AMOS file format afg LIB iid 453 eid

AMOS file format (. afg) {LIB iid: 453 eid: 17000001585820 {DST mea: 3000. 000 std: 166. 667 } This is an insert “library” with mean insert length of 3000 bp, and a standard deviation of 166. 667 bp. The library ID is 453

AMOS file format (. afg) {FRG iid: 456 eid: 90 lib: 453 rds: 88, 89 typ: I } This “fragment” is a clone insert, from which both ends have been sequenced Its internal ID is 456 It came from Library 453 (which has an insert length of 3000 bp) Its ends are identified by the two reads with internal IDs 88 and 89 Its ends face “Inward”, 5’ on the outside and 3’ on the inside

AMOS file format (. afg) This is read ID 88 {RED iid: 88 eid: 17000001585880 f seq: GCCACGTAGGCGTTTTGGAAATTAGCCGCCTCGGGCGTCGCATTGCTCAAGGGACTAATTTCAGCG GCCCTGTGATGTGGCCTGTCGGTGGGGGTGTGGTGAGGAGTTCGCGAACCTGATCGTCGAGTAGATCTGT CCAACCGTCATCAAACGCGGATATCAATGGGTTGCGCACACCACATCGTAGGCTTCGTGCGATCTCACGG CCAGGCTGTTGGCCCGACCGGTATCGTGACAATTATTGATTTGGGGGGTCGAGCGGGTCTCGTGGC CCGTAAGTTACGGCGGCCGTCAGCATGCTGGCGCCGGTGGCTATGCCGTCATCGACGGGGGTCAC GGTCCTGCCGTGTGGGTCGGCCGACGGTGCGCTTGCCCCTATACATCCGTTTGCATCGCATGAGTGCCAC TGTCTCCTTGTCAATCACTCGTGCGAGTCAGCATCGGACGGGGCATTGTTGGGGTATTGAGGCCTTGGGT GGTGGTGTTGTG. qlt: KKKKK 7 IK: KKKKKKA 9 KKKK 5 KKKKKKKT; KKKKQKLKKKKKKFKKKK<E<K: KKKKNKK 9= FK<KKK@KKLKOKKKKK: KKKKJK 5? KKKKMLKKK 8 IKKTKKKKF@KKTK=KK 5@UKBKKUADDKKEKH< EKDUKKK; KPKKKBKK 9 TKKPKK@? KKGKKKKTKKKKKUK 9 KKK>LK 5 KKKKK 9 KK 8 KFO; KKKQK KKKKKKKTKKK 5 FKKKKKUKKKK 8 RKKKQTKKKFKKPSKKKKKK: KKKK<KKOKKK= KPKKKKKIHBKKKK<NKBKKKKK; KKKKK 6 DKKKKK=KKKKSKKKKKUKKEKKKKKKHKPIKRKKG KOKKMKKKK 5 K>KKOKS 6: KKCKKSK<KKKN@TKKKKK? QKKK>PK>KLGKKKKMUKKDKKK KKKKKKK 9 KKKKKKK; KK 7 KTNKQKKKKJBKNKUKK 7 K 99 OKKKKK 7 KKKKKKDKKKKKPK 7 HAKK KKKKKKUTKUKK. frg: 456 clr: 0, 502 } It comes from fragment (pair) 456 The high-quality (“clear”) part of the read is from 0 -502

Lab 2: identifying the species just BLAST it http: //www. ncbi. nlm. nih. gov/BLAST/ Suggestion: If you want a fast answer, set BLAST to use a word size of 15, and set “expect” to a small value such as 0. 00001. Or use Megablast.

Running AMOScmp Input files: lab 02. afg, lab 02. 1 con $ AMOScmp lab 02 The log file is: lab 02. run. Amos. log Doing step 10: Building AMOS bank Doing step 20: Collecting clear range sequences Doing step 30: Running nucmer Doing step 40: Running layout Doing step 50: Running consensus Doing step 60: Outputting contigs Doing step 70: Outputting fasta Files created: lab 02. bnk (a directory) lab 02. conflict lab 02. delta (created by nucmer) lab 02. layout lab 02. seq lab 02. cluster (created by nucmer) lab 02. contig lab 02. fasta lab 02. run. Amos. log
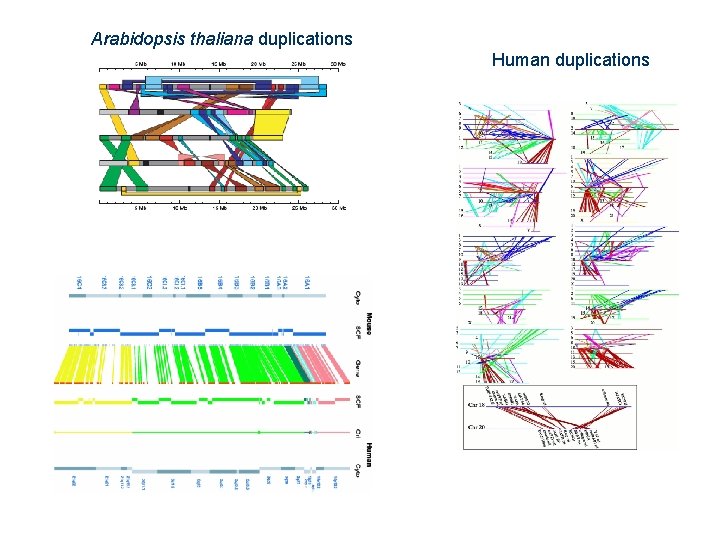
Arabidopsis thaliana duplications Human duplications
- Slides: 6