AMINO ACIDS Structure of amino acids carbon Aminoacetic

AMINO ACIDS
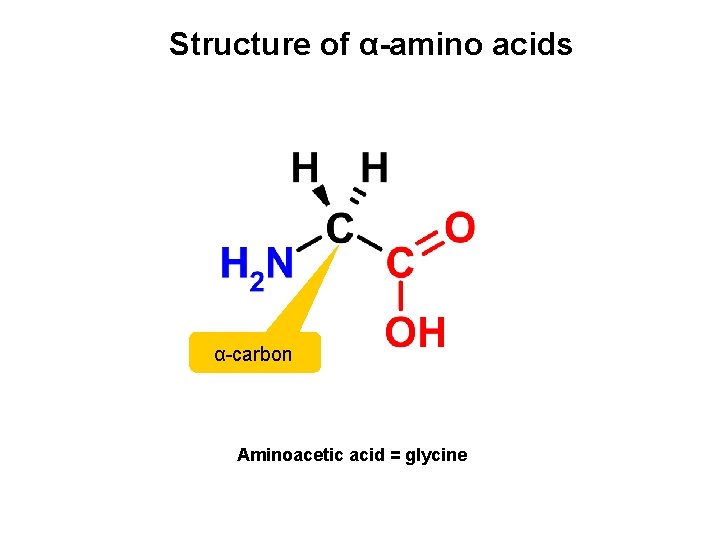
Structure of α-amino acids α-carbon Aminoacetic acid = glycine
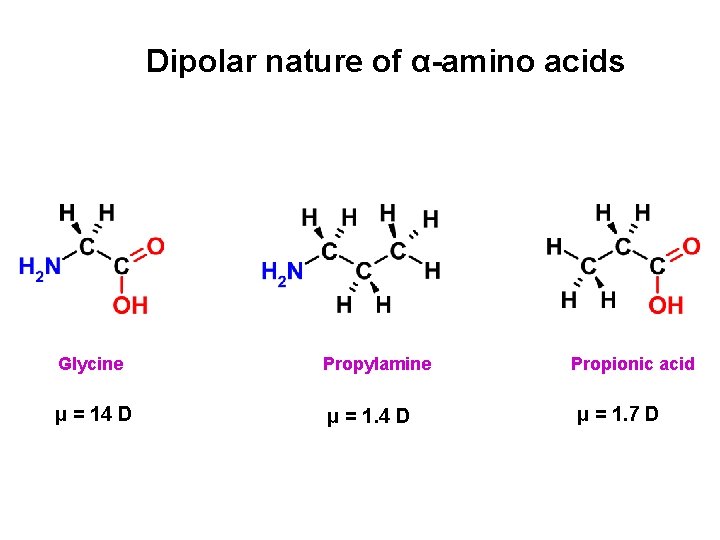
Dipolar nature of α-amino acids Glycine Propylamine μ = 14 D μ = 1. 4 D Propionic acid μ = 1. 7 D
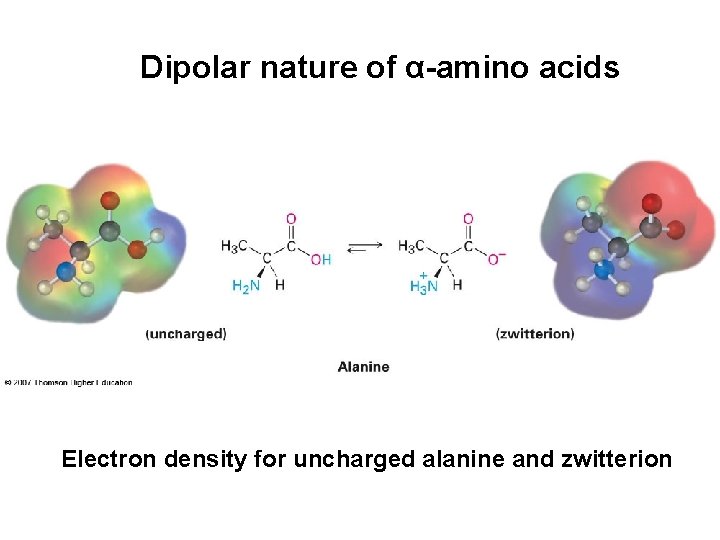
Dipolar nature of α-amino acids Electron density for uncharged alanine and zwitterion

Dipolar nature of α-amino acids determines their properties • They are solids with high melting points – glycine m. p. 262°C • They are much better soluble in water than in organic solvents • Their dipole moments are high • They are less acidic than carboxylic acids and less basic than amines
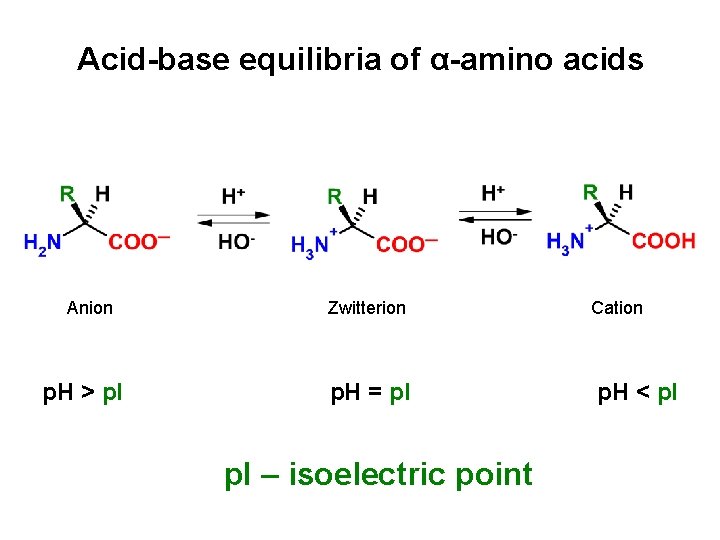
Acid-base equilibria of α-amino acids Anion p. H > p. I Zwitterion p. H = p. I – isoelectric point Cation p. H < p. I

At p. H correspondig to p. I value of given amino acid: • Predominating form is zwitterion • Molecule is electrically uncharged • Amino acid shows the lowest solubility in water • At p. H < p. I predominating form is a cation • At p. H > p. I predominating form is an anion
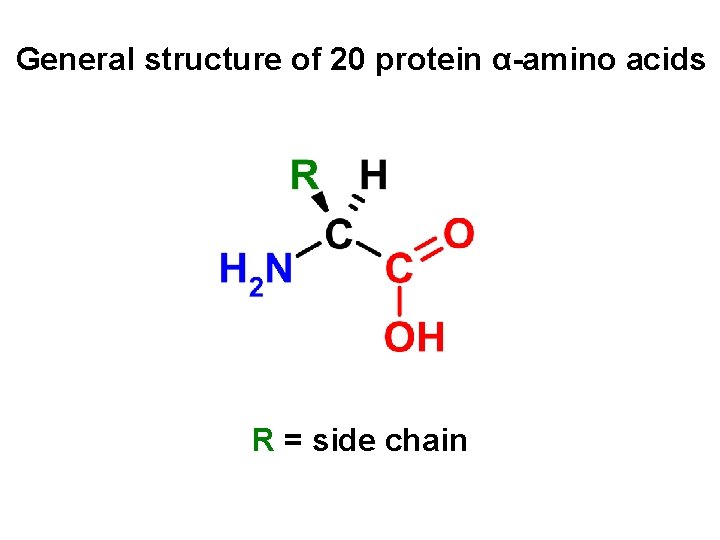
General structure of 20 protein α-amino acids R = side chain
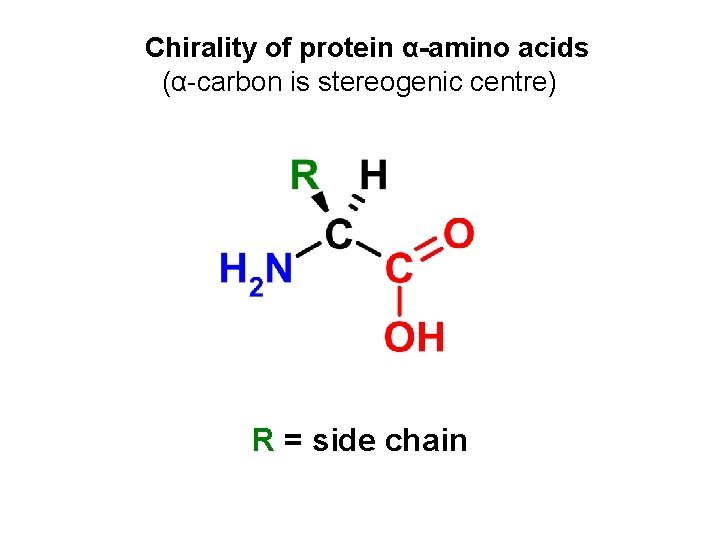
Chirality of protein α-amino acids (α-carbon is stereogenic centre) R = side chain
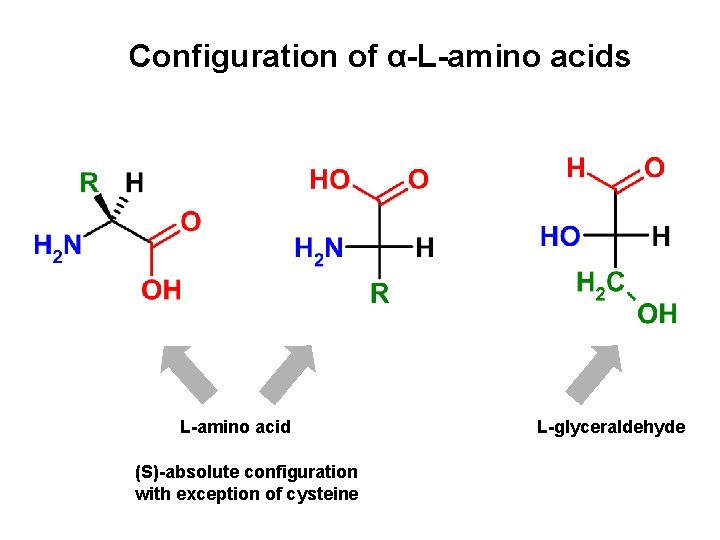
Configuration of α-L-amino acids L-amino acid (S)-absolute configuration with exception of cysteine L-glyceraldehyde
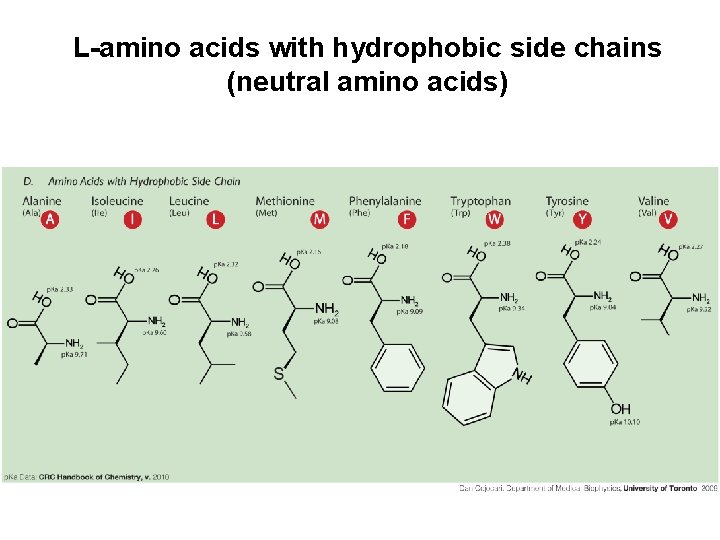
L-amino acids with hydrophobic side chains (neutral amino acids)
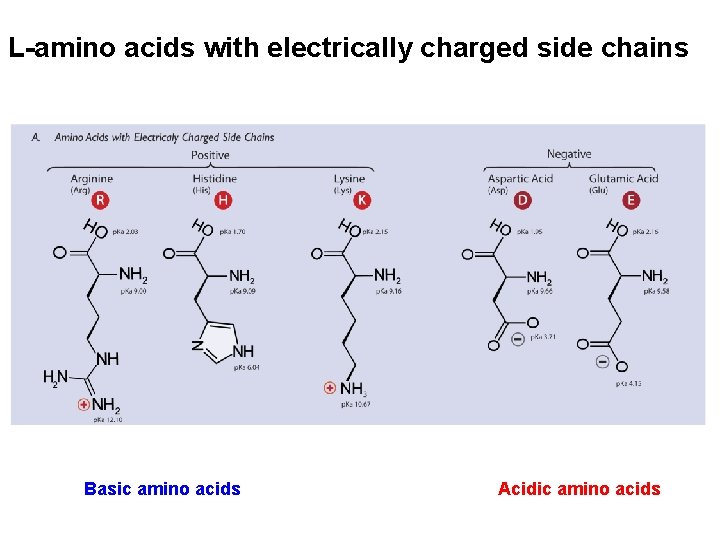
L-amino acids with electrically charged side chains Basic amino acids Acidic amino acids
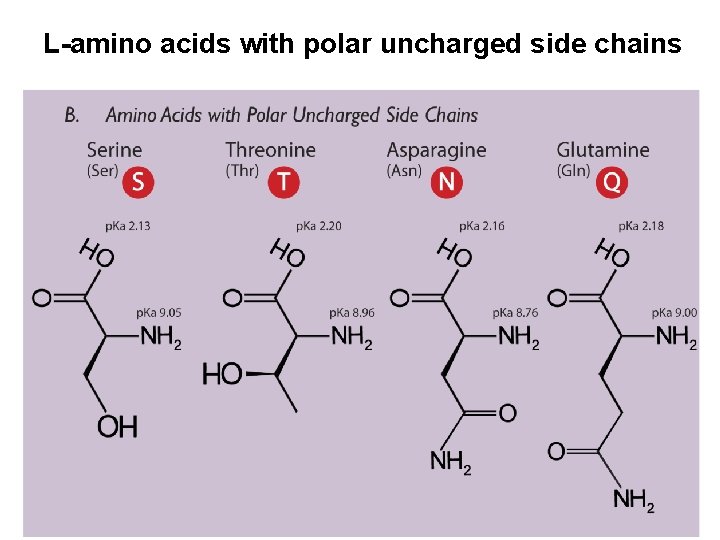
L-amino acids with polar uncharged side chains
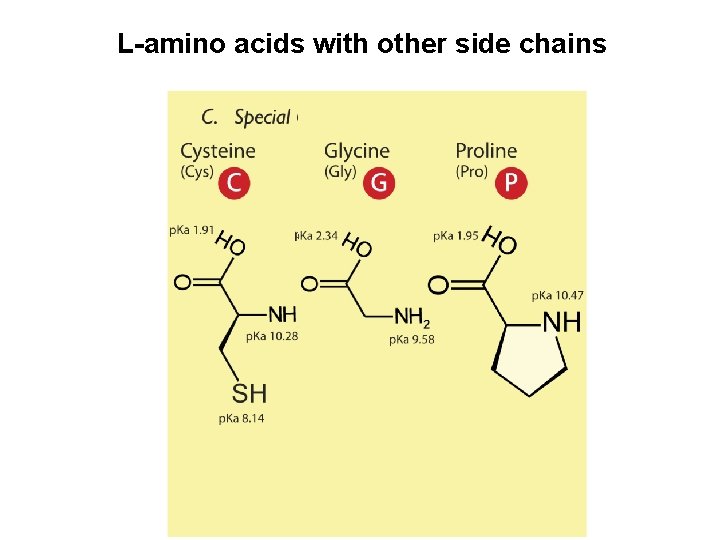
L-amino acids with other side chains
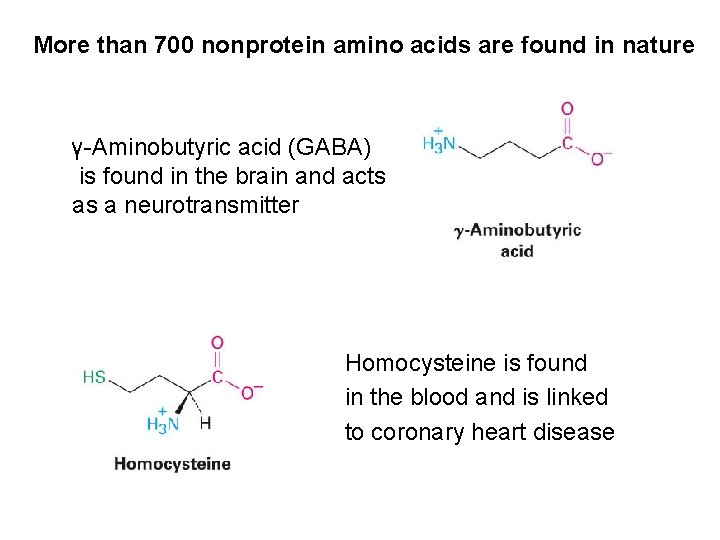
More than 700 nonprotein amino acids are found in nature γ-Aminobutyric acid (GABA) is found in the brain and acts as a neurotransmitter Homocysteine is found in the blood and is linked to coronary heart disease
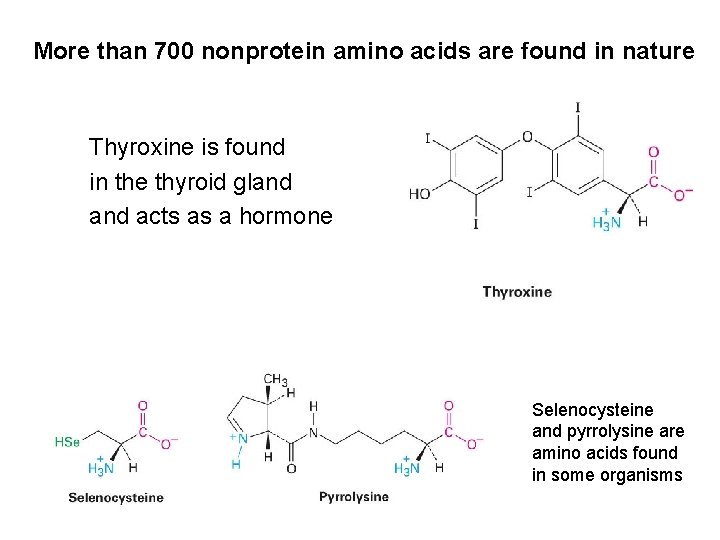
More than 700 nonprotein amino acids are found in nature Thyroxine is found in the thyroid gland acts as a hormone Selenocysteine and pyrrolysine are amino acids found in some organisms
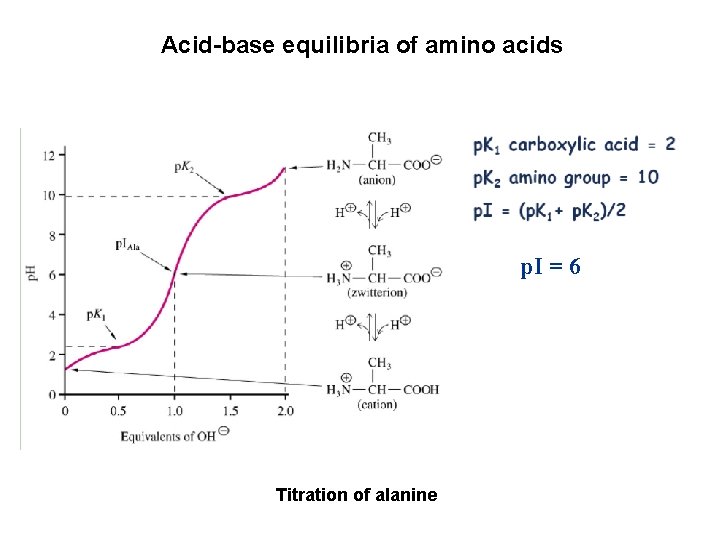
Acid-base equilibria of amino acids p. I = 6 Titration of alanine
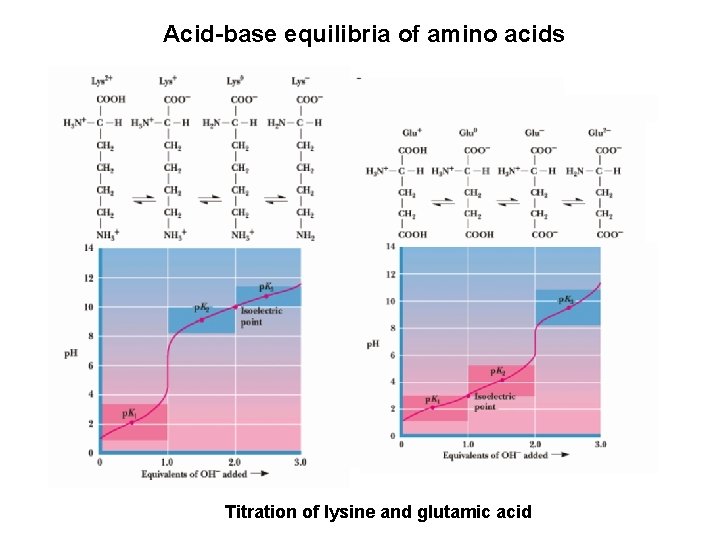
Acid-base equilibria of amino acids Titration of lysine and glutamic acid
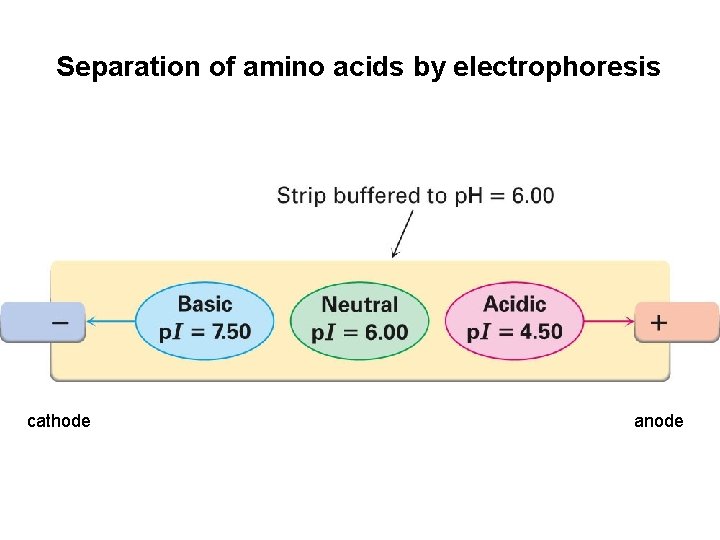
Separation of amino acids by electrophoresis cathode anode
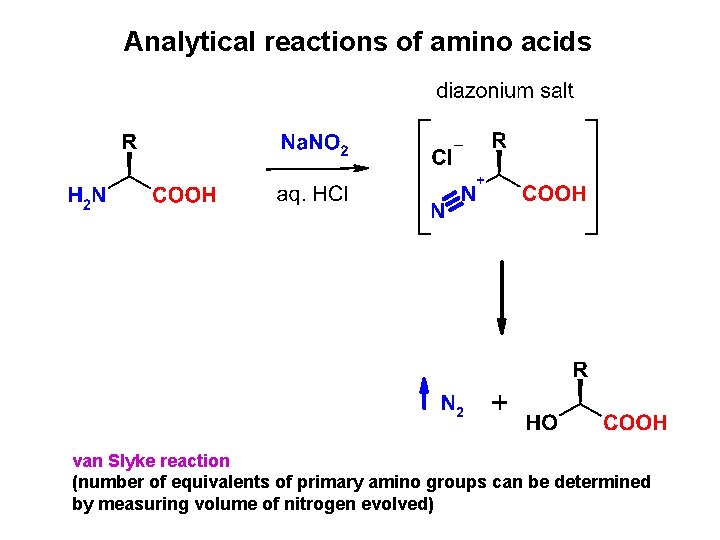
Analytical reactions of amino acids van Slyke reaction (number of equivalents of primary amino groups can be determined by measuring volume of nitrogen evolved)
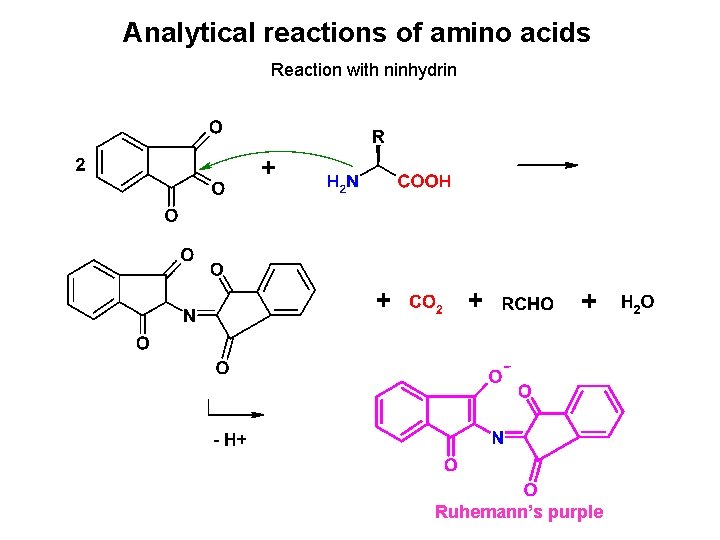
Analytical reactions of amino acids Reaction with ninhydrin Ruhemann’s purple
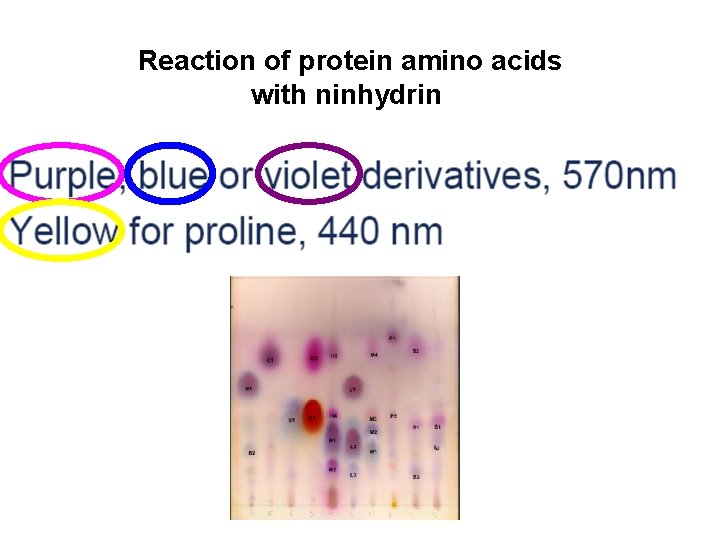
Reaction of protein amino acids with ninhydrin
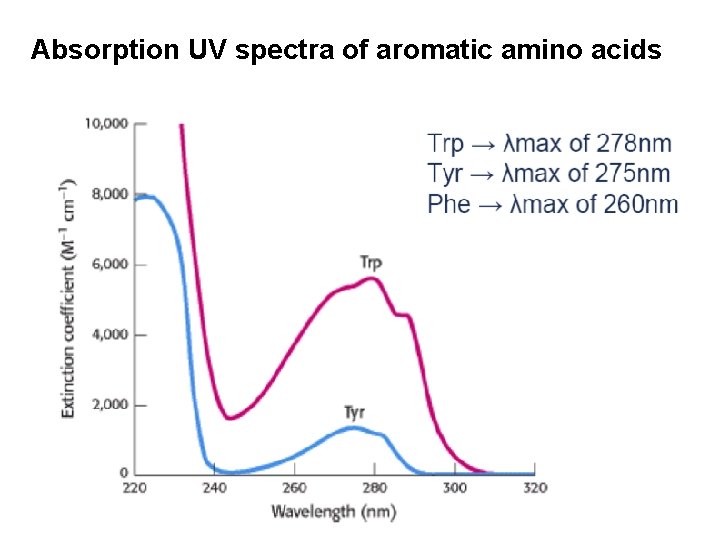
Absorption UV spectra of aromatic amino acids
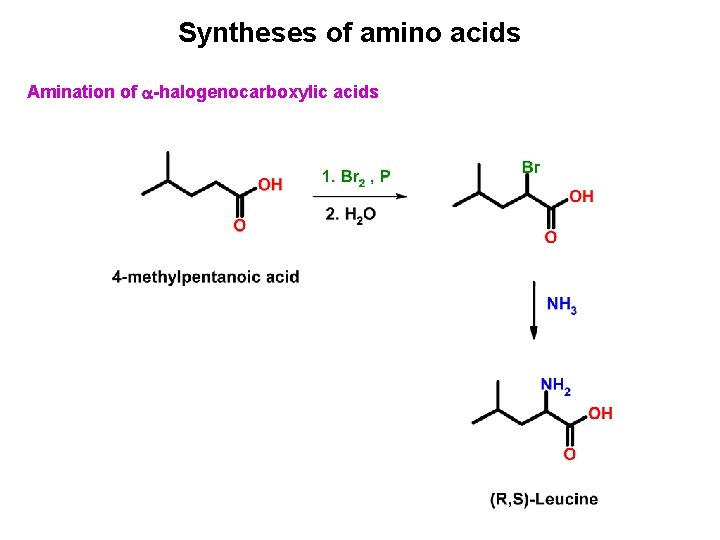
Syntheses of amino acids Amination of -halogenocarboxylic acids
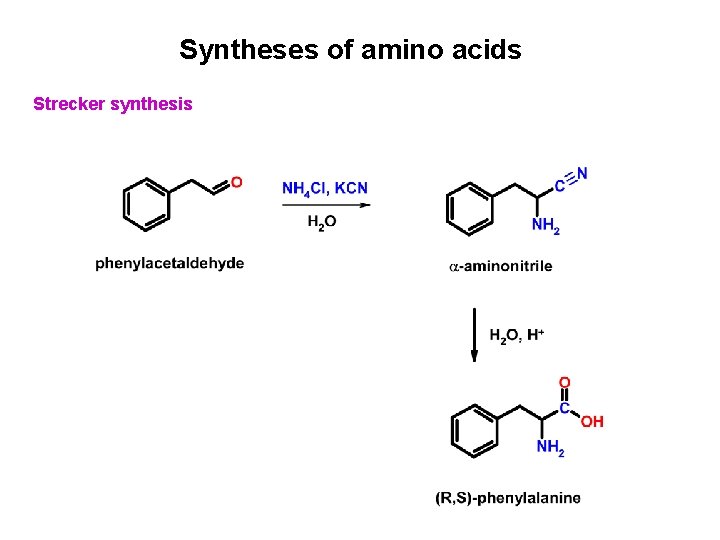
Syntheses of amino acids Strecker synthesis

Syntheses of amino acids Gabriel synthesis
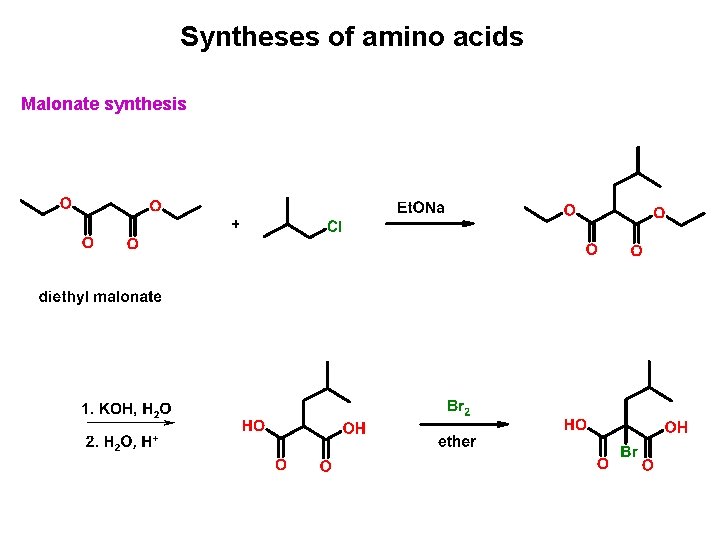
Syntheses of amino acids Malonate synthesis
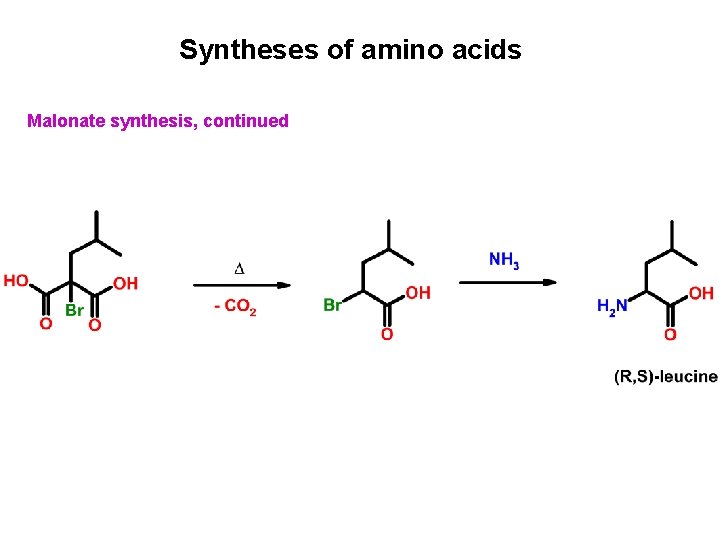
Syntheses of amino acids Malonate synthesis, continued
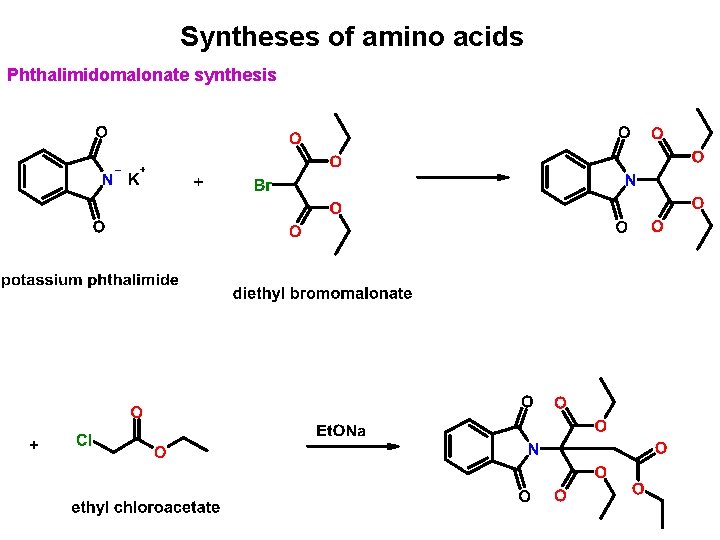
Syntheses of amino acids Phthalimidomalonate synthesis
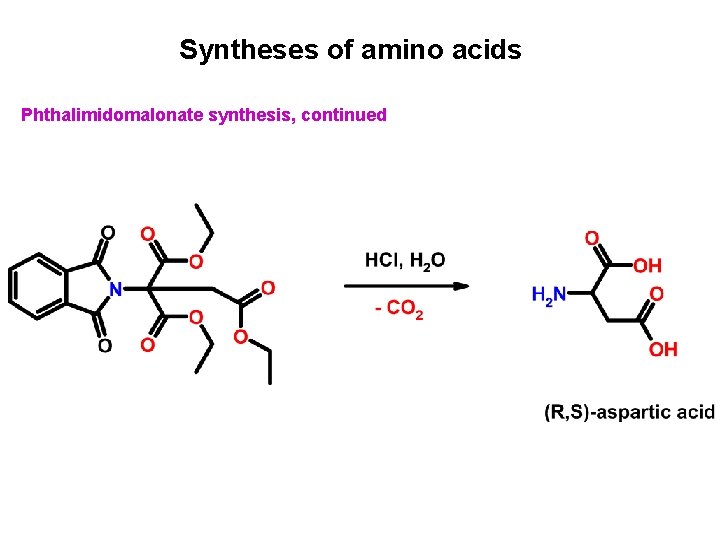
Syntheses of amino acids Phthalimidomalonate synthesis, continued
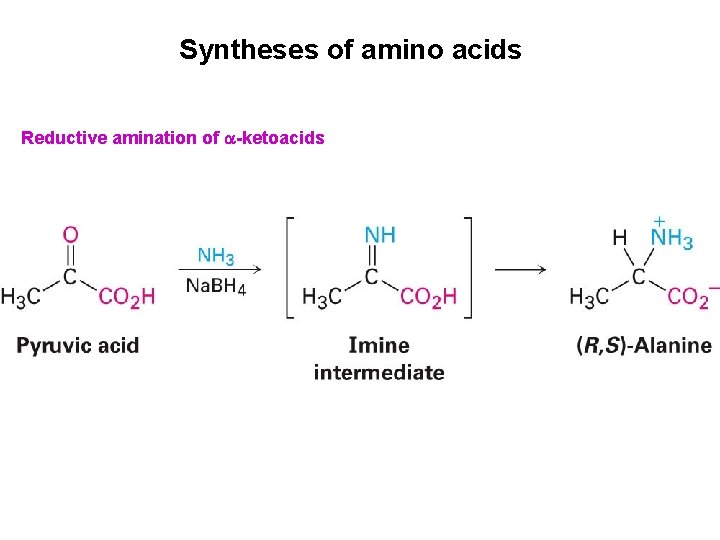
Syntheses of amino acids Reductive amination of -ketoacids
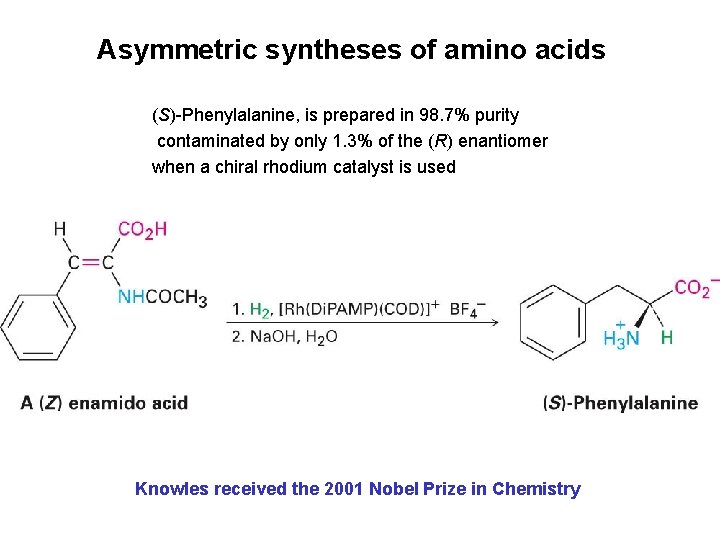
Asymmetric syntheses of amino acids (S)-Phenylalanine, is prepared in 98. 7% purity contaminated by only 1. 3% of the (R) enantiomer when a chiral rhodium catalyst is used Knowles received the 2001 Nobel Prize in Chemistry
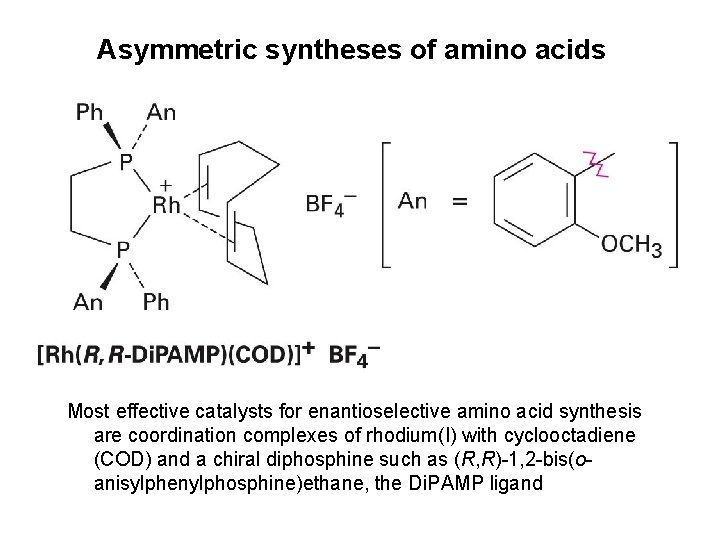
Asymmetric syntheses of amino acids Most effective catalysts for enantioselective amino acid synthesis are coordination complexes of rhodium(I) with cyclooctadiene (COD) and a chiral diphosphine such as (R, R)-1, 2 -bis(oanisylphenylphosphine)ethane, the Di. PAMP ligand
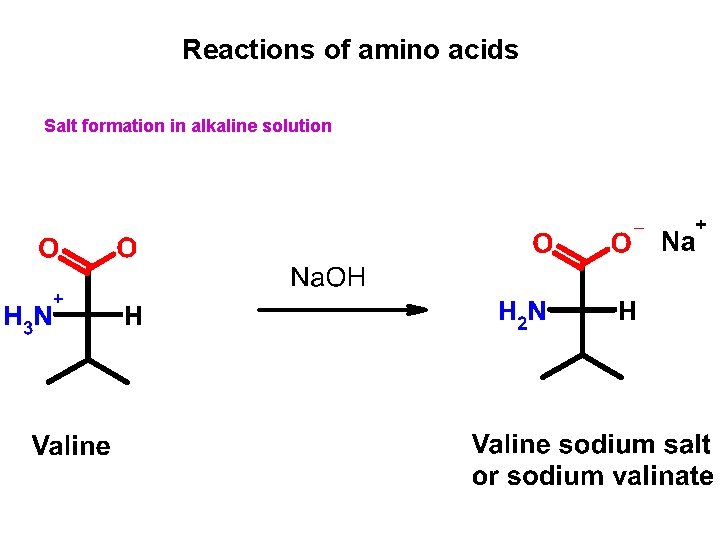
Reactions of amino acids Salt formation in alkaline solution
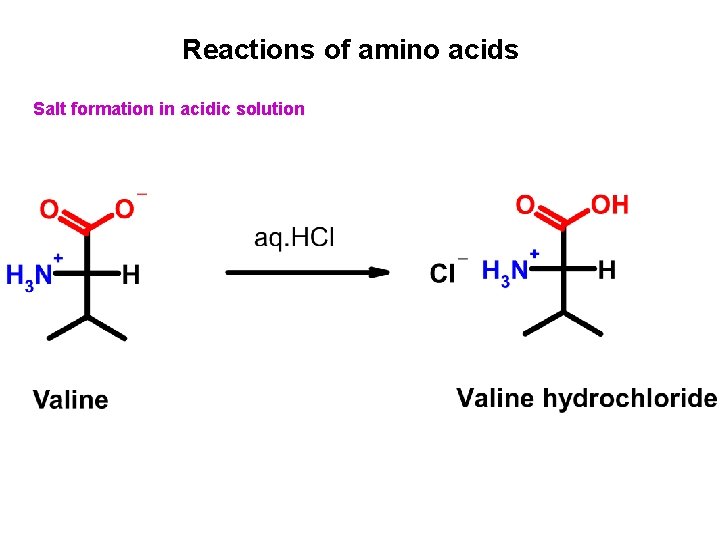
Reactions of amino acids Salt formation in acidic solution
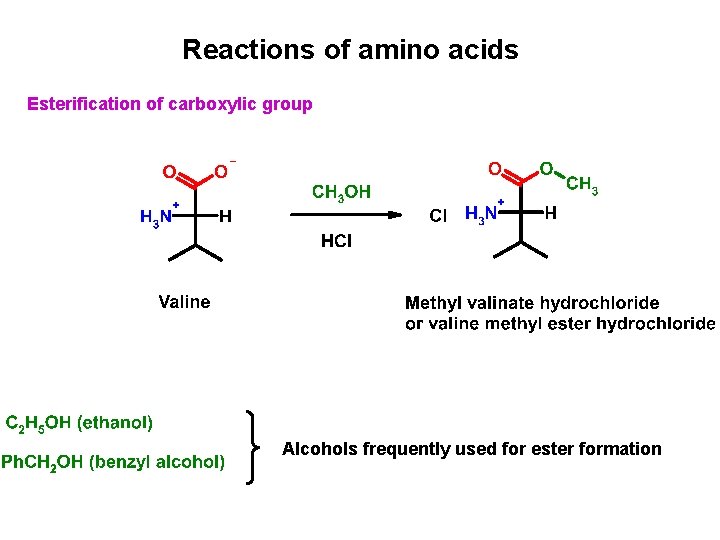
Reactions of amino acids Esterification of carboxylic group Alcohols frequently used for ester formation
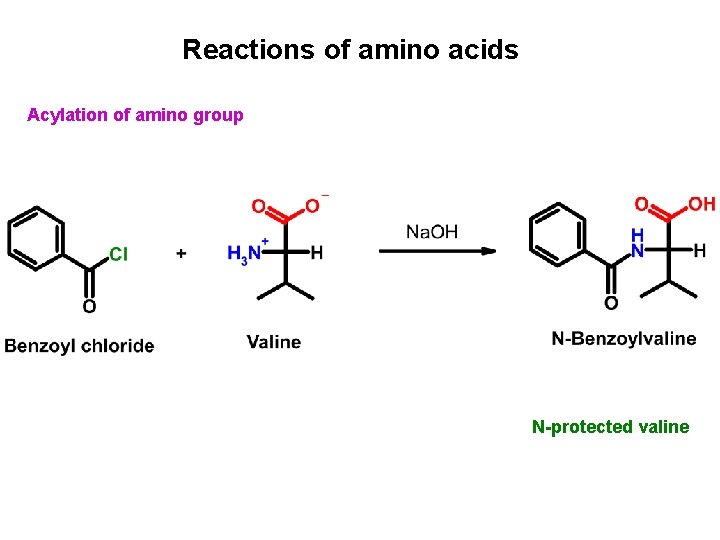
Reactions of amino acids Acylation of amino group N-protected valine
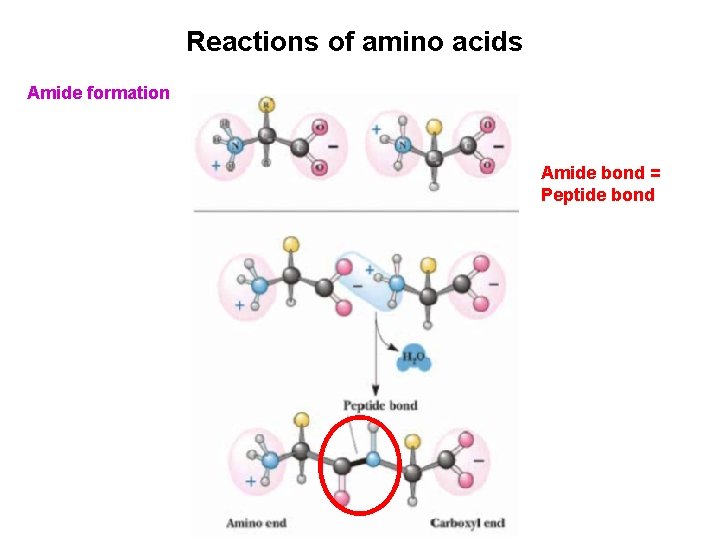
Reactions of amino acids Amide formation Amide bond = Peptide bond

PEPTIDES
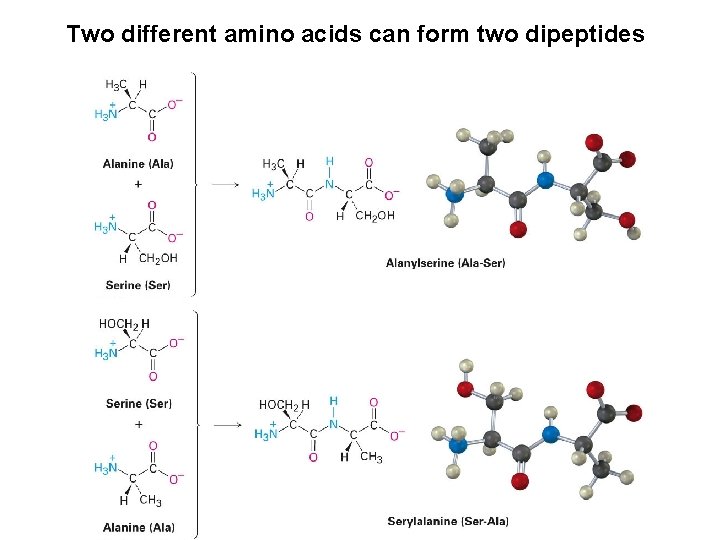
Two different amino acids can form two dipeptides

Convention for writing peptides: • N-terminal amino acid on the left Amino acid with the free –NH 2 group • C-terminal amino acid on the right Amino acid with the free –CO 2 H group The name of the peptide is indicated using abbreviations: Alanylserine is abbreviated Ala-Ser or A-S Serylalanine is abbreviated Ser-Ala or S-A

Three different amino acids can form six tripeptides Example: Asp, Gly, Ser as components of tripeptide can be linked in six ways yielding molecules of different properties Asp-Gly-Ser Asp-Ser-Gly Gly-Asp-Ser Gly-Ser-Asp-Gly Ser-Gly-Asp
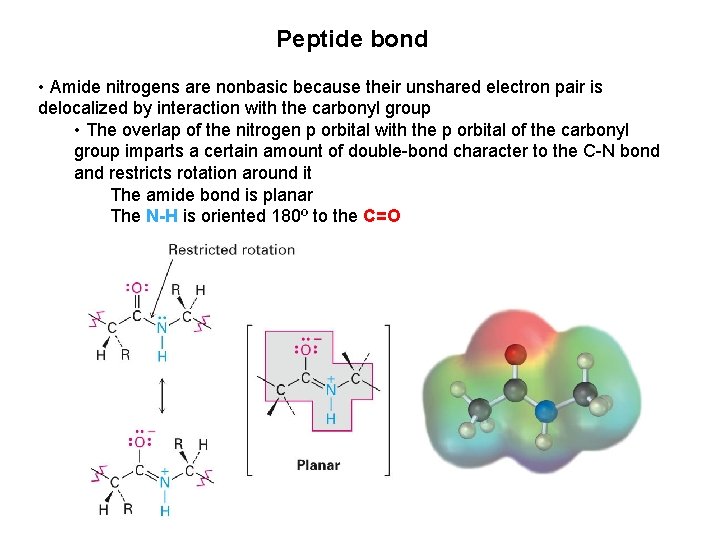
Peptide bond • Amide nitrogens are nonbasic because their unshared electron pair is delocalized by interaction with the carbonyl group • The overlap of the nitrogen p orbital with the p orbital of the carbonyl group imparts a certain amount of double-bond character to the C-N bond and restricts rotation around it The amide bond is planar The N-H is oriented 180º to the C=O
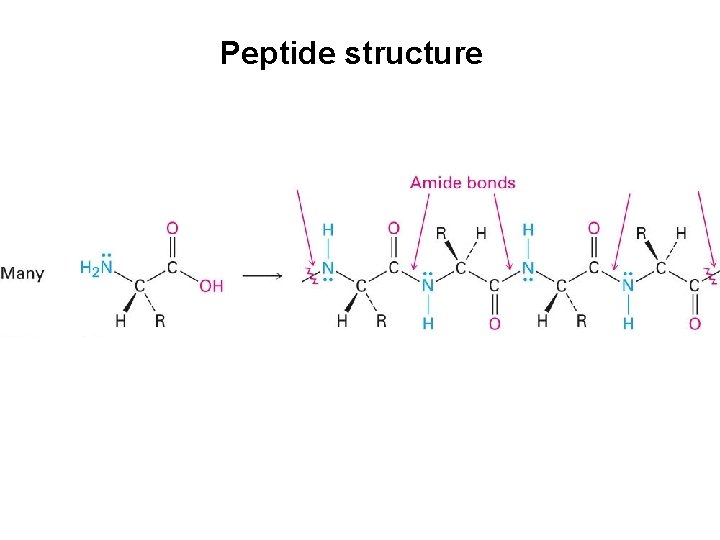
Peptide structure
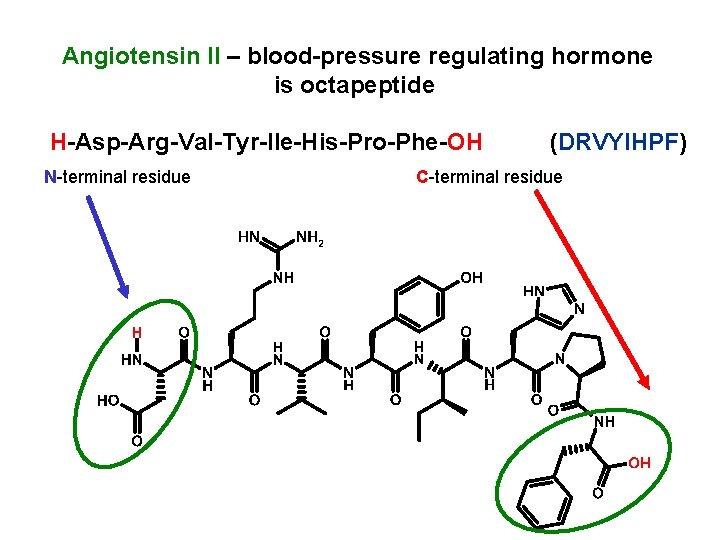
Angiotensin II – blood-pressure regulating hormone is octapeptide H-Asp-Arg-Val-Tyr-Ile-His-Pro-Phe-OH N-terminal residue (DRVYIHPF) C-terminal residue
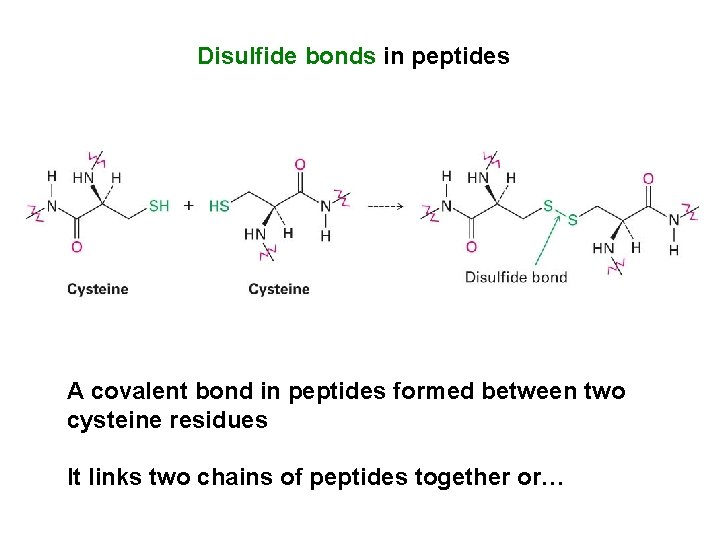
Disulfide bonds in peptides A covalent bond in peptides formed between two cysteine residues It links two chains of peptides together or…
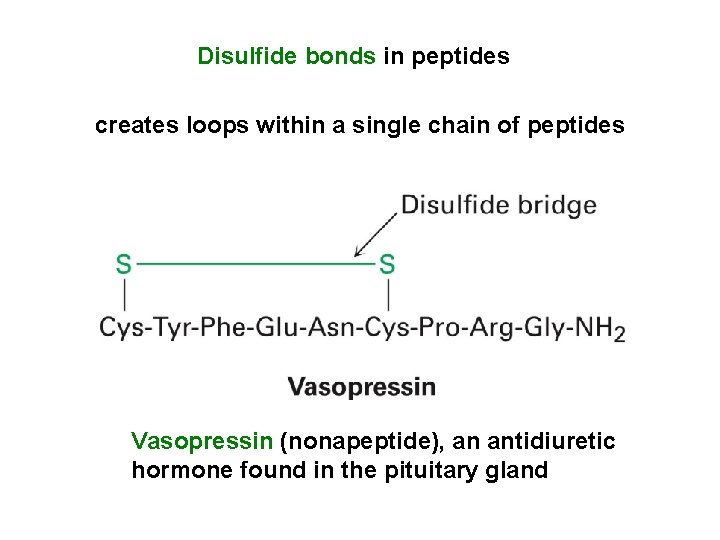
Disulfide bonds in peptides creates loops within a single chain of peptides Vasopressin (nonapeptide), an antidiuretic hormone found in the pituitary gland

Determination of a peptide structure 1. What amino acids are present? 2. How much of each is present? 3. In what sequence do amino acids occur?
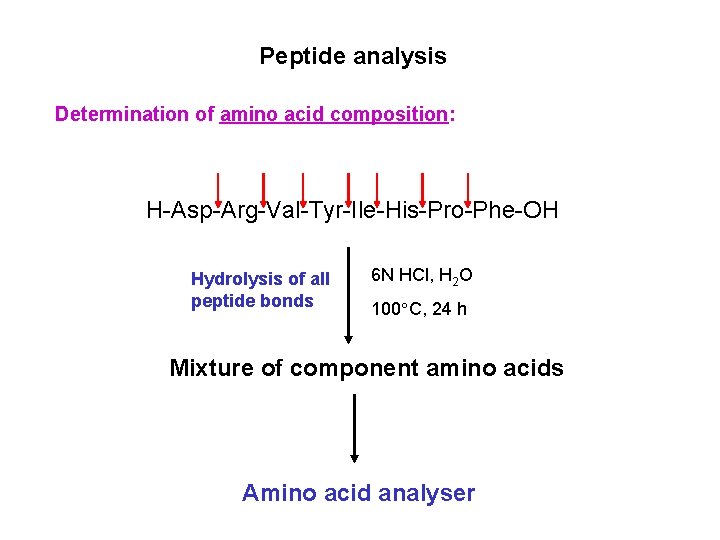
Peptide analysis Determination of amino acid composition: H-Asp-Arg-Val-Tyr-Ile-His-Pro-Phe-OH Hydrolysis of all peptide bonds 6 N HCl, H 2 O 100 C, 24 h Mixture of component amino acids Amino acid analyser

Amino acid analyser
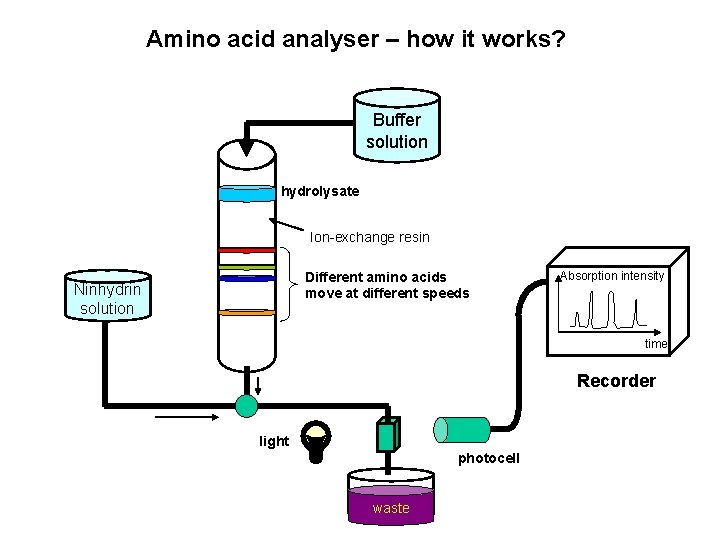
Amino acid analyser – how it works? Buffer solution hydrolysate Ion-exchange resin Different amino acids move at different speeds Ninhydrin solution Absorption intensity time Recorder light photocell waste
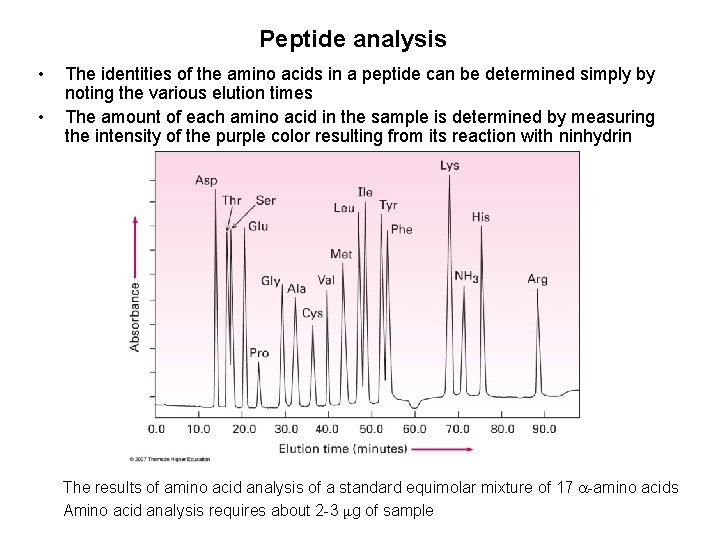
Peptide analysis • • The identities of the amino acids in a peptide can be determined simply by noting the various elution times The amount of each amino acid in the sample is determined by measuring the intensity of the purple color resulting from its reaction with ninhydrin The results of amino acid analysis of a standard equimolar mixture of 17 -amino acids Amino acid analysis requires about 2 -3 μg of sample

Peptide sequencing With the known identities and proportions of amino acids, the peptide is sequenced to find out in what order the amino acids are linked together • Mass spectrometry (molecular weight measurement) – Edman degradation • Cleave one amino acid at a time from an end of the peptide chain • Cleaved amino acid is separated and identified • Repeated until entire peptide sequence is known

Automated protein sequencers • Allow as many as 50 repetitive sequencing cycles to be carried out before a buildup of unwanted by-products interferes with the results • Provide information from as little as 1 to 5 picomoles of sample – less than 0. 1 mg Edman degradation for N-terminal analysis – Involves treatment of a peptide with phenyl isothiocyanate (PITC), C 6 H 5–N=C=S, followed by treatment with trifluoroacetic acid – The phenylthiohydantoin (PTH) is identified chromatographically by comparison of its elution time with the known elution times of PTH derivatives of all 20 protein amino acids
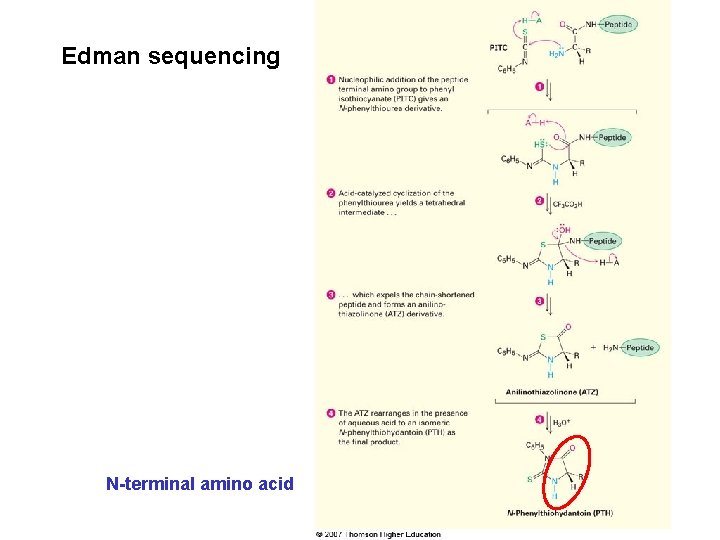
Edman sequencing N-terminal amino acid
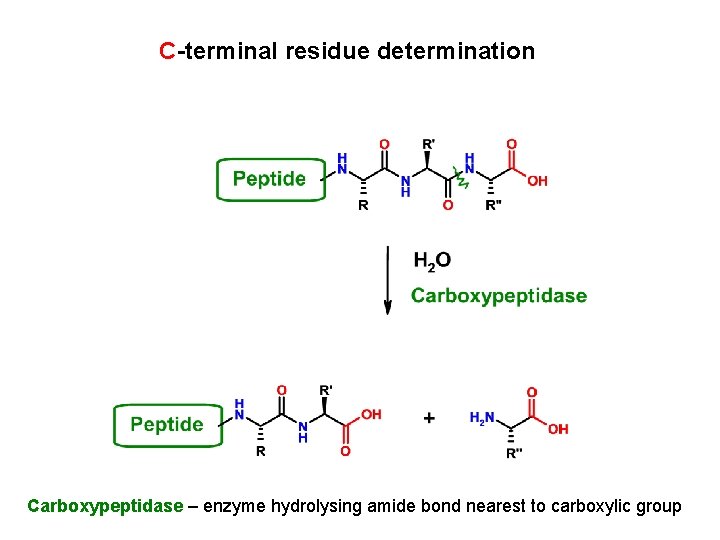
C-terminal residue determination Carboxypeptidase – enzyme hydrolysing amide bond nearest to carboxylic group

Large peptide sequencing 1. Amino acid composition and molecular weight determination 2. Partial hydrolysis (acid or enzyme) and separation of shorter fragments 3. Edman sequencing of shorter fragments 4. Matching smaller fragments to establish full sequence
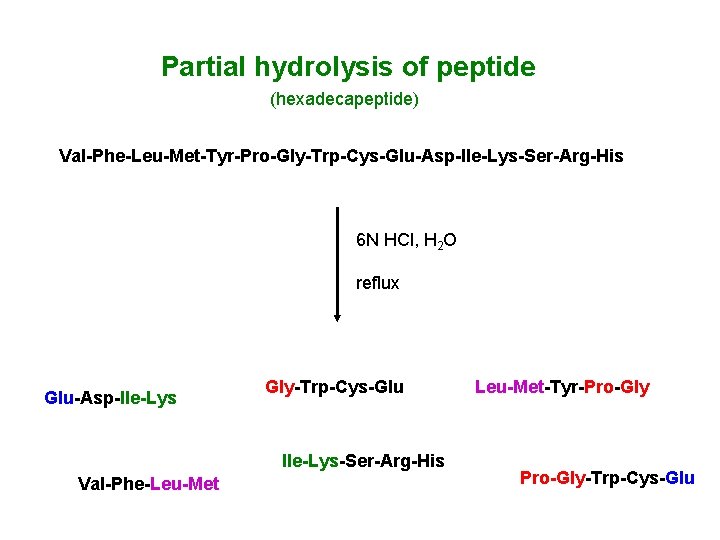
Partial hydrolysis of peptide (hexadecapeptide) Val-Phe-Leu-Met-Tyr-Pro-Gly-Trp-Cys-Glu-Asp-Ile-Lys-Ser-Arg-His 6 N HCl, H 2 O reflux Glu-Asp-Ile-Lys Gly-Trp-Cys-Glu Ile-Lys-Ser-Arg-His Val-Phe-Leu-Met-Tyr-Pro-Gly-Trp-Cys-Glu
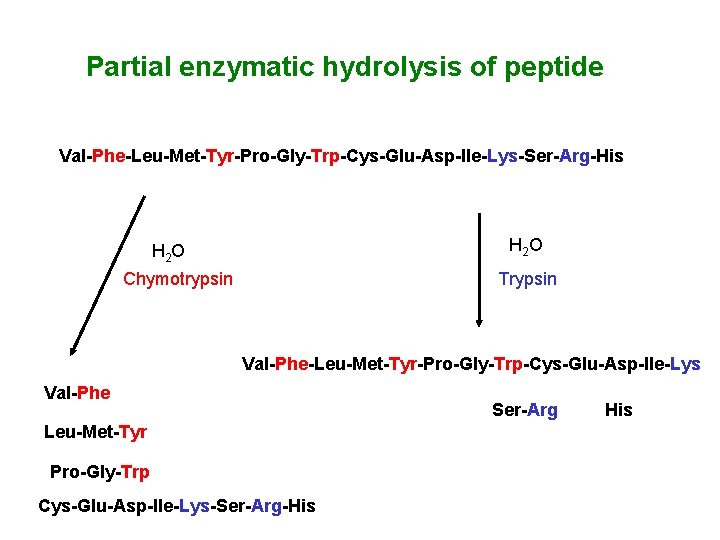
Partial enzymatic hydrolysis of peptide Val-Phe-Leu-Met-Tyr-Pro-Gly-Trp-Cys-Glu-Asp-Ile-Lys-Ser-Arg-His H 2 O Chymotrypsin Trypsin Val-Phe-Leu-Met-Tyr-Pro-Gly-Trp-Cys-Glu-Asp-Ile-Lys Val-Phe Leu-Met-Tyr Pro-Gly-Trp Cys-Glu-Asp-Ile-Lys-Ser-Arg-His Ser-Arg His
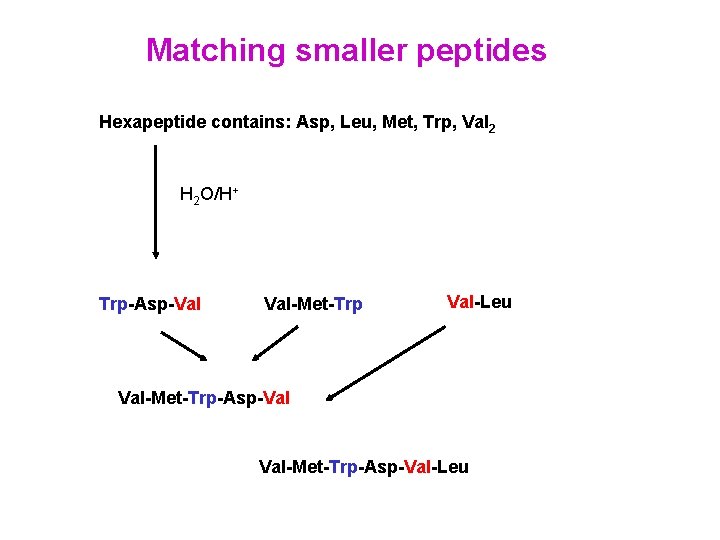
Matching smaller peptides Hexapeptide contains: Asp, Leu, Met, Trp, Val 2 H 2 O/H+ Trp-Asp-Val Val-Met-Trp Val-Leu Val-Met-Trp-Asp-Val-Leu
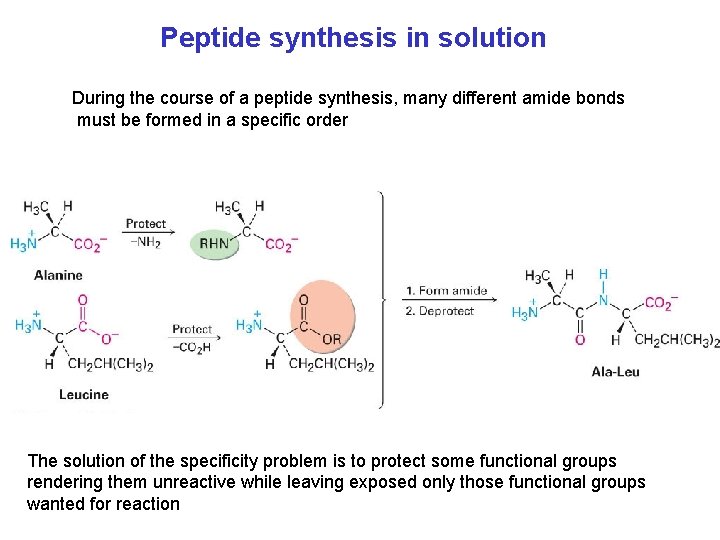
Peptide synthesis in solution During the course of a peptide synthesis, many different amide bonds must be formed in a specific order The solution of the specificity problem is to protect some functional groups rendering them unreactive while leaving exposed only those functional groups wanted for reaction
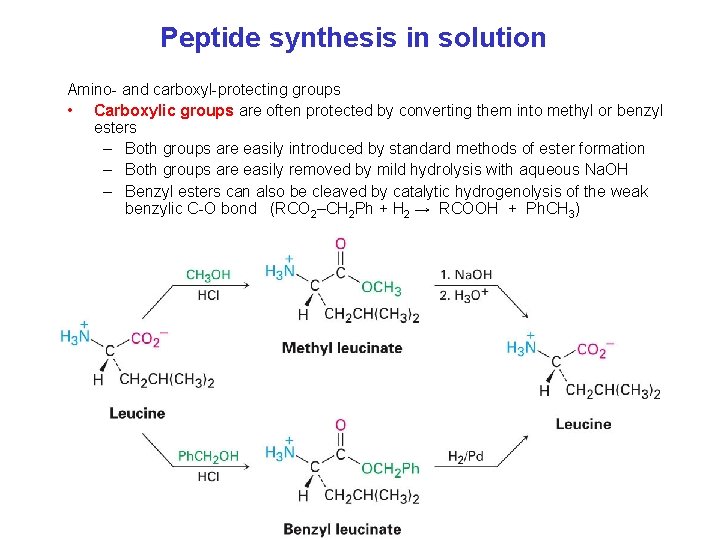
Peptide synthesis in solution Amino- and carboxyl-protecting groups • Carboxylic groups are often protected by converting them into methyl or benzyl esters – Both groups are easily introduced by standard methods of ester formation – Both groups are easily removed by mild hydrolysis with aqueous Na. OH – Benzyl esters can also be cleaved by catalytic hydrogenolysis of the weak benzylic C-O bond (RCO 2–CH 2 Ph + H 2 → RCOOH + Ph. CH 3)
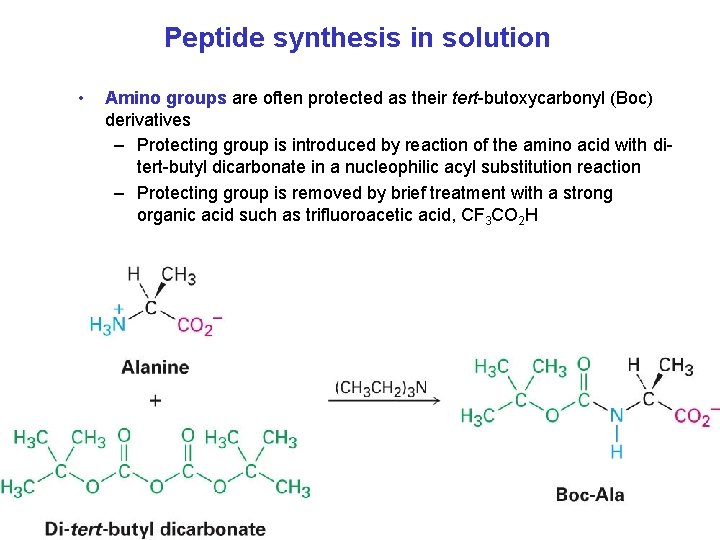
Peptide synthesis in solution • Amino groups are often protected as their tert-butoxycarbonyl (Boc) derivatives – Protecting group is introduced by reaction of the amino acid with ditert-butyl dicarbonate in a nucleophilic acyl substitution reaction – Protecting group is removed by brief treatment with a strong organic acid such as trifluoroacetic acid, CF 3 CO 2 H
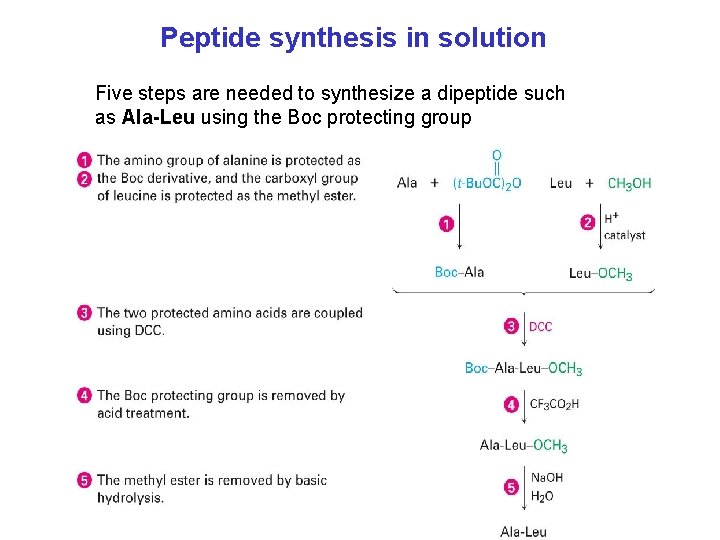
Peptide synthesis in solution Five steps are needed to synthesize a dipeptide such as Ala-Leu using the Boc protecting group
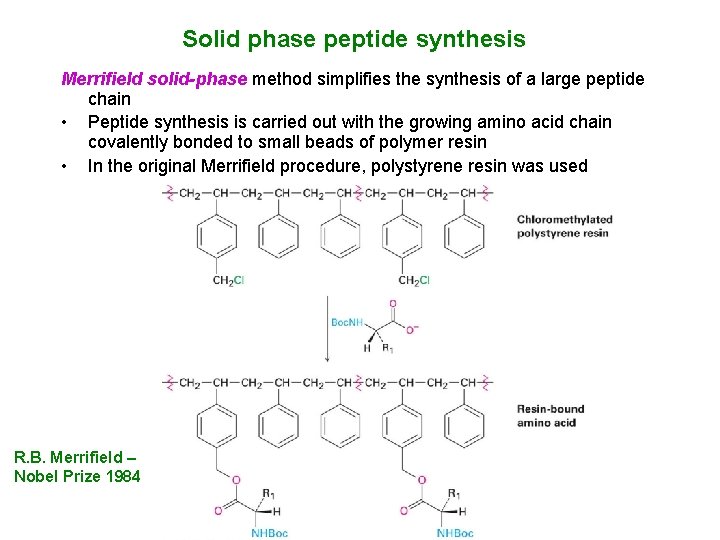
Solid phase peptide synthesis Merrifield solid-phase method simplifies the synthesis of a large peptide chain • Peptide synthesis is carried out with the growing amino acid chain covalently bonded to small beads of polymer resin • In the original Merrifield procedure, polystyrene resin was used R. B. Merrifield – Nobel Prize 1984
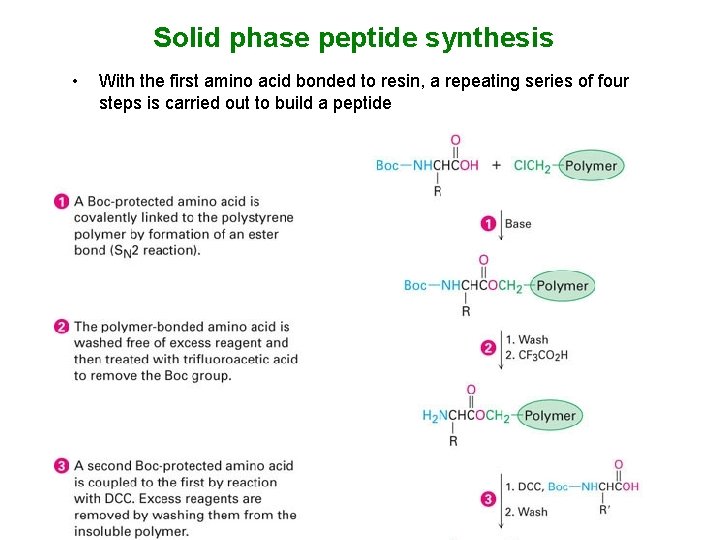
Solid phase peptide synthesis • With the first amino acid bonded to resin, a repeating series of four steps is carried out to build a peptide
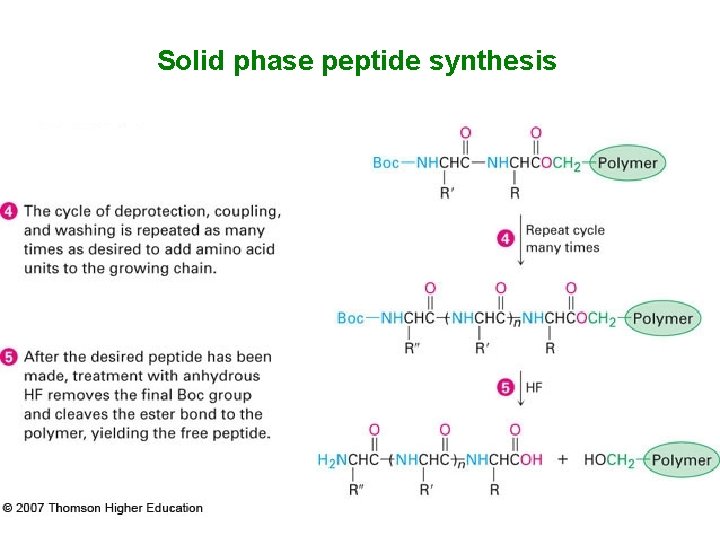
Solid phase peptide synthesis
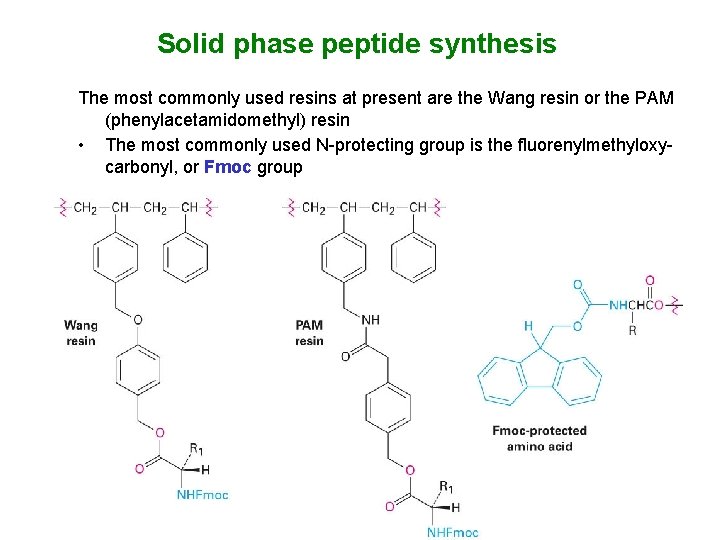
Solid phase peptide synthesis The most commonly used resins at present are the Wang resin or the PAM (phenylacetamidomethyl) resin • The most commonly used N-protecting group is the fluorenylmethyloxycarbonyl, or Fmoc group

Peptide synthesis Robotic, computer-controlled peptide synthesis used to automatically repeat the coupling, washing, and deprotection steps with different amino acids • Each step occurs in high yield • The peptide intermediates are never removed from the insoluble polymer until the final step • Using this procedure, up to 30 mg of a peptide with 20 amino acids can be routinely prepared

PROTEINS
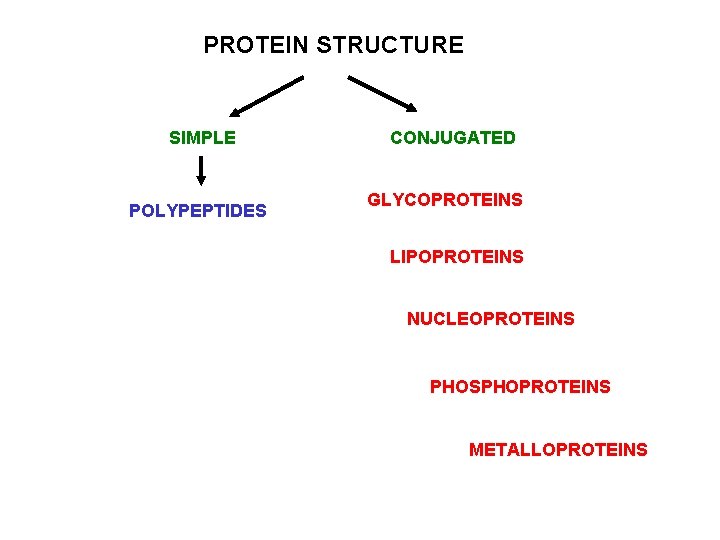
PROTEIN STRUCTURE SIMPLE POLYPEPTIDES CONJUGATED GLYCOPROTEINS LIPOPROTEINS NUCLEOPROTEINS PHOSPHOPROTEINS METALLOPROTEINS
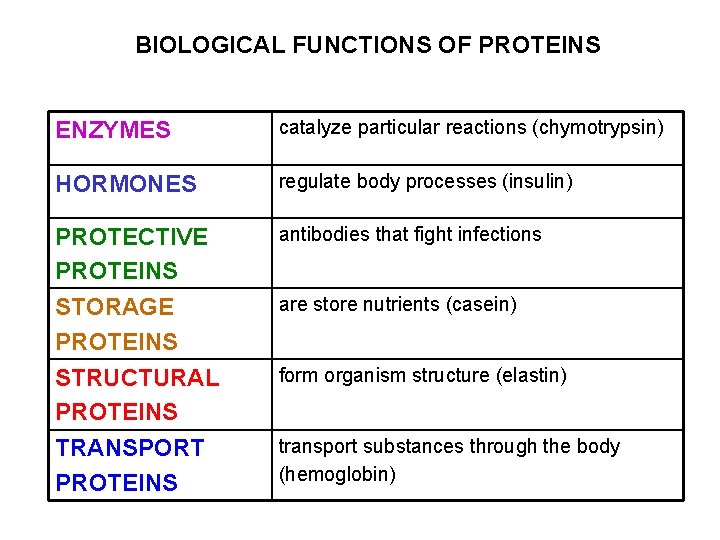
BIOLOGICAL FUNCTIONS OF PROTEINS ENZYMES catalyze particular reactions (chymotrypsin) HORMONES regulate body processes (insulin) PROTECTIVE PROTEINS antibodies that fight infections STORAGE PROTEINS STRUCTURAL PROTEINS TRANSPORT PROTEINS are store nutrients (casein) form organism structure (elastin) transport substances through the body (hemoglobin)

PROTEIN STRUCTURE Proteins are classified according to their threedimensional shape • Fibrous proteins – Consist of polypeptide chains arranged side by side in long filaments – Tough and insoluble in water – Are used in nature as structural materials – – Collagen, elastin in tendons and connective tissue Myosin in muscle tissue -Keratin in nails and hair Silk protein

PROTEIN STRUCTURE • Globular proteins – Usually coiled into compact, roughly spherical shapes – Generally soluble in water and are mobile within cells – Most of the 3000 or so enzymes are globular proteins – Insulin, hormone regulating glucose level in blood – Hemoglobin, protein transporting oxygen

PROTEIN STRUCTURE Four levels for the structure of proteins: • • The primary structure of a protein is simply the amino acid sequence The secondary structure of a protein describes how segments of the peptide backbone orient into a regular pattern The tertiary structure describes how the entire protein molecule coils into an overall three-dimensional shape The quaternary structure describes how several protein molecules come together to yield large aggregate structures Primary structure is determined by sequencing the protein Secondary, tertiary, and quaternary structures are determined by X -ray crystallography Computer aided modelling is also helpful in determining protein structure
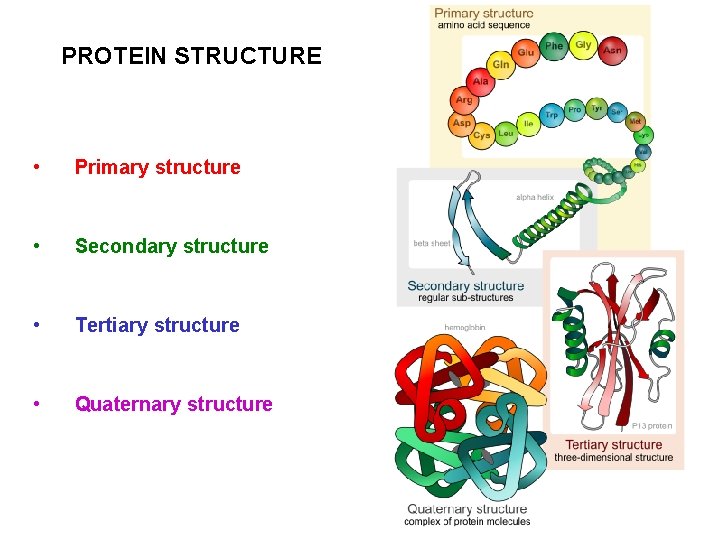
PROTEIN STRUCTURE • Primary structure • Secondary structure • Tertiary structure • Quaternary structure
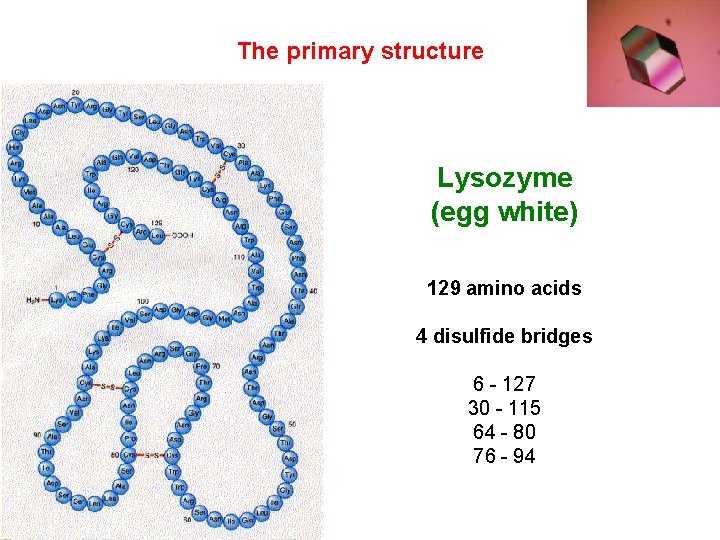
The primary structure Lysozyme (egg white) 129 amino acids 4 disulfide bridges 6 - 127 30 - 115 64 - 80 76 - 94
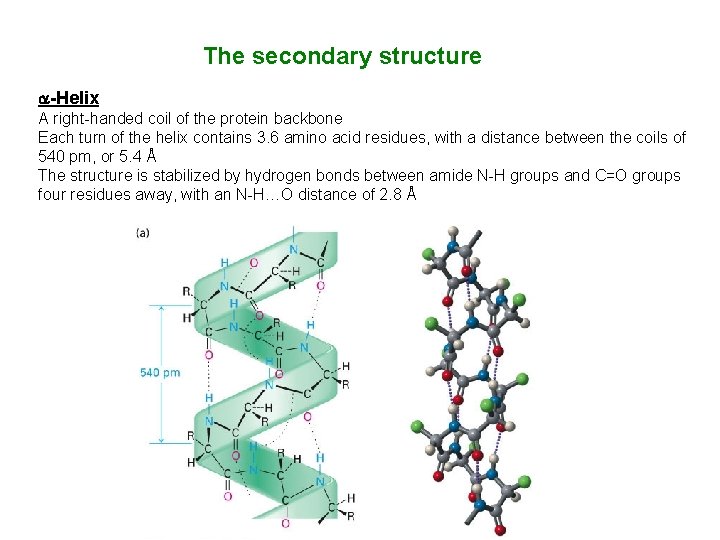
The secondary structure -Helix A right-handed coil of the protein backbone Each turn of the helix contains 3. 6 amino acid residues, with a distance between the coils of 540 pm, or 5. 4 Å The structure is stabilized by hydrogen bonds between amide N-H groups and C=O groups four residues away, with an N-H…O distance of 2. 8 Å
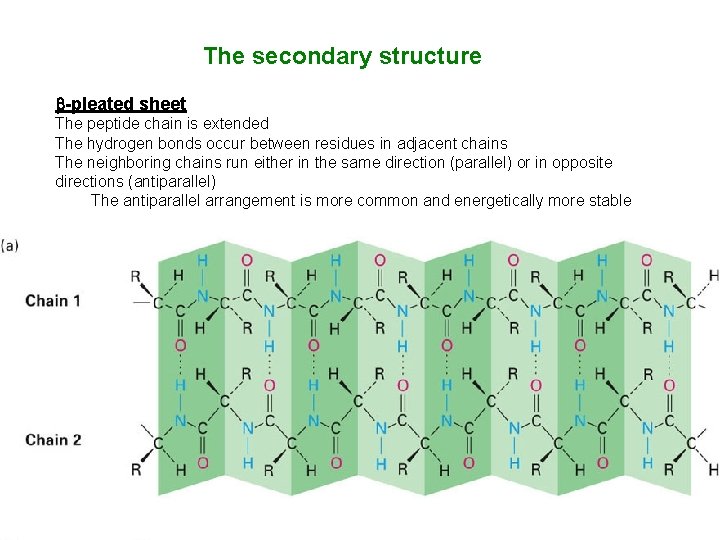
The secondary structure -pleated sheet The peptide chain is extended The hydrogen bonds occur between residues in adjacent chains The neighboring chains run either in the same direction (parallel) or in opposite directions (antiparallel) The antiparallel arrangement is more common and energetically more stable
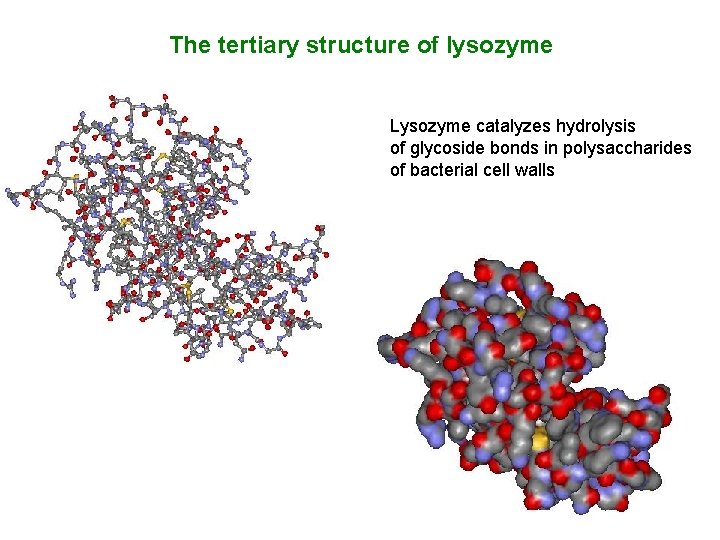
The tertiary structure of lysozyme Lysozyme catalyzes hydrolysis of glycoside bonds in polysaccharides of bacterial cell walls
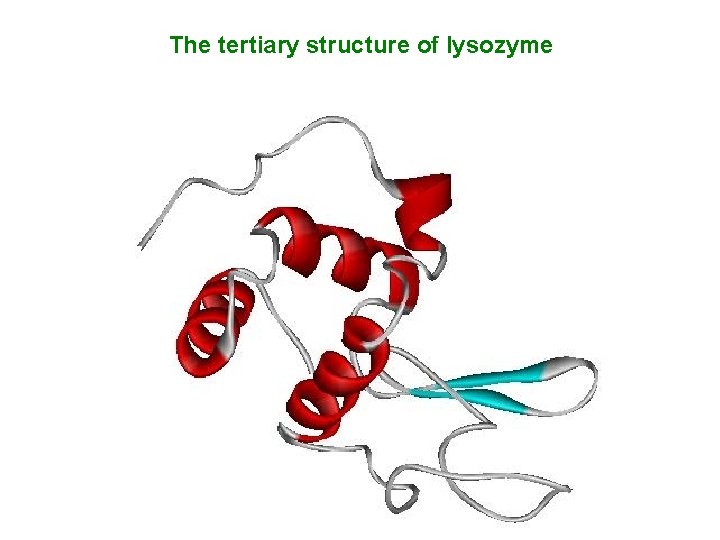
The tertiary structure of lysozyme
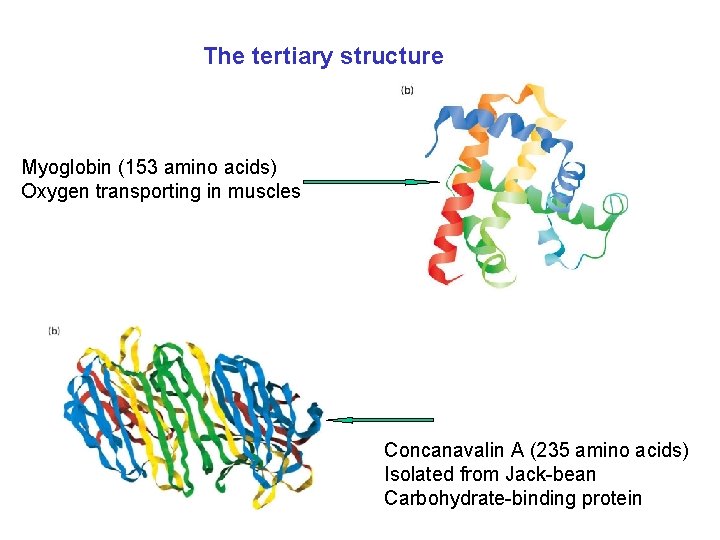
The tertiary structure Myoglobin (153 amino acids) Oxygen transporting in muscles Concanavalin A (235 amino acids) Isolated from Jack-bean Carbohydrate-binding protein
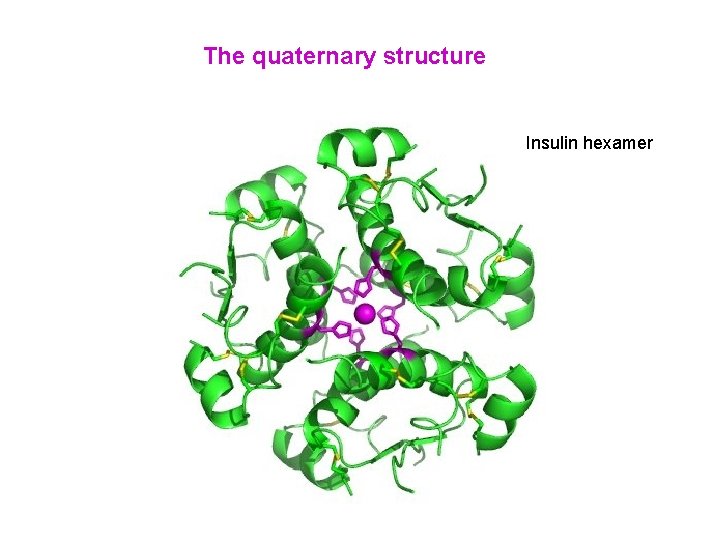
The quaternary structure Insulin hexamer

PROTEIN STRUCTURE stabilization The forces that determine the tertiary structures of a protein are the same forces that act on all molecules, regardless of size, to provide maximum stability • Hydrophilic (water loving) interactions of the polar side chains of acidic or basic amino acids – Acidic or basic amino acids with charged side chains congregate on the exterior of the protein, where they can be solvated by water – Amino acids with neutral, nonpolar side chains congregate on the hydrocarbon-like interior of a protein molecule away from the aqueous medium Other factors for stabilization of a protein’s tertiary structure • Formation of disulfide bridges between cysteine residues • The formation of hydrogen bonds between nearby amino acid residues • The presence of ionic attractions, called salt bridges, between positively and negatively charged sites on various amino acid side chains within the protein
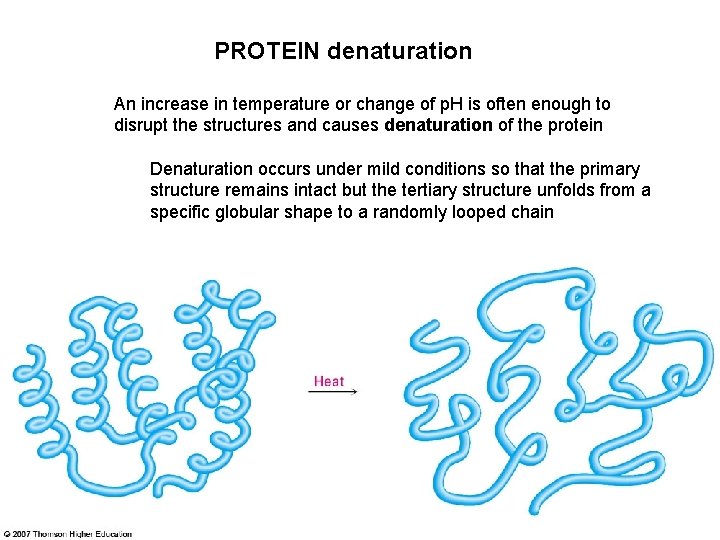
PROTEIN denaturation An increase in temperature or change of p. H is often enough to disrupt the structures and causes denaturation of the protein Denaturation occurs under mild conditions so that the primary structure remains intact but the tertiary structure unfolds from a specific globular shape to a randomly looped chain

PROTEIN denaturation • It is accompanied by changes in both physical and biological properties – Solubility is drastically decreased • Most enzymes lose all catalytic activity • Most denaturation is irreversible – In some cases spontaneous renaturation of an unfolded protein to its stable tertiary structure occurs and is accompanied by a full recovery of biological functions

ENZYMES and COENZYMES Enzyme • • Usually a large protein A substance that acts as a catalyst for a biological reaction Does not affect the equilibrium constant of a reaction Acts only to lower the activation energy for a reaction to make the reaction proceed more rapidly – Glycosidase enzymes that hydrolyze polysaccharides increase the reaction rate by a factor of more than 1017, changing the time required for the reaction from millions of years to milliseconds • Usually specific in their actions – Often, an enzyme will catalyze only a single reaction of a single compound, the enzyme’s substrate
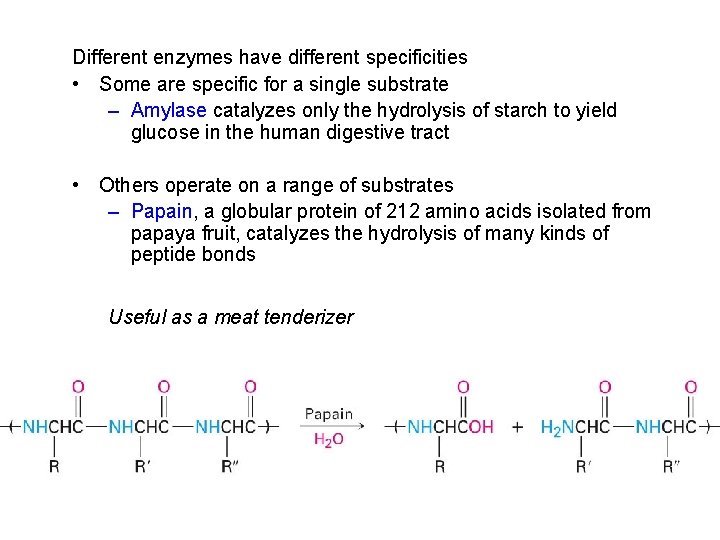
Different enzymes have different specificities • Some are specific for a single substrate – Amylase catalyzes only the hydrolysis of starch to yield glucose in the human digestive tract • Others operate on a range of substrates – Papain, a globular protein of 212 amino acids isolated from papaya fruit, catalyzes the hydrolysis of many kinds of peptide bonds Useful as a meat tenderizer

How do enzymes work? Enzymes function through a pathway that involves initial formation of an enzymesubstrate complex E • S, a multistep chemical conversion of the enzyme-bound substrate into enzyme-bound product E • P, and final release of product from the complex E+S E • P E+P
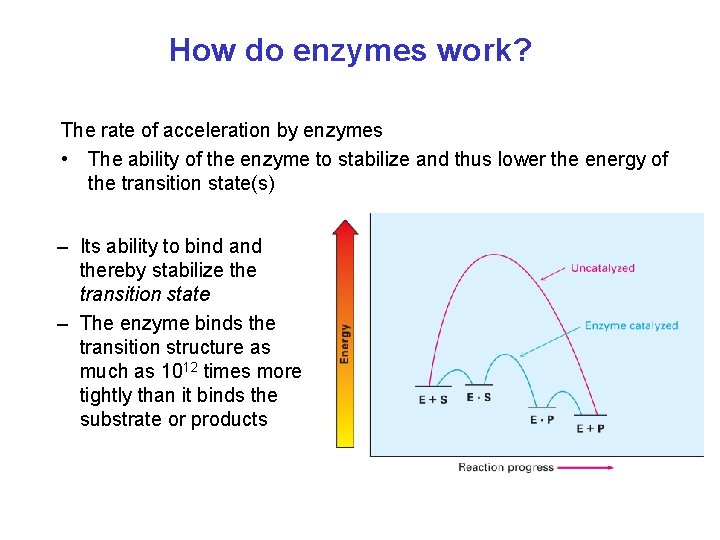
How do enzymes work? The rate of acceleration by enzymes • The ability of the enzyme to stabilize and thus lower the energy of the transition state(s) – Its ability to bind and thereby stabilize the transition state – The enzyme binds the transition structure as much as 1012 times more tightly than it binds the substrate or products

Enzymes are classified into six categories depending on the kind of reaction they catalyze

Most enzymes also contain a small part called the cofactor Cofactor • Can be an inorganic ion, such as Zn 2+ • Can be a small organic molecule called a coenzyme Coenzyme • A reactant that undergoes chemical change during the reaction and requires an additional step to return to its initial state Many coenzymes are derived from vitamins
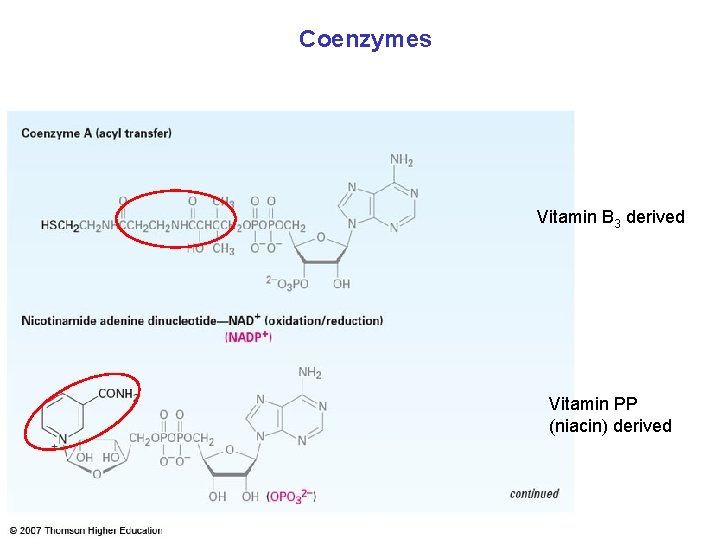
Coenzymes Vitamin B 3 derived Vitamin PP (niacin) derived

How do enzymes work? Enzymes • Bring reactant molecules together • Hold molecules in orientation necessary for reaction • Provide any necessary acidic or basic sites to catalyze specific steps – Citrate synthase • An enzyme that catalyzes the aldol-like addition of acetyl Co. A to oxaloacetate to give citrate • The reaction is the first step in the citric acid cycle, in which acetyl groups produced by degradation of food molecules are metabolized to yield CO 2 and H 2 O
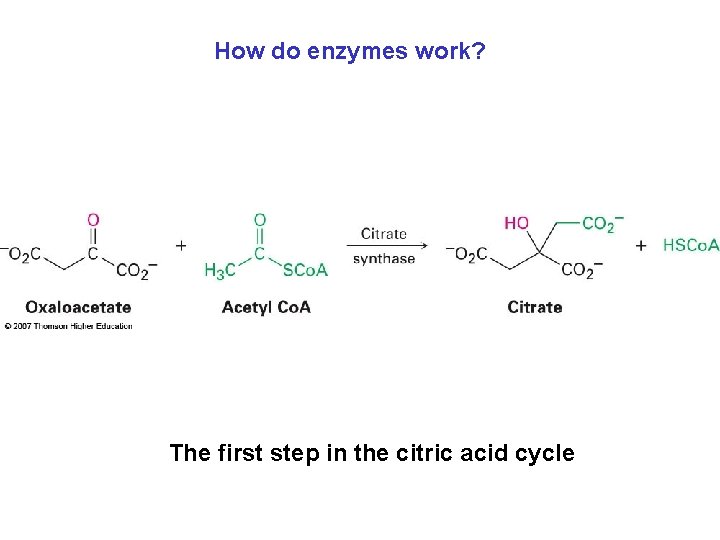
How do enzymes work? The first step in the citric acid cycle
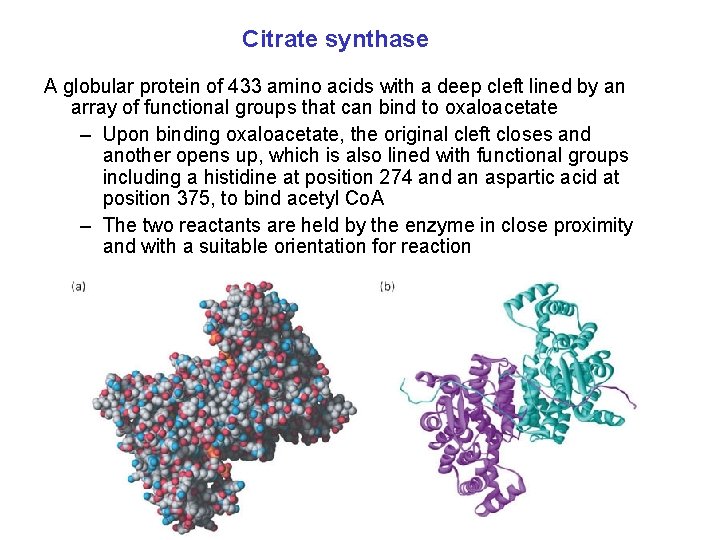
Citrate synthase A globular protein of 433 amino acids with a deep cleft lined by an array of functional groups that can bind to oxaloacetate – Upon binding oxaloacetate, the original cleft closes and another opens up, which is also lined with functional groups including a histidine at position 274 and an aspartic acid at position 375, to bind acetyl Co. A – The two reactants are held by the enzyme in close proximity and with a suitable orientation for reaction
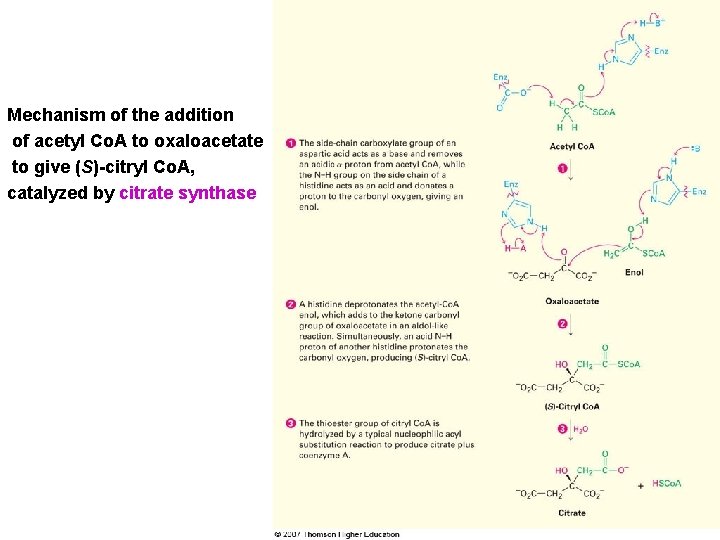
Mechanism of the addition of acetyl Co. A to oxaloacetate to give (S)-citryl Co. A, catalyzed by citrate synthase
- Slides: 97