Amino acid metabolism II nucleic acid chemistry I

Amino acid metabolism II; nucleic acid chemistry I Andy Howard Biochemistry Lectures, Fall 2010 17 November 2010 Biochem: AAmetab 2; Nuc 1 11/17/2010

Amino acid metabolism Amino acid synthetic pathways are complex but we can understand them if we’re patient n Amino acid degradation falls into two broad categories n Nucleic acid chemistry: N-rings n 11/17/2010 Biochem: AAmetab 2; Nuc 1 2

What we’ll cover n Amino acid synthesis n n n n n Reactions Glutamate n Cellular localization Glutamine n Nucleic acid chemistry Other simple aas n Pyrimidines: C, U, T Essential amino acids n Purines: A, G Nucleosides Amino acid catabolism n Urea cycle Degradation products Interconversions Specifics n n Nucleotides Oligo- and polynucleotides Duplex DNA Helicity 11/17/2010 Biochem: AAmetab 2; Nuc 1 3

Glutamate n n Glutamate is a critical metabolite because so many of the transaminations start with it as the amine donor It is produced in E. coli, etc. via glutamate dehydrogenase using ammonium ion as nitrogen donor: -ketoglutarate + NH 4+ + NAD(P)H + H+ NAD(P)+ + H 2 O + glutamate 1 BGV 296 k. Da hexamer monomer shown EC 1. 4. 1. 2, 1. 9Å Clostridium 11/17/2010 Biochem: AAmetab 2; Nuc 1 4

Glutamine n n Glutamate can be aminated with expenditure of ATP to form glutamine: glutamate + NH 4+ + ATP glutamine + ADP + Pi Note that glutamine synthetase is a ligase: the ATP is an energy-provider, not a phosphate donor 11/17/2010 Biochem: AAmetab 2; Nuc 1 Glutamine synthetase PDB 2 OJW 211 k. Da pentamer Human EC 6. 3. 1. 2, 2. 05Å 5

Aspartate and asparagine n n Asp is simple: transamination of oxaloacetate Asn is straightforward too asparagine synthetase moves the amine from gln to asp, leaving glu (another ligase) Gln + asp + ATP AMP + PPi + glu + asn 11/17/2010 Biochem: AAmetab 2; Nuc 1 Asparagine synthetase B PDB 1 CT 9 243 k. Da EC 6. 3. 5. 4, 2Å tetramer E. coli 6

Simple: ala, gly, ser n n n Alanine by transamination from pyruvate Glycine from serine by SHMT (q. v. ) Serine from 3 -phosphoglycerate: n n n Phosphoserine 3 -phosphoglycerate + NAD phosphatase NADH + H+ + 3 PDB 1 NNL phosphohydroxypyruvate 49 k. Da dimer 3 -phosphohydroxypyruvate + glutamate human 3 -phoserine + -ketoglutarate EC 3. 1. 3. 3 3 -phoserine + H 2 O serine + Pi 1. 53Å 11/17/2010 Biochem: AAmetab 2; Nuc 1 7

Serine hydroxymethyltransferase n n n Serine + tetrahydrofolate H 2 O + glycine + 5, 10 -methylenetetrahydrofolate This can be viewed as a source of methylene units for other biosyntheses PLP-dependent reaction 11/17/2010 Biochem: AAmetab 2; Nuc 1 SHMT PDB 2 DKJ 90 k. Da dimer EC 2. 1 1. 15Å Thermus thermophilus 8
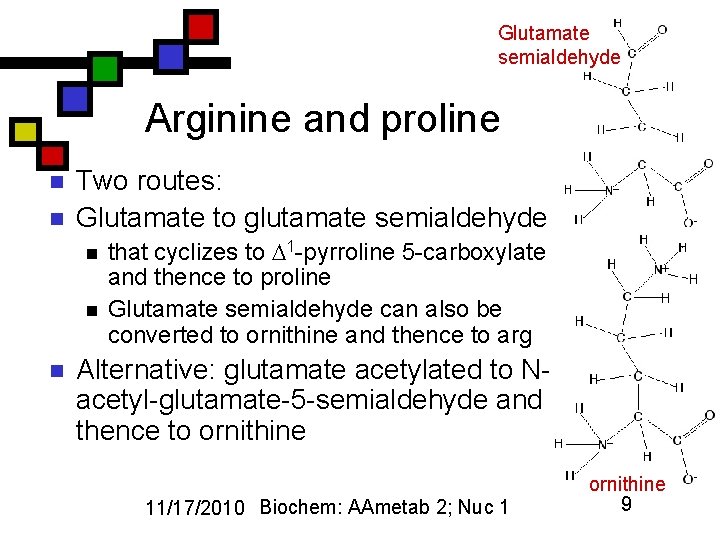
Glutamate semialdehyde Arginine and proline n n Two routes: Glutamate to glutamate semialdehyde n n n that cyclizes to 1 -pyrroline 5 -carboxylate and thence to proline Glutamate semialdehyde can also be converted to ornithine and thence to arg Alternative: glutamate acetylated to Nacetyl-glutamate-5 -semialdehyde and thence to ornithine 11/17/2010 Biochem: AAmetab 2; Nuc 1 ornithine 9

Glutamate to P 5 C n n Single enzyme can interconvert glutamate and 1 -pyrroline carboxylate: 1 -pyrroline-5 -carboxylate dehydrogenase 3 -layer sandwich protein 11/17/2010 Biochem: AAmetab 2; Nuc 1 PDB 2 BJA 300 k. Da hexamer dimer shown EC 1. 5. 1. 12, 1. 9Å Thermus thermophilus 10
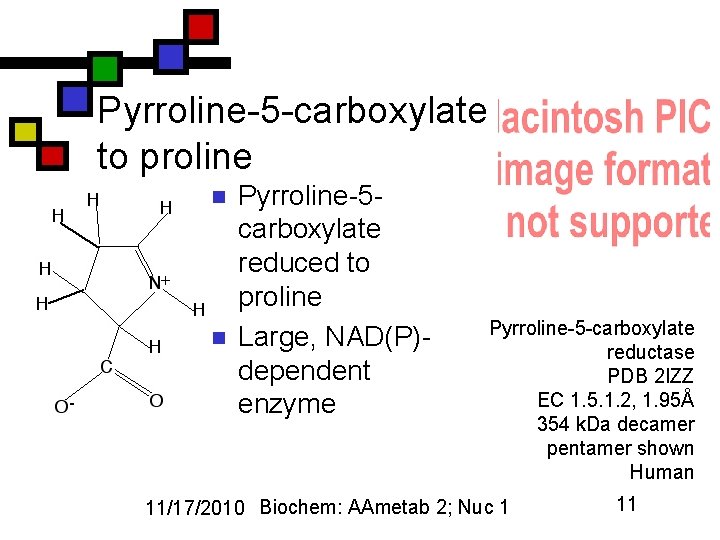
Pyrroline-5 -carboxylate to proline n n Pyrroline-5 carboxylate reduced to proline Large, NAD(P)dependent enzyme Pyrroline-5 -carboxylate reductase PDB 2 IZZ EC 1. 5. 1. 2, 1. 95Å 354 k. Da decamer pentamer shown Human 11/17/2010 Biochem: AAmetab 2; Nuc 1 11
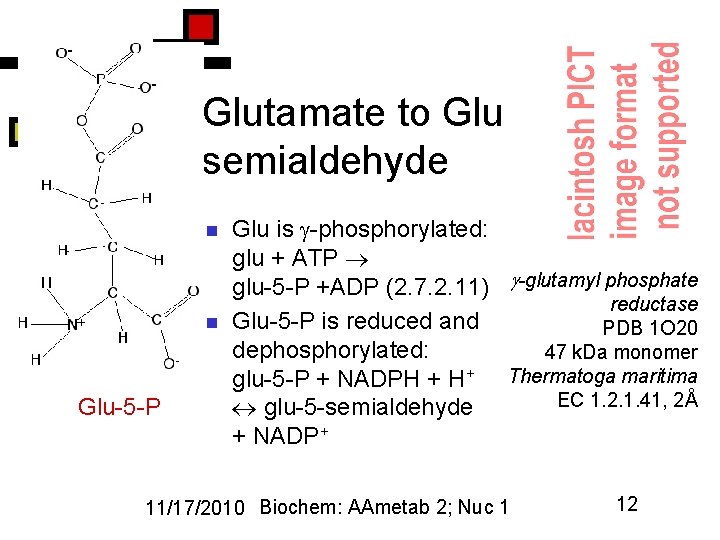
Glutamate to Glu semialdehyde n n Glu-5 -P Glu is -phosphorylated: glu + ATP glu-5 -P +ADP (2. 7. 2. 11) -glutamyl phosphate reductase Glu-5 -P is reduced and PDB 1 O 20 dephosphorylated: 47 k. Da monomer glu-5 -P + NADPH + H+ Thermatoga maritima EC 1. 2. 1. 41, 2Å glu-5 -semialdehyde + NADP+ 11/17/2010 Biochem: AAmetab 2; Nuc 1 12
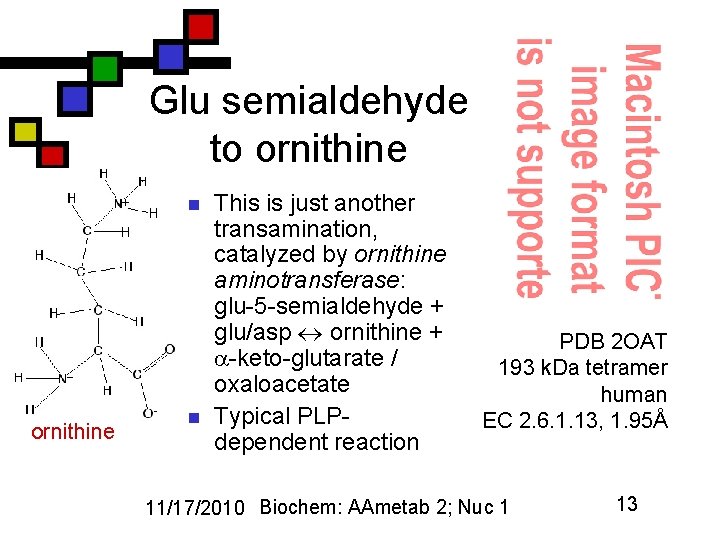
Glu semialdehyde to ornithine n This is just another transamination, catalyzed by ornithine aminotransferase: glu-5 -semialdehyde + glu/asp ornithine + -keto-glutarate / oxaloacetate Typical PLPdependent reaction PDB 2 OAT 193 k. Da tetramer human EC 2. 6. 1. 13, 1. 95Å 11/17/2010 Biochem: AAmetab 2; Nuc 1 13
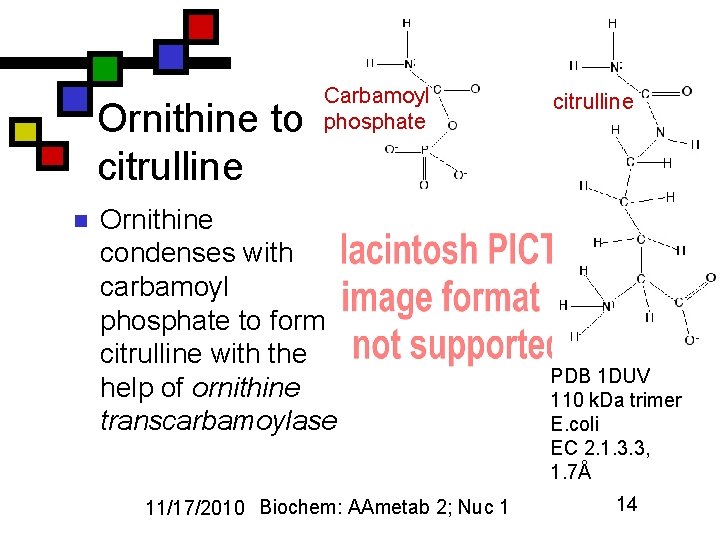
Ornithine to citrulline n Carbamoyl phosphate Ornithine condenses with carbamoyl phosphate to form citrulline with the help of ornithine transcarbamoylase 11/17/2010 Biochem: AAmetab 2; Nuc 1 citrulline PDB 1 DUV 110 k. Da trimer E. coli EC 2. 1. 3. 3, 1. 7Å 14
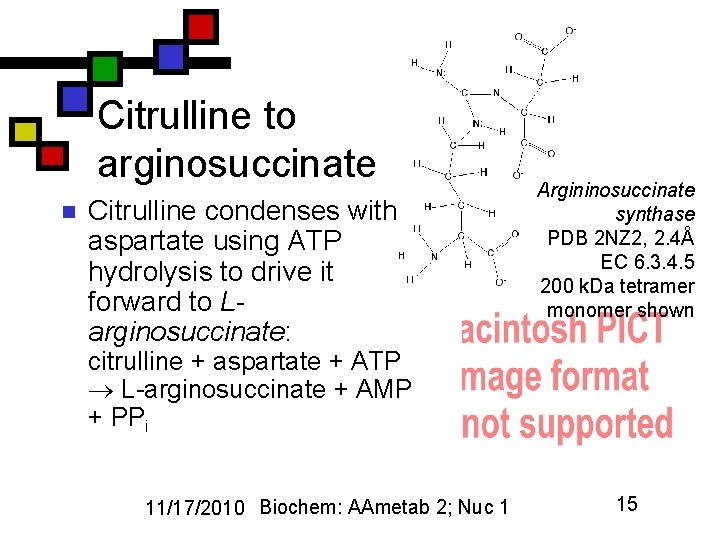
Citrulline to arginosuccinate n Citrulline condenses with aspartate using ATP hydrolysis to drive it forward to Larginosuccinate: Argininosuccinate synthase PDB 2 NZ 2, 2. 4Å EC 6. 3. 4. 5 200 k. Da tetramer monomer shown citrulline + aspartate + ATP L-arginosuccinate + AMP + PPi 11/17/2010 Biochem: AAmetab 2; Nuc 1 15
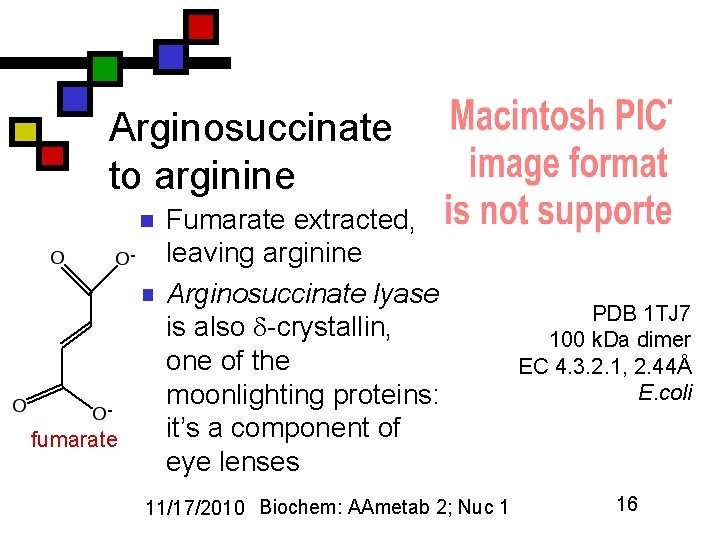
Arginosuccinate to arginine n n fumarate Fumarate extracted, leaving arginine Arginosuccinate lyase is also -crystallin, one of the moonlighting proteins: it’s a component of eye lenses 11/17/2010 Biochem: AAmetab 2; Nuc 1 PDB 1 TJ 7 100 k. Da dimer EC 4. 3. 2. 1, 2. 44Å E. coli 16

Why all that detail? n n These reactions form 75% of the urea cycle, which is an important path for amino acid and nucleic acid degradation. So we’ll need this later. 11/17/2010 Biochem: AAmetab 2; Nuc 1 17
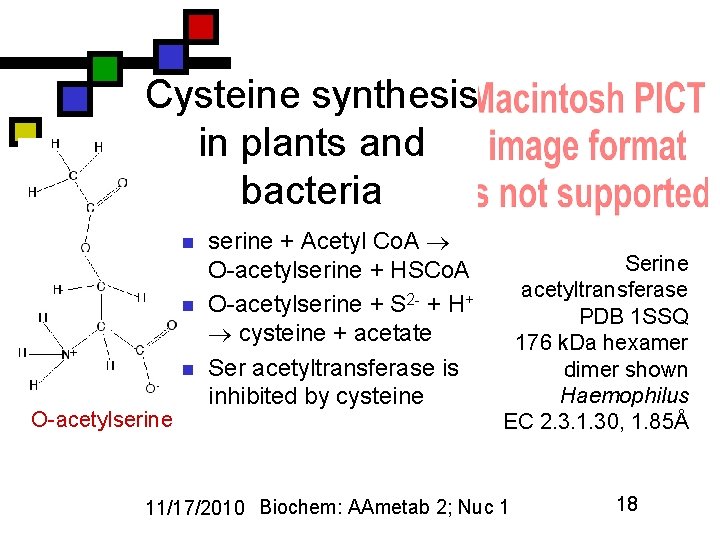
Cysteine synthesis in plants and bacteria n n n O-acetylserine + Acetyl Co. A O-acetylserine + HSCo. A O-acetylserine + S 2 - + H+ cysteine + acetate Ser acetyltransferase is inhibited by cysteine Serine acetyltransferase PDB 1 SSQ 176 k. Da hexamer dimer shown Haemophilus EC 2. 3. 1. 30, 1. 85Å 11/17/2010 Biochem: AAmetab 2; Nuc 1 18
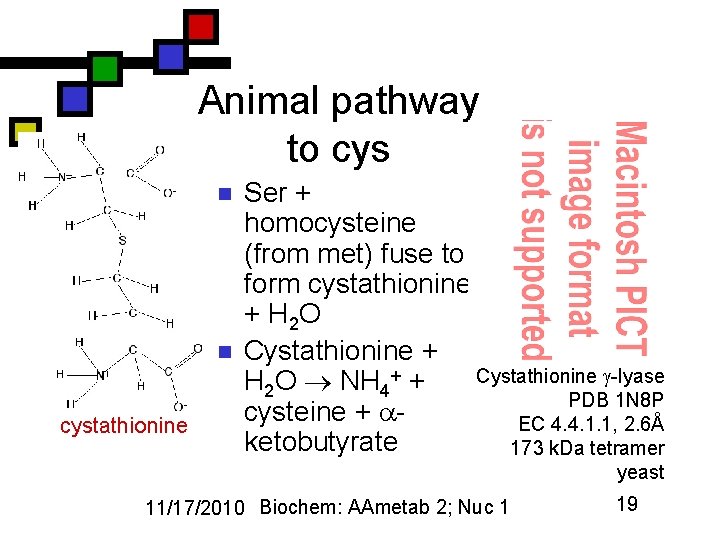
Animal pathway to cys n n cystathionine Ser + homocysteine (from met) fuse to form cystathionine + H 2 O Cystathionine + Cystathionine -lyase H 2 O NH 4+ + PDB 1 N 8 P cysteine + EC 4. 4. 1. 1, 2. 6Å ketobutyrate 173 k. Da tetramer yeast 11/17/2010 Biochem: AAmetab 2; Nuc 1 19

Marching through the list of twenty amino acids n Amino acids we’ve already covered n n Essential but straightforward Acids and amides: n lys, met, thr glu, gln, asp, asn n val, leu, ile Simple: n Essential & Ugly ala, ser, gly n phe, tyr, trp Other non-essential amino acids: n his arg, pro, cys 11/17/2010 Biochem: AAmetab 2; Nuc 1 20

Lys, met, thr n n n asp gets phosphorylated and becomes a source for all of these: asp + ATP -aspartyl phosphate + ADP via aspartate kinase -asp P + NADPH + H+ -> Pi + aspartate -semialdehyde +NADP+ This heads to lys or to homoserine Homoserine converts in a few steps to met or thr 11/17/2010 Biochem: AAmetab 2; Nuc 1 Aspartate kinase 112 k. Da EC 2. 7. 2. 4 2. 85Å PDB 2 CDQ dimer Arabidopsis 21
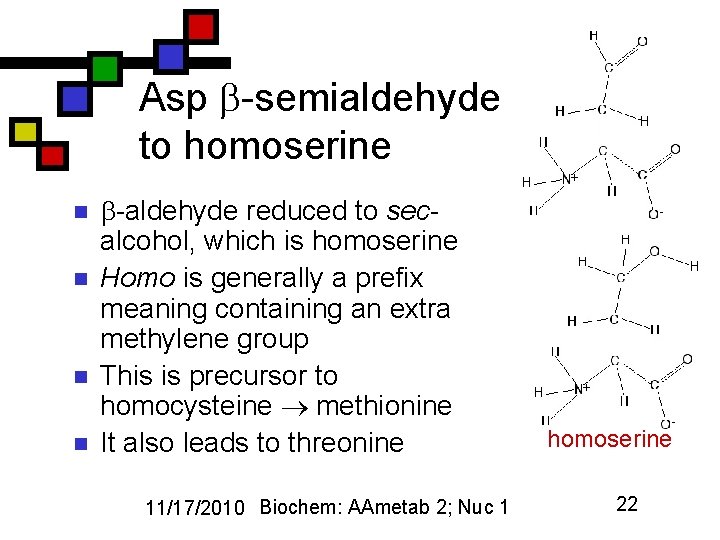
Asp -semialdehyde to homoserine n n -aldehyde reduced to secalcohol, which is homoserine Homo is generally a prefix meaning containing an extra methylene group This is precursor to homocysteine methionine It also leads to threonine 11/17/2010 Biochem: AAmetab 2; Nuc 1 homoserine 22
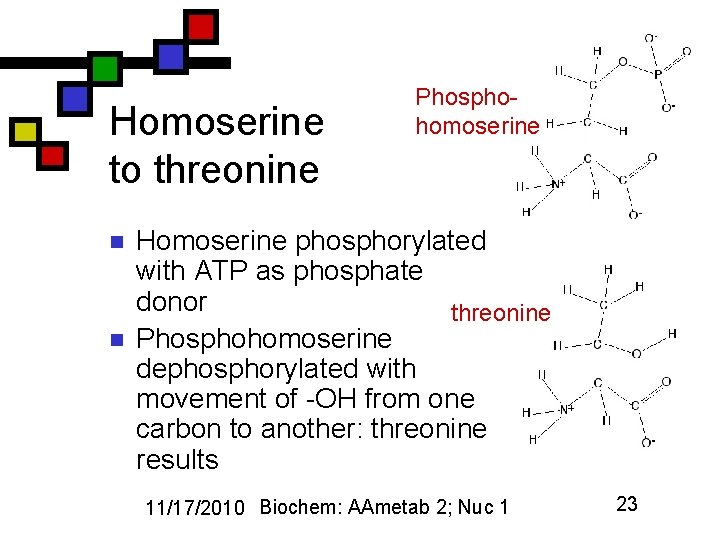
Homoserine to threonine n n Phosphohomoserine Homoserine phosphorylated with ATP as phosphate donor threonine Phosphohomoserine dephosphorylated with movement of -OH from one carbon to another: threonine results 11/17/2010 Biochem: AAmetab 2; Nuc 1 23
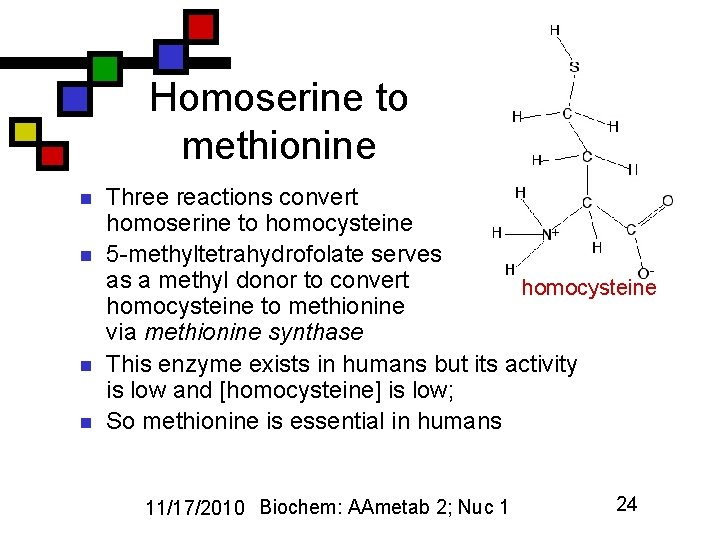
Homoserine to methionine n n Three reactions convert homoserine to homocysteine 5 -methyltetrahydrofolate serves as a methyl donor to convert homocysteine to methionine via methionine synthase This enzyme exists in humans but its activity is low and [homocysteine] is low; So methionine is essential in humans 11/17/2010 Biochem: AAmetab 2; Nuc 1 24
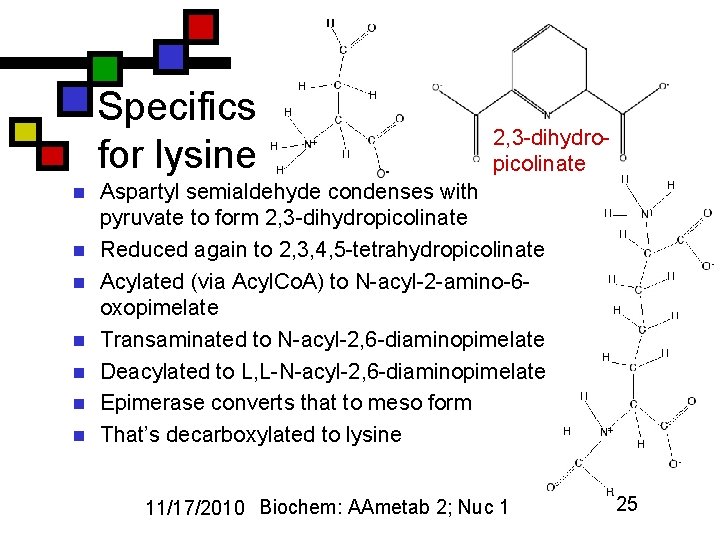
Specifics for lysine n n n n 2, 3 -dihydropicolinate Aspartyl semialdehyde condenses with pyruvate to form 2, 3 -dihydropicolinate Reduced again to 2, 3, 4, 5 -tetrahydropicolinate Acylated (via Acyl. Co. A) to N-acyl-2 -amino-6 oxopimelate Transaminated to N-acyl-2, 6 -diaminopimelate Deacylated to L, L-N-acyl-2, 6 -diaminopimelate Epimerase converts that to meso form That’s decarboxylated to lysine 11/17/2010 Biochem: AAmetab 2; Nuc 1 25

The human list (cf. box 17. 3) AA Asp Asn Lys Met Thr Ala Val Leu Ile Glu Gln moles ATP essen 21 no 22 -24 no 50 -51 yes 44 yes 31 yes 20 no 39 yes 47 yes 55 yes 30 no 31 no AA tial? Arg Pro Ser Gly Cys Phe Tyr Trp His moles ATP tial? 44 no 39 no 18 no 12 no 19 no 65 yes 62 no* 78 yes 42 yes 11/17/2010 Biochem: AAmetab 2; Nuc 1 essen- 26
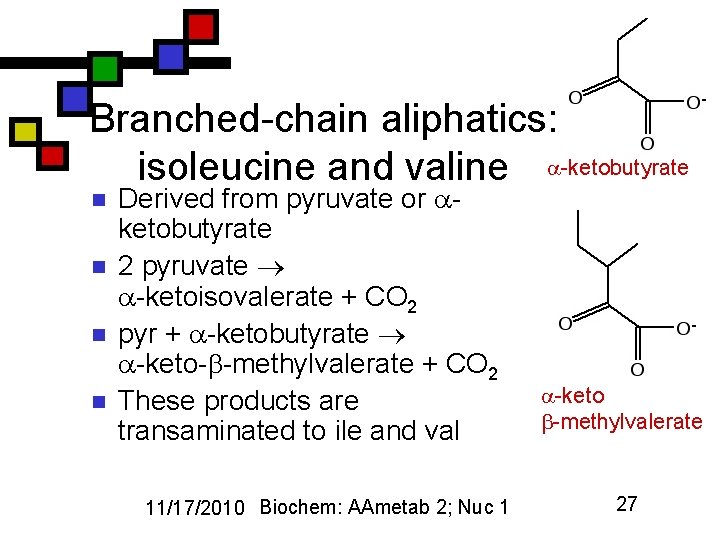
Branched-chain aliphatics: isoleucine and valine -ketobutyrate n n Derived from pyruvate or ketobutyrate 2 pyruvate -ketoisovalerate + CO 2 pyr + -ketobutyrate -keto- -methylvalerate + CO 2 These products are transaminated to ile and val 11/17/2010 Biochem: AAmetab 2; Nuc 1 -keto -methylvalerate 27
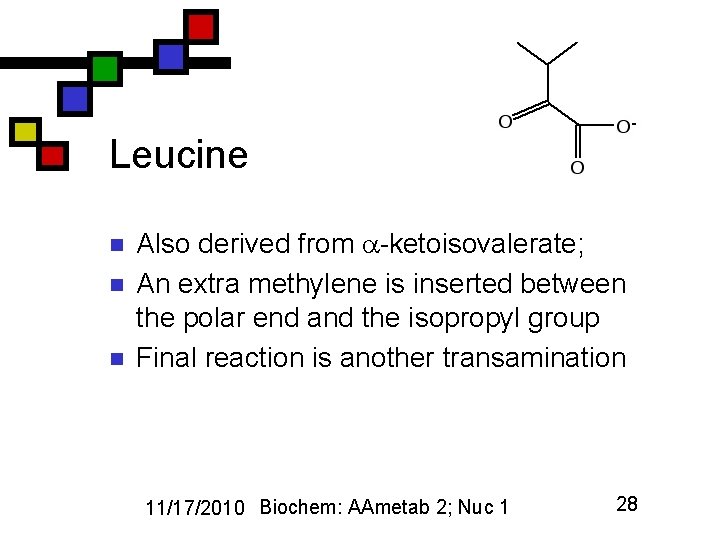
Leucine n n n Also derived from -ketoisovalerate; An extra methylene is inserted between the polar end and the isopropyl group Final reaction is another transamination 11/17/2010 Biochem: AAmetab 2; Nuc 1 28
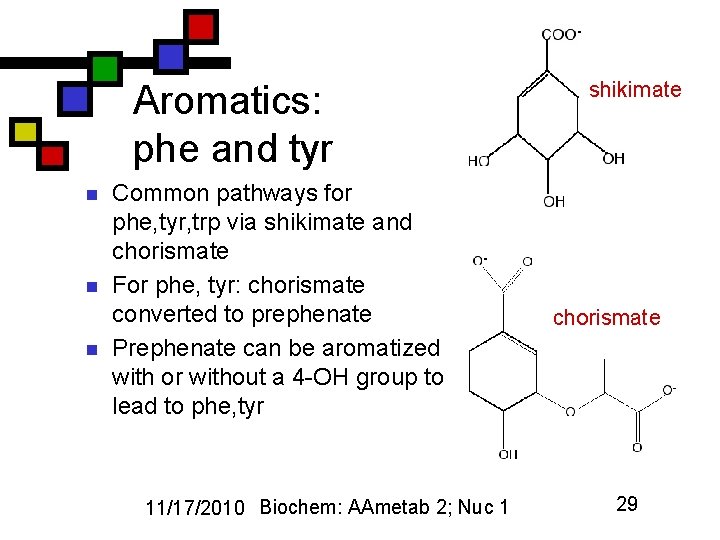
Aromatics: phe and tyr n n n Common pathways for phe, tyr, trp via shikimate and chorismate For phe, tyr: chorismate converted to prephenate Prephenate can be aromatized with or without a 4 -OH group to lead to phe, tyr 11/17/2010 Biochem: AAmetab 2; Nuc 1 shikimate chorismate 29
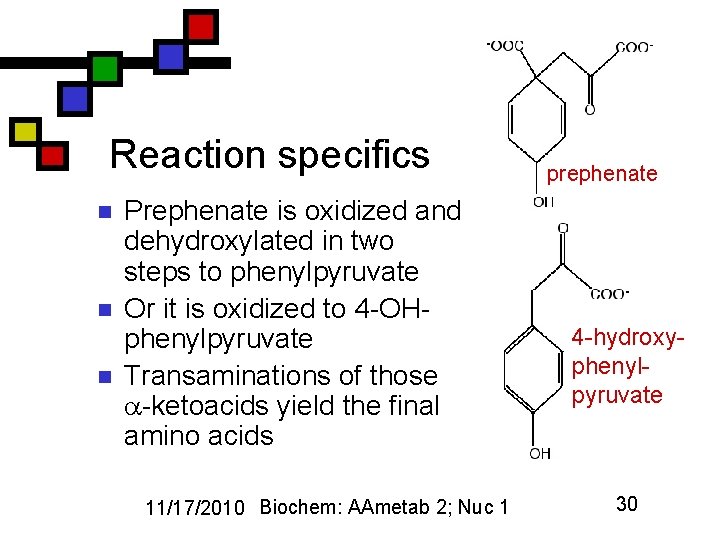
Reaction specifics n n n Prephenate is oxidized and dehydroxylated in two steps to phenylpyruvate Or it is oxidized to 4 -OHphenylpyruvate Transaminations of those -ketoacids yield the final amino acids 11/17/2010 Biochem: AAmetab 2; Nuc 1 prephenate 4 -hydroxyphenylpyruvate 30

Chorismate mutase n n Isomerase, converts chorismate to prephenate In E. coli: 2 versions depending on which path the product is heading to Active sites are similar in all organisms but architecture is very different Catalytic triad similar to serine proteases PDB 1 DBF, 1. 3Å 42 k. Da trimer EC 5. 4. 99. 5 B. subtilis 11/17/2010 Biochem: AAmetab 2; Nuc 1 31
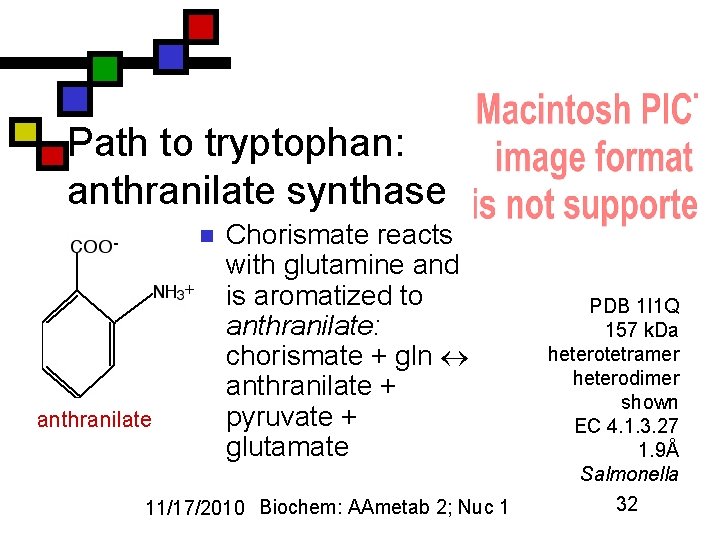
Path to tryptophan: anthranilate synthase n anthranilate Chorismate reacts with glutamine and is aromatized to anthranilate: chorismate + gln anthranilate + pyruvate + glutamate 11/17/2010 Biochem: AAmetab 2; Nuc 1 PDB 1 I 1 Q 157 k. Da heterotetramer heterodimer shown EC 4. 1. 3. 27 1. 9Å Salmonella 32
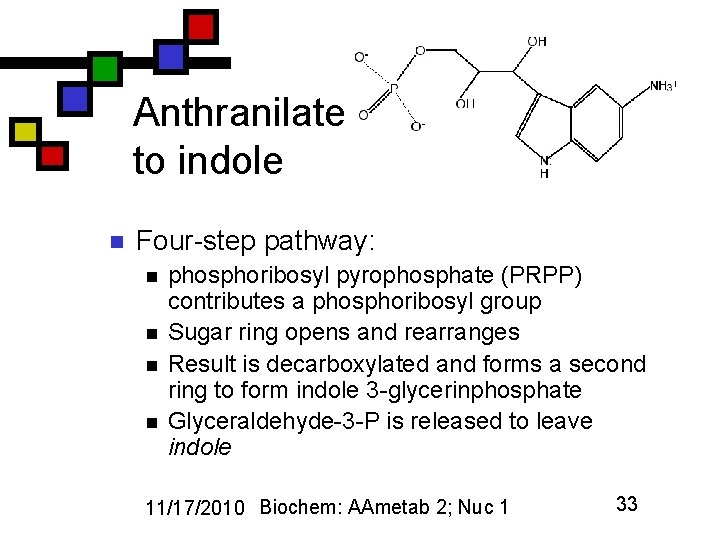
Anthranilate to indole n Four-step pathway: n n phosphoribosyl pyrophosphate (PRPP) contributes a phosphoribosyl group Sugar ring opens and rearranges Result is decarboxylated and forms a second ring to form indole 3 -glycerinphosphate Glyceraldehyde-3 -P is released to leave indole 11/17/2010 Biochem: AAmetab 2; Nuc 1 33

Tryptophan synthase n n Indole + ser tryptophan + H 2 O PLP-dependent enzyme, but different in how it uses PLP from the transaminases 11/17/2010 Biochem: AAmetab 2; Nuc 1 PDB 2 CLF 146 k. Da heterotetramer; heterodimer shown EC 4. 2. 1. 20 1. 7Å Salmonella 34

Genetic control of aromatic aa synthesis n In E. coli and many other bacteria, a single operon controls several chorismate -related genes: the aro pathway 11/17/2010 Biochem: AAmetab 2; Nuc 1 35

Histidine (fig. 17. 22) n n n Start with PRPP and ATP: form phosphoribosyl ATP 3 reactions involving glutamine as nitrogen donor for ring lead to imidazole glycerol phosphate That gets modified and transaminated to make histidine 11/17/2010 Biochem: AAmetab 2; Nuc 1 36

What do we do with amino acids? n n Obviously a lot of them serve as buildingblocks for protein and peptide synthesis via ribosomal mechanisms Also serve as metabolites, getting converted to other compounds or getting oxidized as fuel 11/17/2010 Biochem: AAmetab 2; Nuc 1 37
Transaminations n Generally two stages: n n n amino acid + -ketoglutarate -keto acid + glutamate Glutamate + NAD+ + H 2 O -ketoglutarate + NADH + H+ + NH 4+ Net reaction is amino acid + NAD+ + H 2 O -keto acid + NADH + H+ + NH 4+ 11/17/2010 Biochem: AAmetab 2; Nuc 1 38

Glucogenic and ketogenic amino acids n Degradation of many amino acids lead to TCA cycle intermediates or pyruvate n n n therefore these can be built back up to glucose; these are called glucogenic Degradation of others leads to acetyl Co. A and related compounds n n these cannot be built back up to glucose except via the glyoxalate shuttle these are called ketogenic 11/17/2010 Biochem: AAmetab 2; Nuc 1 39

Glucogenic amino acids n n Amino acids that can be catabolized to produce building blocks that lead to glucose without help of glyoxalate pathway Most produce succinate, succinyl Co. A, fumarate, -ketoglutarate, or oxaloacetate 11/17/2010 Biochem: AAmetab 2; Nuc 1 40

Ketogenic amino acids n n These do not produce TCA cycle intermediates, but rather produce acetyl Co. A or its close relatives Can be built back up into fats or ketone bodies By convention, amino acids that produce both acetyl Co. A and TCA cycle intermediates or pyruvate are considered glucogenic On that basis, only leu and lys are purely ketogenic 11/17/2010 Biochem: AAmetab 2; Nuc 1 41

Serine-based metabolites n n Serine is a building block for sphinganine and therefore for sphingolipids Serine also leads to phosphatidylserine, which is important by itself and can be metabolized to phosphatidylethanolamine and phosphatidylcholine 11/17/2010 Biochem: AAmetab 2; Nuc 1 42

Serine degradation n Two paths for degrading serine: n n PLP-dependent serine dehydratase simply deaminates ser to pyruvate; this enzyme is like trp synthase More common: SHMT transfers hydroxymethyl group to THF, leaving glycine; we’ve seen that one as a biosynthetic enzyme for making glycine 11/17/2010 Biochem: AAmetab 2; Nuc 1 Serine dehydratase PDB 1 P 5 J 41 k. Da monomer Human EC 4. 3. 1. 17, 2. 5Å 43

Glycine-based metabolites n porphobilinogen Glycine is a source for purines, glyoxylate, creatine phosphate, and (with the help of succinyl Co. A) porphobilinogen, whence we get porphyrins, and from those we get chlorophyll, heme, and cobalamin 11/17/2010 Biochem: AAmetab 2; Nuc 1 44

Glycine cleavage system n n n Glycine + H 2 O + NAD+ + THF NADH + H+ + HCO 3 - + NH 4+ + 5, 10 -methylene. THF Complex system: PLP, lipoamide, FAD prosthetic groups Lipoamide swinging arm works as in pyruvate dehydrogenase 11/17/2010 Biochem: AAmetab 2; Nuc 1 T protein (aminomethyltransferase) of glycine cleavage system PDB 1 V 5 V 93 k. Da dimer Pyrococcus EC 2. 10, 1. 5Å 45

asp, glu, ala degradation I n Standard transmination converts aspartate to oxaloacetate with release of glutamate, which then can be deaminated to re-form -ketoglutarate: asp + -kg oxaloacetate + glu n glu + NAD+ + H 2 O -kg + NADH + H+ + NH 4+ n 11/17/2010 Biochem: AAmetab 2; Nuc 1 46

Asp, glu, ala degradation II n n Deamination converts glutamate to -ketoglutarate, as above Standard transamination converts alanine to pyruvate according to the same logic as asp 11/17/2010 Biochem: AAmetab 2; Nuc 1 47

All three of these are glucogenic! n n -ketoglutarate and oxaloacetate are TCA cycle intermediates Pyruvate can be regarded as a glucose precursor via the gluconeogenesis pathways (pyr -> OAA -> PEP -> …) 11/17/2010 Biochem: AAmetab 2; Nuc 1 48

Degradation of asn, gln n n Asparagine and glutamine are deaminated on their side-chains to asp and glu Thus they lead to oxaloacetate and -ketoglutarate, respectively So they’re glucogenic The initial hydrolyses (deaminations) are catalyzed by asparaginase and glutaminase 11/17/2010 Biochem: AAmetab 2; Nuc 1 Asparaginase PDB 1 O 7 J 144 k. Da tetramer EC 3. 5. 1. 1, 1Å Erwinia chrysanthemi 49

Arginine degradation n Arginine is hydrolyzed to urea and ornithine as part of the urea cycle; enzyme is arginase PLP-dependent enzyme converts ornithine to glu semialdehyde That’s oxidized to glutamate 11/17/2010 Biochem: AAmetab 2; Nuc 1 Arginase PDB 2 AEB 212 k. Da hexamer Dimer shown Human EC 3. 5. 3. 1, 1. 29Å 50
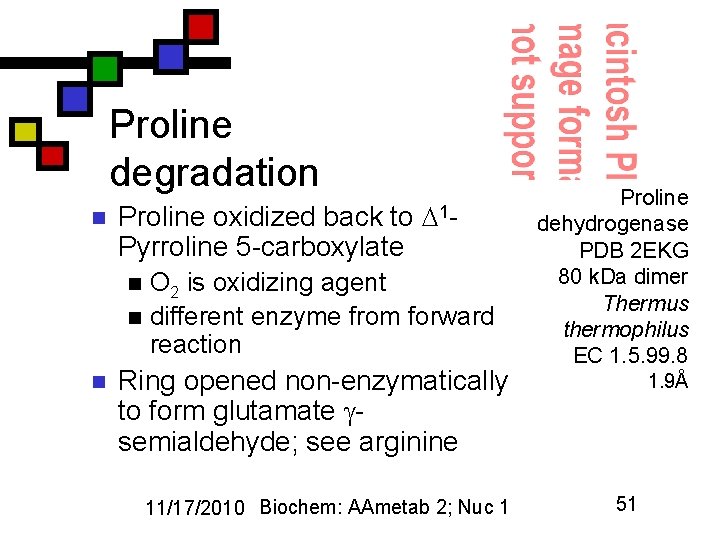
Proline degradation n Proline oxidized back to Pyrroline 5 -carboxylate 1 - O 2 is oxidizing agent n different enzyme from forward reaction n n Ring opened non-enzymatically to form glutamate semialdehyde; see arginine 11/17/2010 Biochem: AAmetab 2; Nuc 1 Proline dehydrogenase PDB 2 EKG 80 k. Da dimer Thermus thermophilus EC 1. 5. 99. 8 1. 9Å 51
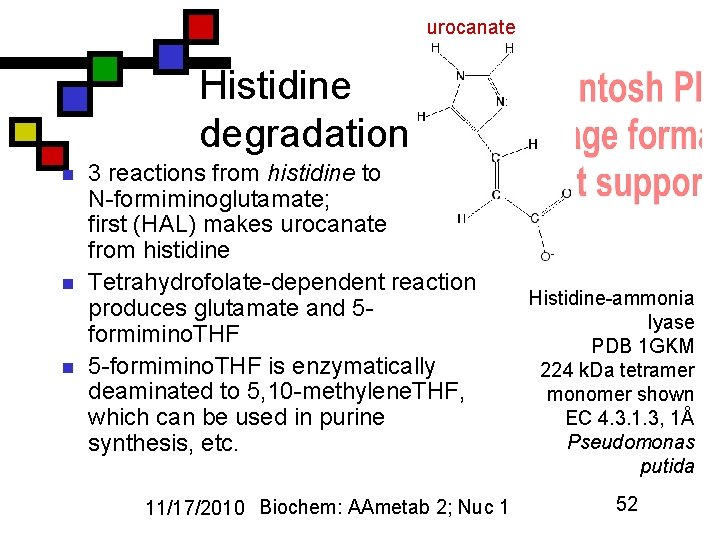
urocanate Histidine degradation n 3 reactions from histidine to N-formiminoglutamate; first (HAL) makes urocanate from histidine Tetrahydrofolate-dependent reaction produces glutamate and 5 formimino. THF 5 -formimino. THF is enzymatically deaminated to 5, 10 -methylene. THF, which can be used in purine synthesis, etc. 11/17/2010 Biochem: AAmetab 2; Nuc 1 Histidine-ammonia lyase PDB 1 GKM 224 k. Da tetramer monomer shown EC 4. 3. 1. 3, 1Å Pseudomonas putida 52

How are we doing so far? n n n We did ser and gly first because they’re so important Then we’ve done a whole bunch that connect up to glutamate (or asp): asp, glu, ala, asn, gln, arg, pro, his So we’re halfway through. 11/17/2010 Biochem: AAmetab 2; Nuc 1 53

Threonine degradation n n Several pathways (fig. 17. 29) Major one: oxidize threonine to 2 -amino-3 -ketobutyrate 2 -amino-3 -ketobutyrate reacts with HS-Co. A to form acetyl Co. A and glycine So this one is ketogenic Other pathways are glucogenic 11/17/2010 Biochem: AAmetab 2; Nuc 1 Threonine dehydrogenase PDB 2 DFV 116 k. Da trimer EC 1. 103 2. 05Å Pyrococcus 54

Valine degradation (fig. 17. 30, center) n n Valine transaminated to -ketoisovalerate Branched-chain -keto acid dehydrogenase PDB 2 VBF (TTP, Lipoamide): 125 k. Da dimer + -ketoisovalerate + NAD + HS-Co. A Lactococcus -ketoisovaleryl Co. A EC 4. 1. 1. 72 Next reaction (acyl Co. A dehydrogenase) 1. 6Å 2 -methyl-1 -propenyl Co. A + NADH + CO 2 Product undergoes 4 reactions to propionyl Co. A and thence to succinyl Co. A: glucogenic 11/17/2010 Biochem: AAmetab 2; Nuc 1 55

Isoleucine and leucine degradation n Same path but products are: Leucine’s products: acetyl Co. A + acetoacetate: ketogenic Isoleucine: Acetyl Co. A + propionyl Co. A: ketogenic and glucogenic 11/17/2010 Biochem: AAmetab 2; Nuc 1 56

Methionine degradation n Considerable methionine is turned into S-adenosylmethionine: n n Methyl donor Leaves behind S-Adenosylhomocysteine S-adenosylhomocysteine can be hydrolyzed to homocysteine and adenosine Homocysteine can condense with serine to form cystathionine, which can yield cysteine and -ketobutyrate… and we know how to turn -ketobutyrate into propionyl Co. A. So met is glucogenic. 11/17/2010 Biochem: AAmetab 2; Nuc 1 57
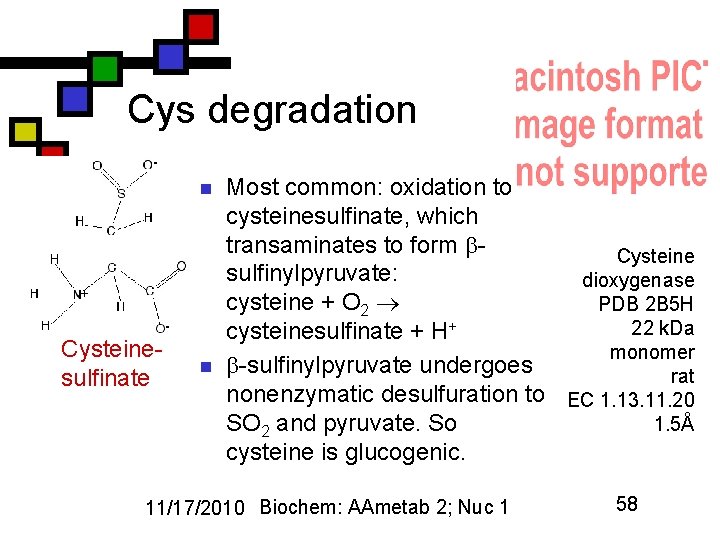
Cys degradation n Cysteinesulfinate n Most common: oxidation to cysteinesulfinate, which transaminates to form Cysteine sulfinylpyruvate: dioxygenase cysteine + O 2 PDB 2 B 5 H 22 k. Da cysteinesulfinate + H+ monomer -sulfinylpyruvate undergoes rat nonenzymatic desulfuration to EC 1. 13. 11. 20 1. 5Å SO 2 and pyruvate. So cysteine is glucogenic. 11/17/2010 Biochem: AAmetab 2; Nuc 1 58

Tetrahydrobiopterin Phenylalanine n n Simple: phenylalanine gets hydroxylated to form tyrosine: phenylalanine + O 2 tyrosine This is a Fe 2+ and tetrahydrobiopterindependent enzyme—a folate-like cofactor Phenylalanine hydroxylase PDB 1 J 8 U 71 k. Da dimer; monomer shown human (residues 103 -427) EC 1. 14. 16. 1, 1. 5Å 11/17/2010 Biochem: AAmetab 2; Nuc 1 59

Phenylketonuria n n n Usually associated with mutation in phenylalanine hydroxylase: Accumulated Phe phenylpyruvate Afflicts 1/15000 newborns Built-up phenylpyruvate causes irreversible mental retardation Type IV PKU related to deficiencies in enzymes that restore tetrahydrobiopterin (see fig. 17. 33, bottom) 11/17/2010 Biochem: AAmetab 2; Nuc 1 60
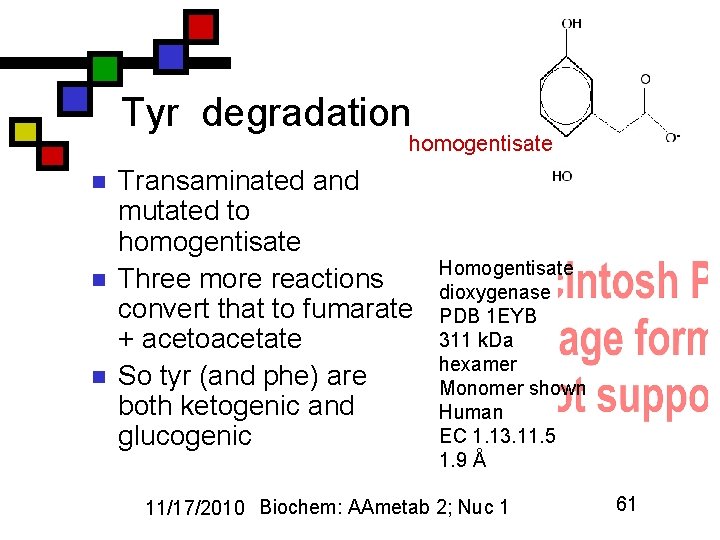
Tyr degradation homogentisate n n n Transaminated and mutated to homogentisate Three more reactions convert that to fumarate + acetoacetate So tyr (and phe) are both ketogenic and glucogenic Homogentisate dioxygenase PDB 1 EYB 311 k. Da hexamer Monomer shown Human EC 1. 13. 11. 5 1. 9 Å 11/17/2010 Biochem: AAmetab 2; Nuc 1 61
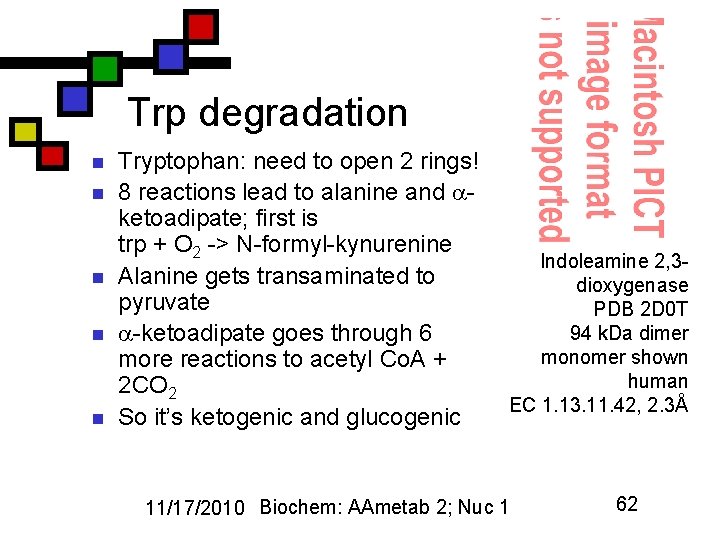
Trp degradation n n Tryptophan: need to open 2 rings! 8 reactions lead to alanine and ketoadipate; first is trp + O 2 -> N-formyl-kynurenine Alanine gets transaminated to pyruvate -ketoadipate goes through 6 more reactions to acetyl Co. A + 2 CO 2 So it’s ketogenic and glucogenic Indoleamine 2, 3 dioxygenase PDB 2 D 0 T 94 k. Da dimer monomer shown human EC 1. 13. 11. 42, 2. 3Å 11/17/2010 Biochem: AAmetab 2; Nuc 1 62

Lys degradation (fig. 17. 35) n n n Condense lysine with -ketoglutarate to form saccharopine Saccharopine That’s deglutamated (? ), oxidized, and dehydrogenase transaminated to -ketoadipate PDB 2 AXQ Six reactions degrade that to 2 acetyl 103 k. Da dimer Co. A molecules plus 2 CO 2 monomer shown Purely ketogenic Yeast EC 1. 5. 1. 10, 1. 7Å Some bacteria decarboxylate it to cadaverine 11/17/2010 Biochem: AAmetab 2; Nuc 1 63
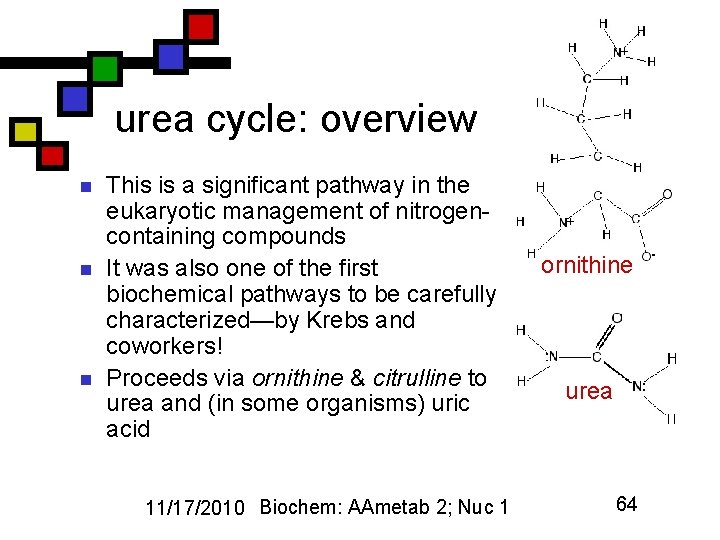
urea cycle: overview n n n This is a significant pathway in the eukaryotic management of nitrogencontaining compounds It was also one of the first biochemical pathways to be carefully characterized—by Krebs and coworkers! Proceeds via ornithine & citrulline to urea and (in some organisms) uric acid 11/17/2010 Biochem: AAmetab 2; Nuc 1 ornithine urea 64
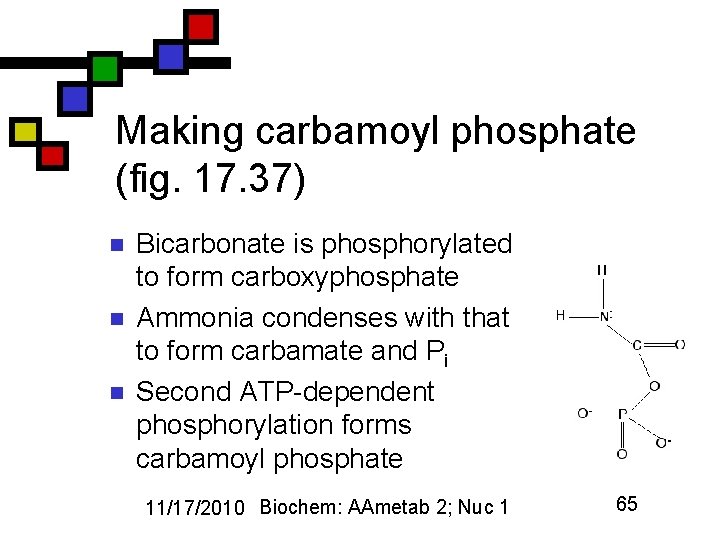
Making carbamoyl phosphate (fig. 17. 37) n n n Bicarbonate is phosphorylated to form carboxyphosphate Ammonia condenses with that to form carbamate and Pi Second ATP-dependent phosphorylation forms carbamoyl phosphate 11/17/2010 Biochem: AAmetab 2; Nuc 1 65

Urea cycle itself n n n In mitochondrion: carbamoyl phosphate condenses with ornithine to form citrulline Citrulline condenses with urea to form arginosuccinate Arginosuccinate is cleaved nonhydrolytically to fumarate and arginine Arginine yields urea and citrulline Citrulline re-enters cycle 11/17/2010 Biochem: AAmetab 2; Nuc 1 66

i. Clicker quiz: question 1 n n n 1. Glutamate + ammonia glutamine + H 2 O is only slightly endergonic ( Go’ = +14 k. J mol-1), yet it is coupled to ATP hydrolysis. Why? (a) You can never run a reaction with a positive Go’ (b) [glutamate] ~ [glutamine] in the cell (c) If you heat the substrates, they disintegrate (d) ammonia is toxic in the absence of ATP 11/17/2010 Biochem: AAmetab 2; Nuc 1 67

i. Clicker quiz #2 n 2. Which ribosomal amino acid’s biosynthesis is closely associated with the urea cycle? n n n (a) alanine (b) serine (c) ornithine (d) arginine (e) none of the above. 11/17/2010 Biochem: AAmetab 2; Nuc 1 68

6 1 5 Pyrimidines n n n 4 2 3 Single-ring nucleic acid bases 6 -atom ring; always two nitrogens in the ring, meta to one another Based on pyrimidine, although pyrimidine itself is not a biologically important molecule Variations depend on oxygens and nitrogens attached to ring carbons Tautomerization possible Note line of symmetry in pyrimidine structure 11/17/2010 Biochem: AAmetab 2; Nuc 1 69

Uracil and thymine n n Uracil is a simple dioxo derivative of pyrimidine: 2, 4 -dioxopyrimidine Thymine is 5 -methyluracil Uracil is found in RNA; Thymine is found in DNA We can draw other tautomers where we move the protons to the oxygens 11/17/2010 Biochem: AAmetab 2; Nuc 1 70

Tautomers n n Lactam and Lactim forms Getting these right was essential to Watson & Crick’s development of the DNA double helical model 11/17/2010 Biochem: AAmetab 2; Nuc 1 71

Cytosine n n This is 2 -oxo, 4 -aminopyrimidine It’s the other pyrimidine base found in DNA & RNA Spontaneous deamination (C U) Again, other tautomers can be drawn 11/17/2010 Biochem: AAmetab 2; Nuc 1 72
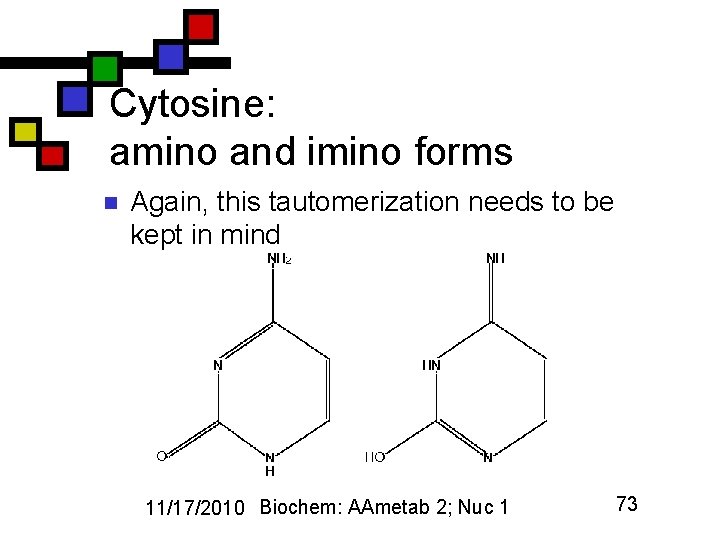
Cytosine: amino and imino forms n Again, this tautomerization needs to be kept in mind 11/17/2010 Biochem: AAmetab 2; Nuc 1 73
- Slides: 73