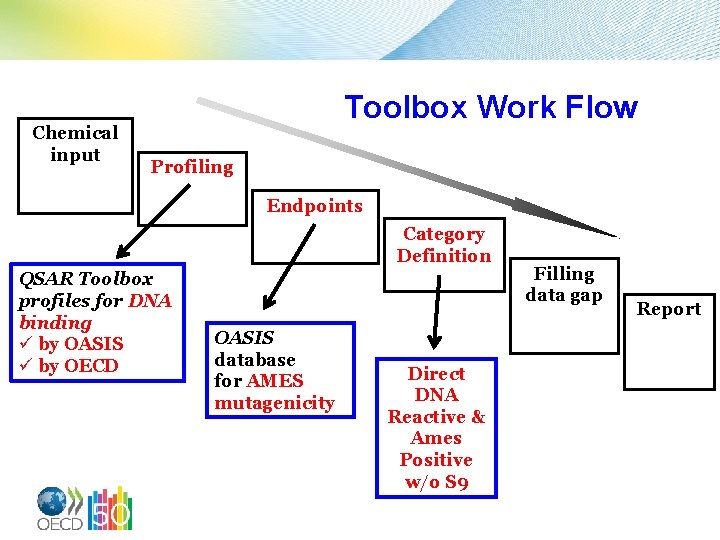
Chemical input Toolbox Work Flow Profiling Endpoints Category Definition QSAR Toolbox profiles for DNA binding ü by OASIS ü by OECD OASIS database for AMES mutagenicity Direct DNA Reactive & Ames Positive w/o S 9 Filling data gap Report

Factors Which Effect Makeup of Chemical Category How one defines the initial target chemical. For Example: Using CH 4 as the initial target chemical and profiling for chemicals containing a C-atom one retrieves 8684 chemicals from the Toolbox. Whether one used DNA-binding OASIS or DNAbinding OECD. This impacts the number and definitions of the SAs which are fired. 9
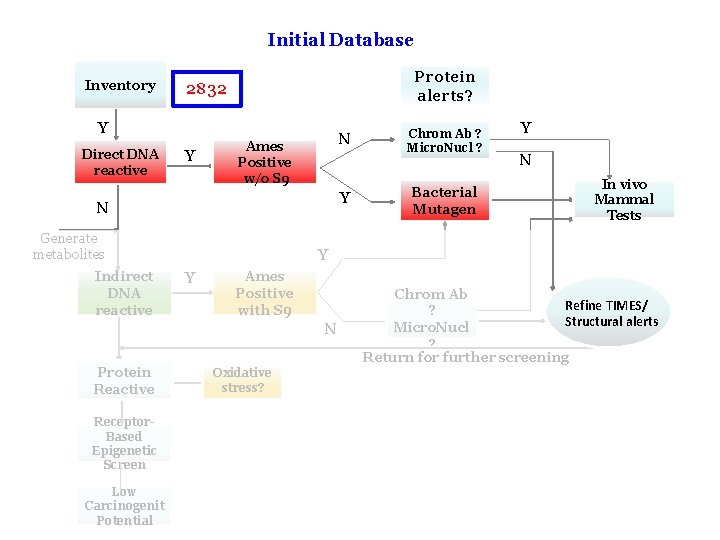
Initial Database Inventory Protein alerts? 2832 Y Direct DNA reactive Y N Ames Positive w/o S 9 Y N Generate metabolites Indirect DNA reactive Bacterial Mutagen Y N In vivo Mammal Tests Y Y Ames Positive with S 9 N Protein Reactive Chrom Ab ? Micro. Nucl ? Chrom Ab Refine TIMES/ ? Structural alerts Micro. Nucl ? Return for further screening Oxidative stress? Receptor. Based Epigenetic Screen Low Carcinogenit Potential 10
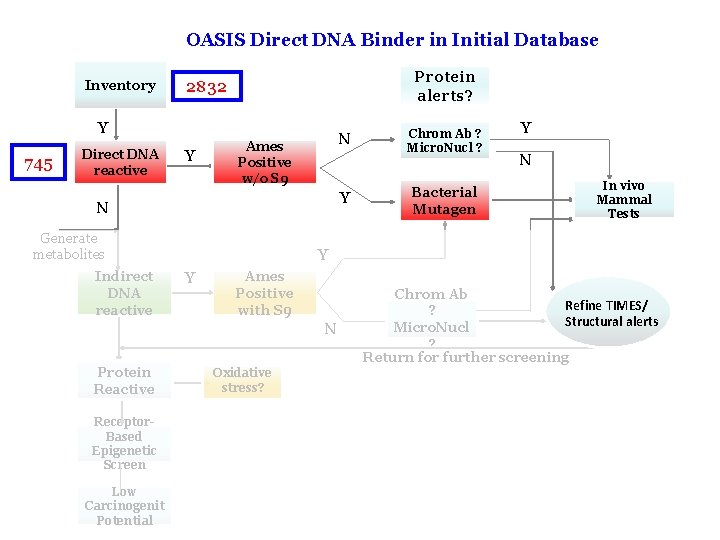
OASIS Direct DNA Binder in Initial Database Inventory Protein alerts? 2832 Y 745 Direct DNA reactive Y N Ames Positive w/o S 9 Y N Generate metabolites Indirect DNA reactive Bacterial Mutagen Y N In vivo Mammal Tests Y Y Ames Positive with S 9 N Protein Reactive Chrom Ab ? Micro. Nucl ? Chrom Ab Refine TIMES/ ? Structural alerts Micro. Nucl ? Return for further screening Oxidative stress? Receptor. Based Epigenetic Screen Low Carcinogenit Potential 11
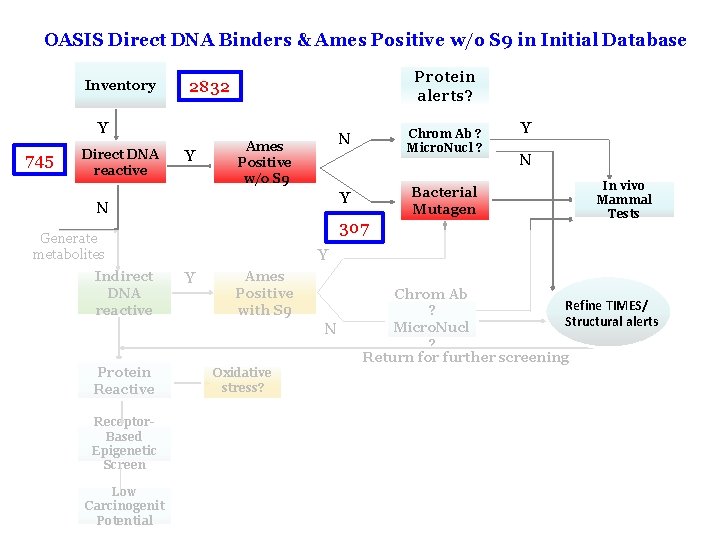
OASIS Direct DNA Binders & Ames Positive w/o S 9 in Initial Database Inventory Protein alerts? 2832 Y 745 Direct DNA reactive Y Ames Positive w/o S 9 307 Generate metabolites Y N In vivo Mammal Tests Y Y Ames Positive with S 9 N Protein Reactive Bacterial Mutagen Y N Indirect DNA reactive Chrom Ab ? Micro. Nucl ? N Chrom Ab Refine TIMES/ ? Structural alerts Micro. Nucl ? Return for further screening Oxidative stress? Receptor. Based Epigenetic Screen Low Carcinogenit Potential 12
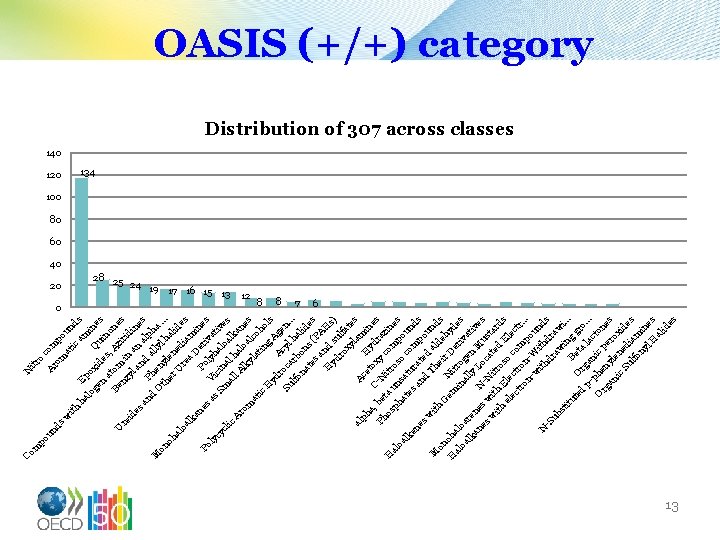
nd po u N itr o Ar com om p sw at oun ith ic d am s ha Ep lo ge oxid Qu ines n at es, ino Az ne o B U en m s ir re i z yl n a idin id es an n al es an d P ph a d h M O en llyl a. on th. y oh er len hal. ed ide al U oa re a iam s lk D an Po er ine es Po s i as Vi lyh vat ly c cy i a Sm in lo ve cl ic al al h alk s l. A a Ar a om lk loa nes y l la co at tin ho ic H l g yd A s ro Ac gen Su ca yl lfo rb ha. . . na on lid te s ( es sa P A H nd Hs yd su ) ro lfa xy t la es al Ac m ph H in e a, C- tox ydr es b Ph e a N y H os ta u itr com zin al o ph n s p es oa at sat o c ou lk es o n u en an rat mp ds es e o d w Th d al und M ith d ei on s e G em Ni r D hyd H oha t e al in rog riv es oa loa al lk re ly en ativ an ne M L us es es s w N-N oc a t w ith itro ted ard E so El s el l e ec ect c ro om ctr. tr on n. . p -w W ou n i ith th N d -S dr dr s ub a aw w st i. i itu Be ng g. . te O ta r d r p- gan la o. . . c p to ic h O rg en pe ne s an yl r ic ene oxi Su d de lfo iam s ny in l H es al id es m Co OASIS (+/+) category Distribution of 307 across classes 140 120 20 0 134 100 80 60 40 28 25 24 19 17 16 15 13 12 8 8 7 6 13

DNA-binding OECD Eliminated: Nitro Aromatic Compounds, Aromatic Amines, PAHs etc. Classes where it is unclear that a bacterial test would mimic the reactions well. 14

nd 2 Exercise Designed to look for which members of the +/+ category have been evaluated in a Rodent Cancer Assay (RCA) and to report what classes they represent. 15
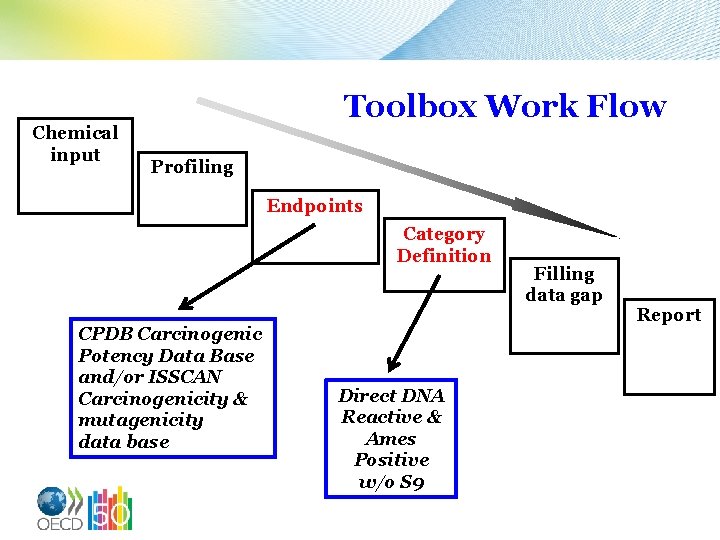
Chemical input Toolbox Work Flow Profiling Endpoints Category Definition CPDB Carcinogenic Potency Data Base and/or ISSCAN Carcinogenicity & mutagenicity data base Direct DNA Reactive & Ames Positive w/o S 9 Filling data gap Report
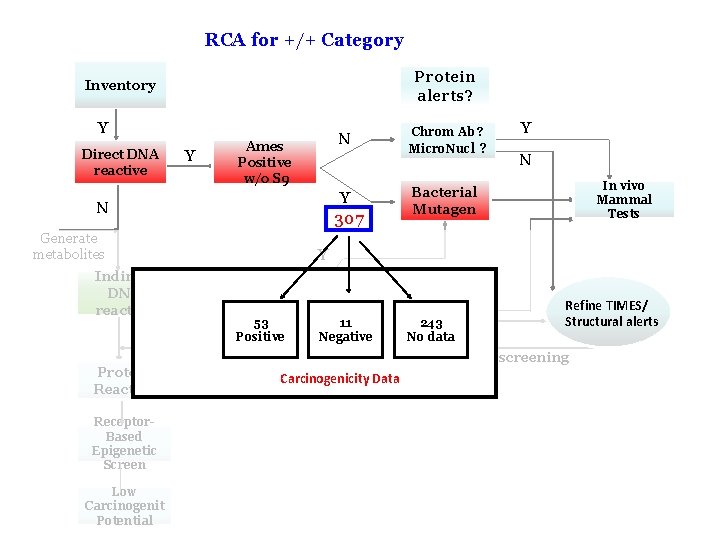
RCA for +/+ Category Protein alerts? Inventory Y Direct DNA reactive Y N Ames Positive w/o S 9 Y 307 N Generate metabolites Indirect DNA reactive Protein Reactive Chrom Ab ? Micro. Nucl ? Bacterial Mutagen Y N In vivo Mammal Tests Y Y Ames Positive with S 9 Chrom Ab Refine TIMES/ ? Structural alerts 53 11 243 Micro. Nucl N Positive Negative No data ? Return for further screening Oxidative Carcinogenicity Data stress? Receptor. Based Epigenetic Screen Low Carcinogenit Potential 17
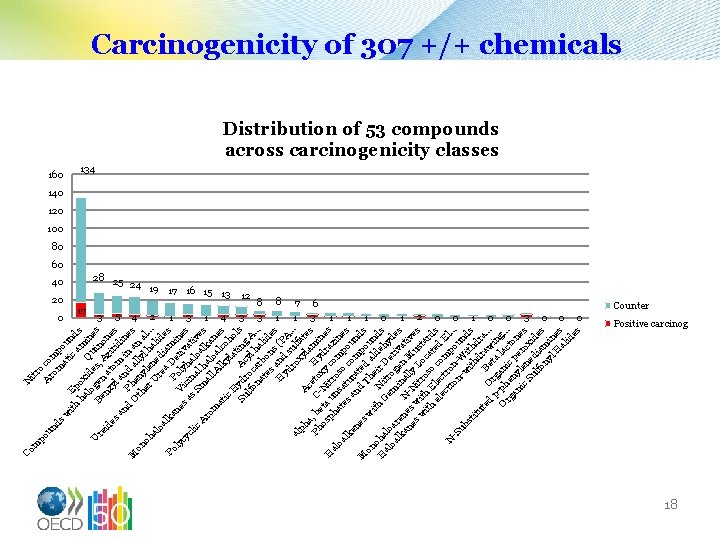
N itr o po c A un ro om ds m po at u w ic nd ith Ep am s ha ox lo ide Qu ine U Be gen s, in s re nz a Az on id es yl tom iri es d an an in ine P d d M O he al an s on th ny ly er le l h al. . oh U ne ali. al re di d oa a am es lk Po D a P ne V o er in ly l cy s a ic yh iva es cl s S ina alo tiv ic m l h al es Ar al al ka om l A oa ne at lk lco s ic yl h a o H Su ydr Ac ting ls lfo oc yl A na ar ha. . . te bo lid s ns es H and (P yd s A ro ul. . . al xy fat ph Ac la e a, Ph b C eto Hy mi s os eta -Ni xy dr nes H ph u tr co az al at nsa oso m ine oa es tu c po s lk an ra om un en d ted p ds es M Th a ou on w i ei ld nd t o h N H ha G itr r D ehy s al lo em o er d oa ar lk en N in gen iva es a ti an e es s w -Ni lly L Mu ves s w ith tro oc ta ith E s o a rd el lec co ted s ec tr m E tr on p l. . N on - ou. -S ub -w W nd st ith s itu dr dra te d Or Bet aw. . . p g a in O -ph ani lac g. . rg e c. an ny pe ton ic len ro es Su ed xi lfo ia des ny mi l H ne al s id es m Co Carcinogenicity of 307 +/+ chemicals Distribution of 53 compounds across carcinogenicity classes 160 20 0 134 140 120 100 80 60 40 28 25 24 19 17 16 15 13 12 17 3 5 4 2 1 3 1 4 3 8 7 1 1 6 3 1 1 1 0 1 2 0 0 1 0 0 3 0 0 0 Counter Positive carcinog 18

RD 3 EXERCISE Designed to look at which DNA-reactions are represented by the (+/+) chemicals that are also positive in the RCA. 19
![Major Classes Aliphatic Halides [CX]; 17 CRA positive; 2 CRA negative (1 -chlorobutane; 2 Major Classes Aliphatic Halides [CX]; 17 CRA positive; 2 CRA negative (1 -chlorobutane; 2](http://slidetodoc.com/presentation_image_h/c3f58a507588cf247ffb51af908d8cbb/image-20.jpg)
Major Classes Aliphatic Halides [CX]; 17 CRA positive; 2 CRA negative (1 -chlorobutane; 2 chloroethanol) Stressed Hetero-Ring Systems (e. g. C 1 CO 1); 13 CRA positive 20

Minor Classes SN 2 addition at an sp 3 carbon atom sulfates and sulfonates; 4 CRA positive Phosphorates; 2 CRA positive Hydazines; 1 CRA positive 21

Summary In this discussion we used a hypothesis testing scheme with the QSAR Toolbox; the DNAbinding profiler generate the initial hypothesis and an in vitro test (AMES testing w/o S 9) tested the hypotheses. 22

Summary We maintain that chemicals that pass this hypothesis testing are “chemicals of high concern” for cancer. 23

Summary We then asked which of these chemicals of high concern have been tested in the RCA? What are the DNA reactions covered by this suite of RCAtested chemical? 24

Conclusion There is experimental evidence that there are opportunities to reduce the use of RCA, especially for chemicals with an aliphatic halide or stressed hetero-ring sub-structure. 25
- Slides: 25