Alignment of large genomic sequences Fragmentbased alignment approach

Alignment of large genomic sequences Fragment-based alignment approach (DIALIGN) useful for alignment of genomic sequences. Possible applications: n Detection of regulatory elements n Identification of pathogenic microorganisms n Gene prediction

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacccctgaataa

The DIALIGN approach atc------taatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaa--gagtatcacc-----cctgaataa

The DIALIGN approach atc------taatagttaaactcccccgtgc-ttag cagtgcgtgtattactaac-----gg-ttcaatcgcg caaa--gagtatcacc-----cctgaataa

The DIALIGN approach Consistency! atc------taatagttaaactcccccgtgc-ttag cagtgcgtgtattactaac-----gg-ttcaatcgcg caaa--gagtatcacc-----cctgaataa

The DIALIGN approach atc------TAATAGTTAaactcccc. CGTGC-TTag cagtgc. GTGTATTACTAAc-----GG-TTCAATcgcg caaa--GAGTATCAcc-----CCTGaa. TTGAATaa

First step in sequence comparison: alignment S 1 S 2 S 3
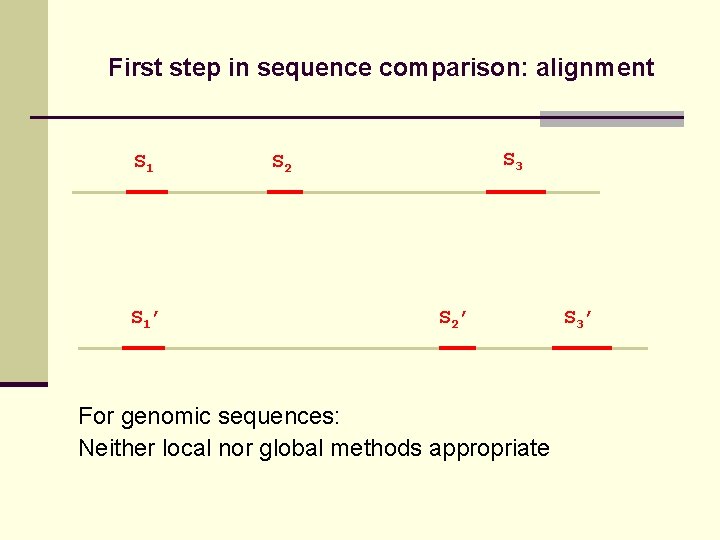
First step in sequence comparison: alignment S 1’ S 3 S 2’ For genomic sequences: Neither local nor global methods appropriate S 3’

First step in sequence comparison: alignment S 1’ S 3 S 2’ S 3’ Local method finds single best local similarity
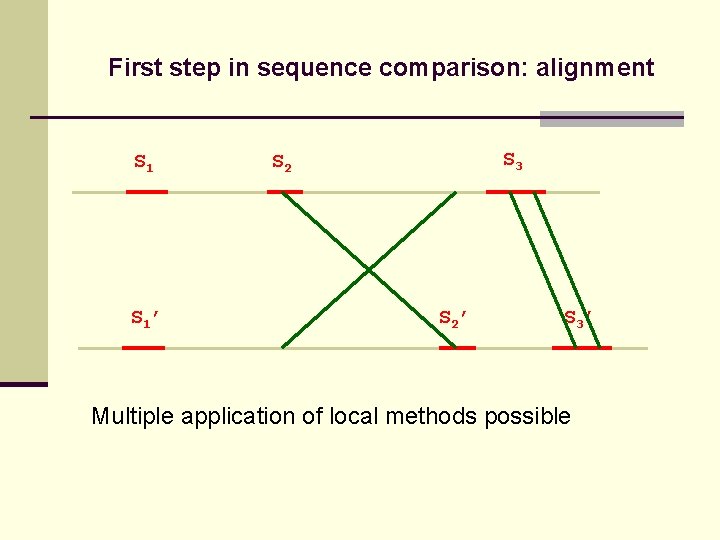
First step in sequence comparison: alignment S 1’ S 3 S 2’ S 3’ Multiple application of local methods possible

First step in sequence comparison: alignment S 1’ S 3 S 2’ S 3’ Multiple application of local methods possible
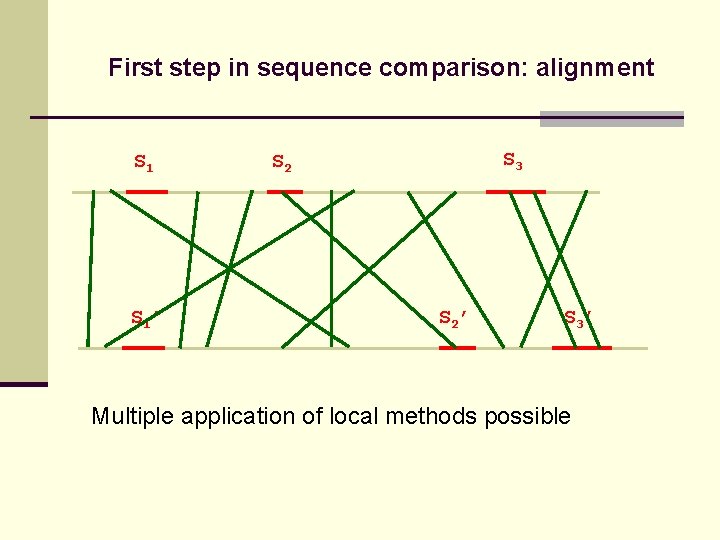
First step in sequence comparison: alignment S 1’ S 3 S 2’ S 3’ Multiple application of local methods possible
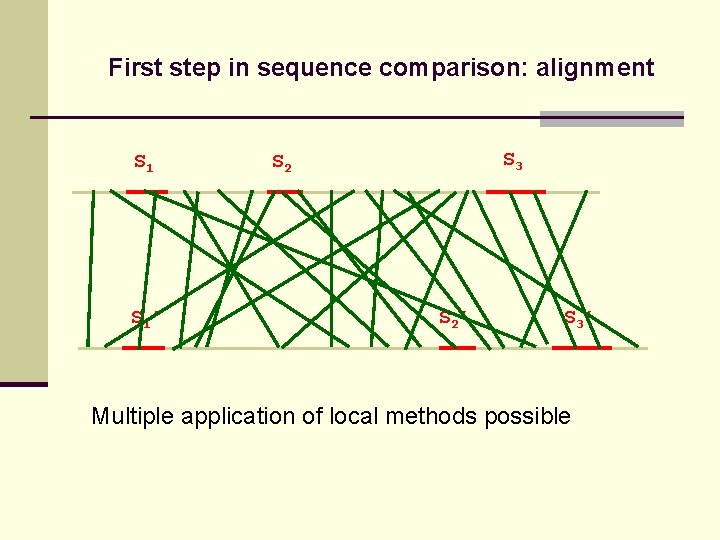
First step in sequence comparison: alignment S 1’ S 3 S 2’ S 3’ Multiple application of local methods possible
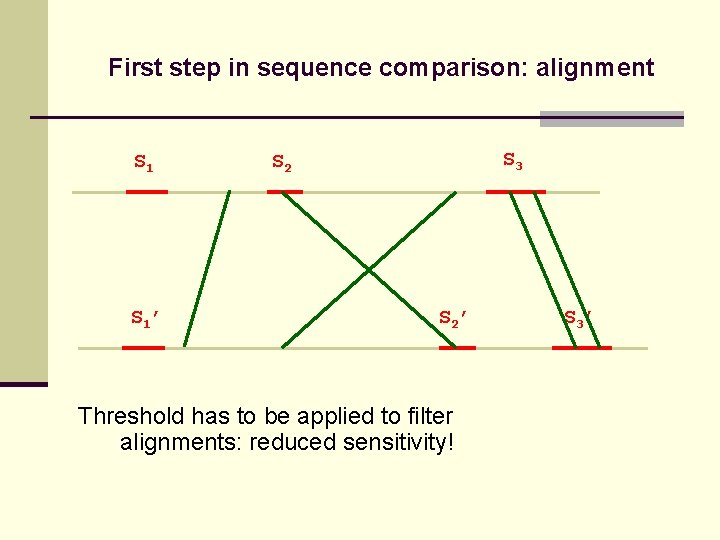
First step in sequence comparison: alignment S 1’ S 3 S 2’ Threshold has to be applied to filter alignments: reduced sensitivity! S 3’

First step in sequence comparison: alignment Alternative approach: n n During evolution few large-scale rearrangements -> relative order homologies conserved Search for chain of local homologies

First step in sequence comparison: alignment S 1’ S 3 S 2’ Genomic alignment: chain of homologies S 3’
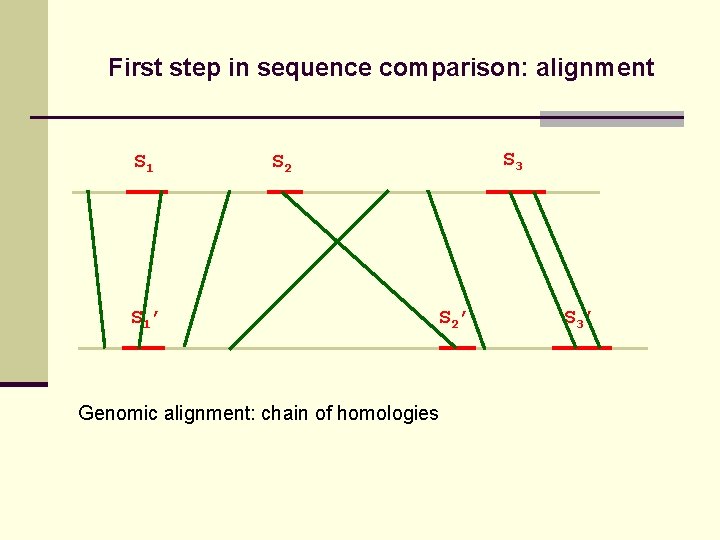
First step in sequence comparison: alignment S 1’ S 3 S 2’ Genomic alignment: chain of homologies S 3’

First step in sequence comparison: alignment S 1’ S 3 S 2 ’ Genomic alignment: chain of homologies S 3’
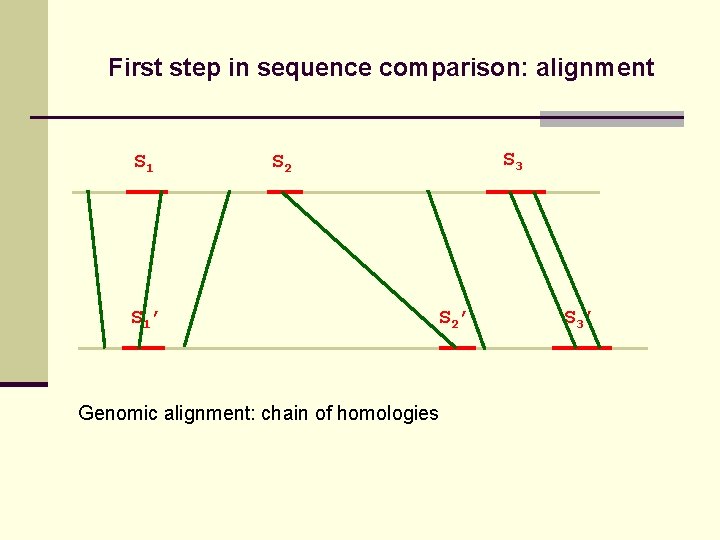
First step in sequence comparison: alignment S 1’ S 3 S 2’ Genomic alignment: chain of homologies S 3’

First step in sequence comparison: alignment Novel approaches for genomic alignment: n n n n WABA Pip. Maker MGA TBA Lagan Avid DIALIGN
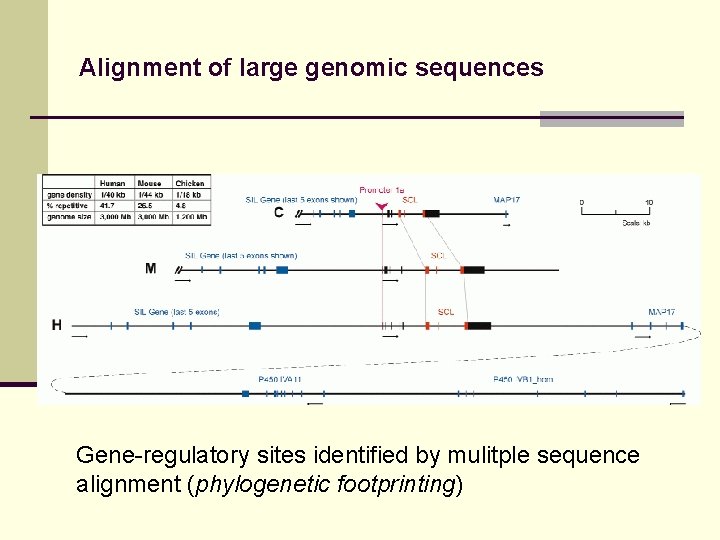
Alignment of large genomic sequences Gene-regulatory sites identified by mulitple sequence alignment (phylogenetic footprinting)
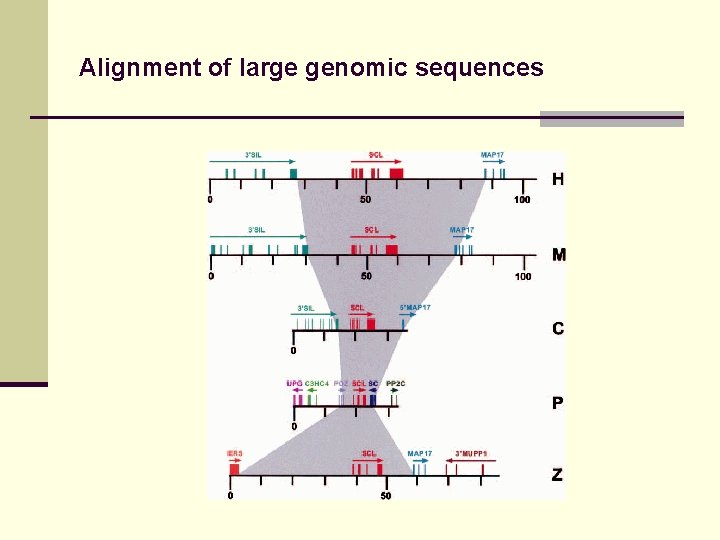
Alignment of large genomic sequences

Objective function for DIALIGN: n Weight score for every possible fragment f based on P-value: P(f) = probability of finding a fragment “like f” by chance in random sequences with same length as input sequences w(f) = -log P(f) (“weight score” of f) ”like f” means: at least same # matches (DNA, RNA) or sum of similarity values (proteins)

Objective function for DIALIGN: n Score of alignment: sum of weight scores of fragments – no gap penalty!

Optimization problem for DIALIGN: n Find consistent collection of fragments with maximum total weight score!

Alternative fragment weight scores for genomic sequences: n Calculate fragment scores at nucleotide level and at peptide level.

catcatatcttacgttaactcccccgt cagtgcgtgatagcccatatccgg

catcatatcttacgttaactcccccgt cagtgcgtgatagcccatatccgg

catcatatcttacgttaactcccccgt cagtgcgtgatagcccatatccgg Standard score: Consider length, # matches, compute probability of random occurrence
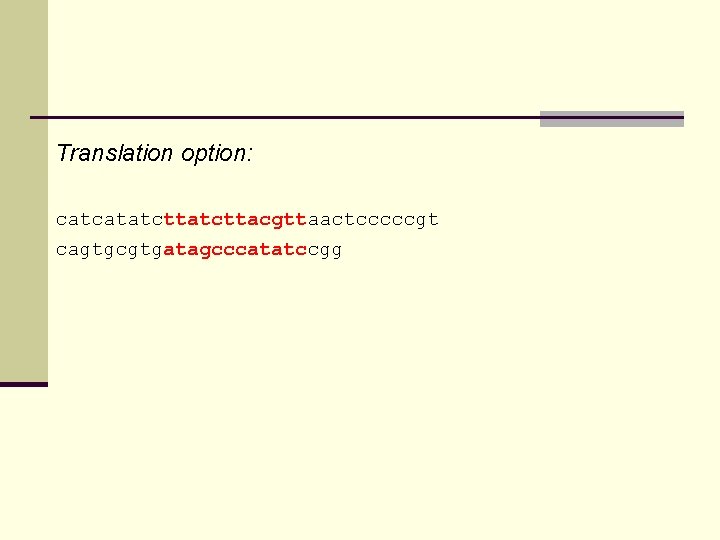
Translation option: catcatatcttacgttaactcccccgt cagtgcgtgatagcccatatccgg

Translation option: L S Y V catcatatc tta tct tac gtt aactcccccgt cagtgcgtg ata gcc cat atc cgg I A H I DNA segments translated to peptide segments; fragment score based on peptide similarity: Calculate probability of finding a fragment of the same length with (at least) the same sum of BLOSUM values

P-fragment (in both orientations) L S Y V catcatatc tta tct tac gtt aactcccccgt cagtgcgtg ata gcc cat atc cgg I A H I N-fragment catcatatc ttatcttacgtt aactcccccgtgct || | cagtgcgtg atagcccatatc cg For each fragment f three probability values calculated; Score of f based on smallest P value.

Alternative fragment weight scores for genomic sequences: n Calculate fragment scores at nucleotide level and at peptide level.
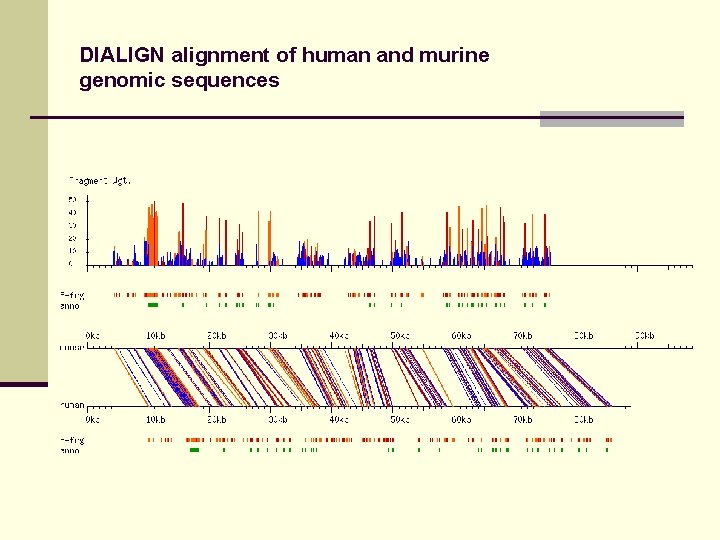
DIALIGN alignment of human and murine genomic sequences
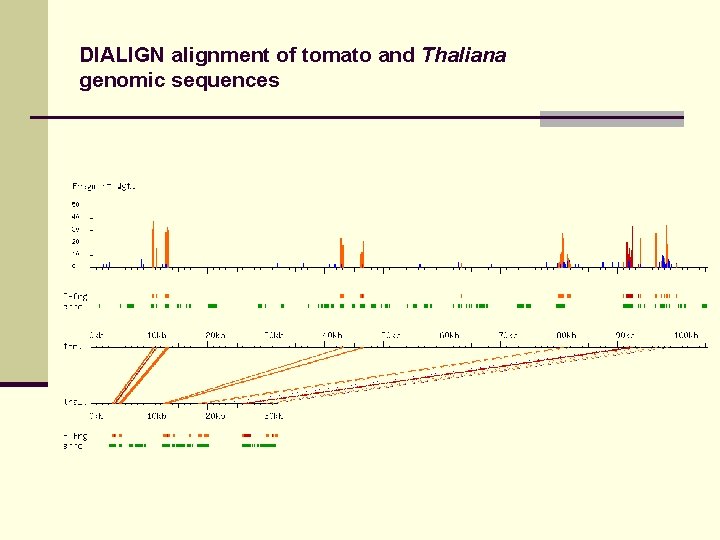
DIALIGN alignment of tomato and Thaliana genomic sequences

Alignment of large genomic sequences Evaluation of signal detection methods: Apply method to data with known signals (correct answer is known!). E. g. experimentally verified genes for gene finding n TP = true positves = # signals correctly predicted (i. e. signal present) n FP = false positives = # signals predicted but wrong (i. e no signal present) n TN = true negative = # no signal predicted, no signal present n FN = false negative = # no signal predicted, signal present!

Alignment of large genomic sequences Sn = Sensitivity = correctly predicted signals / present signals = TP / (TP + FN) Sp = Specificity = correctly predicted signals / predicted signals n = TP / (TP + FP)

Alignment of large genomic sequences Comprehensive evaluation of signal prediction method: Method assigns score to predictions Apply threshold parameter High threshold -> high specificity (Sp), low sensitivity (Sn) Low threshold -> high sensitivity , low specificity ROC curve („receiver-operator curve“) Vary threshold parameter, plot Sn against Sp
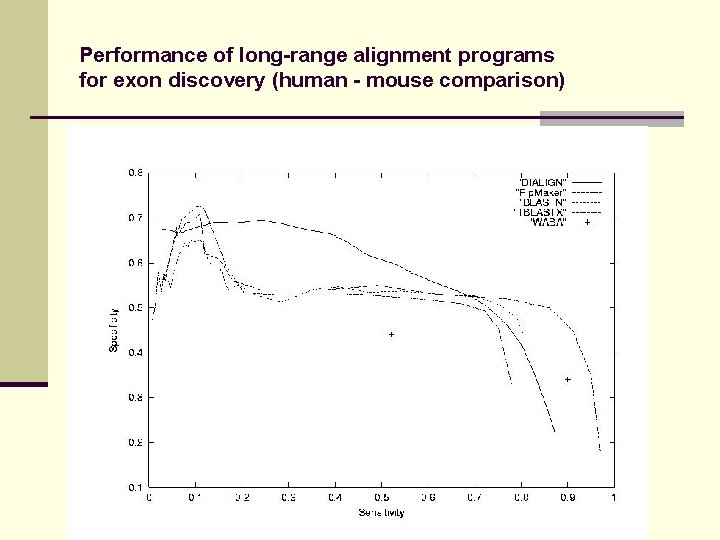
Performance of long-range alignment programs for exon discovery (human - mouse comparison)
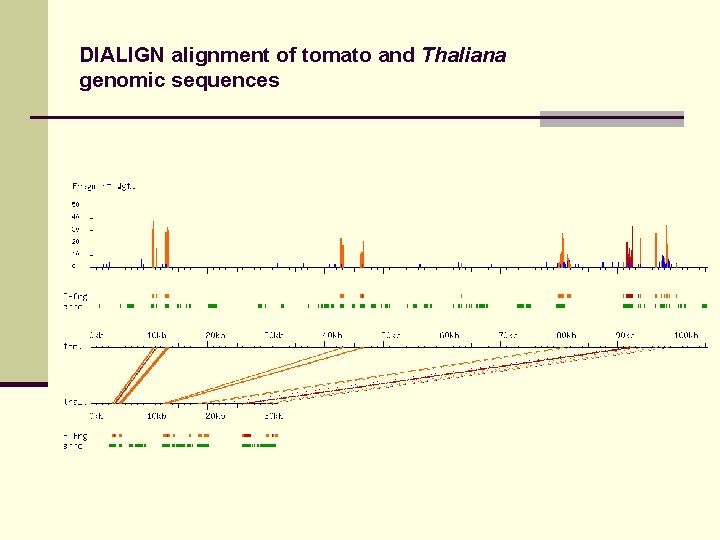
DIALIGN alignment of tomato and Thaliana genomic sequences

AGen. DA: Alignment-based Gene Detection Algorithm n Bridge small gaps between DIALIGN fragments -> cluster of fragments

AGen. DA: Alignment-based Gene Detection Algorithm n Bridge small gaps between DIALIGN fragments -> cluster of fragments n Search conserved splice sites and start/stop codons at cluster boundaries to Identify candidate exons

AGen. DA: Alignment-based Gene Detection Algorithm n Bridge small gaps between DIALIGN fragments -> cluster of fragments n Search conserved splice sites and start/stop codons at cluster boundaries to Identify candidate exons n Recursive algorithm finds biologically consistent chain of potential exons

Identification of candidate exons Fragments in DIALIGN alignment
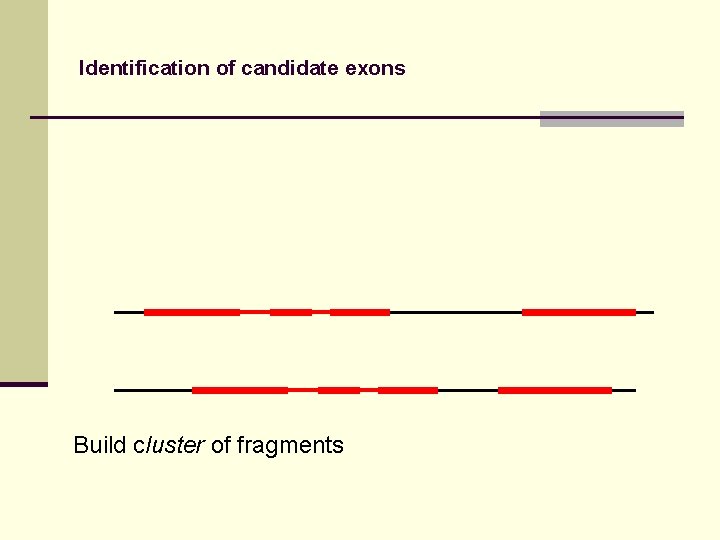
Identification of candidate exons Build cluster of fragments

Identification of candidate exons Identify conserved splice sites

Identification of candidate exons Candidate exons bounded by conserved splice sites

Construct gene models using candidate exons n Score of candidate exon (E) based on DIALIGN scores for fragments, score of splice junctions and penalty for shortening / extending n Find biologically consistent chain of candidate exons (starting with start codon, ending with stop codon, no internal stop codons …) with maximal total score

Find optimal consistent chain of candidate exons
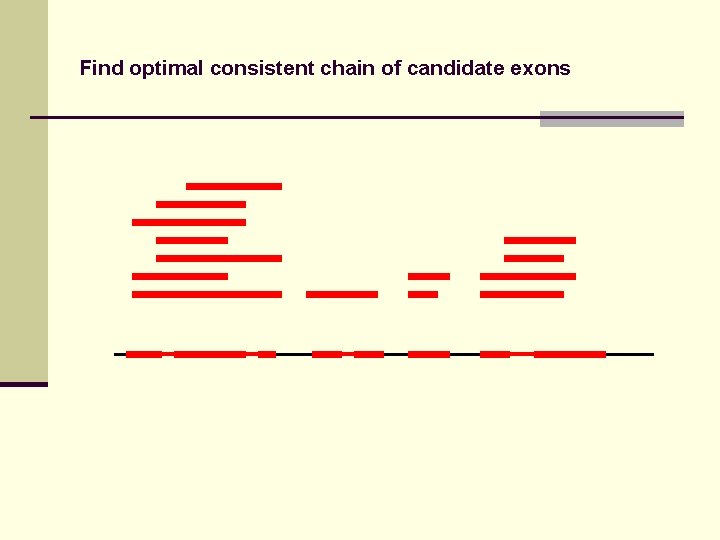
Find optimal consistent chain of candidate exons

Find optimal consistent chain of candidate exons

Find optimal consistent chain of candidate exons

Find optimal consistent chain of candidate exons atg gt ag tga atg tga
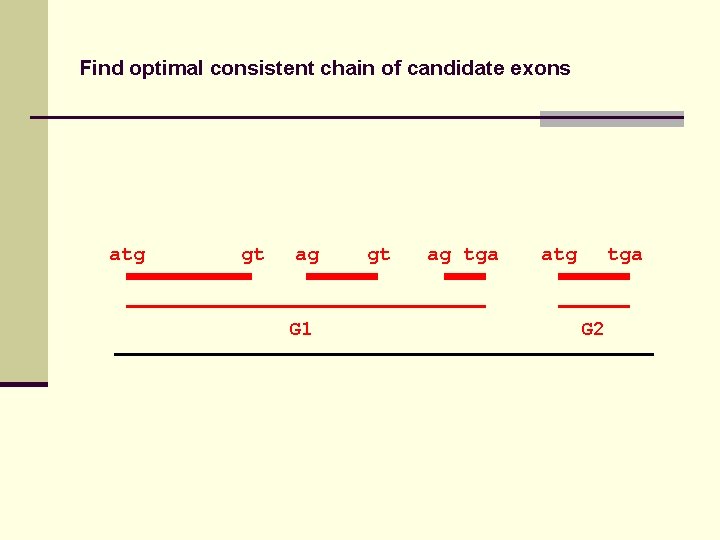
Find optimal consistent chain of candidate exons atg gt ag G 1 gt ag tga atg tga G 2
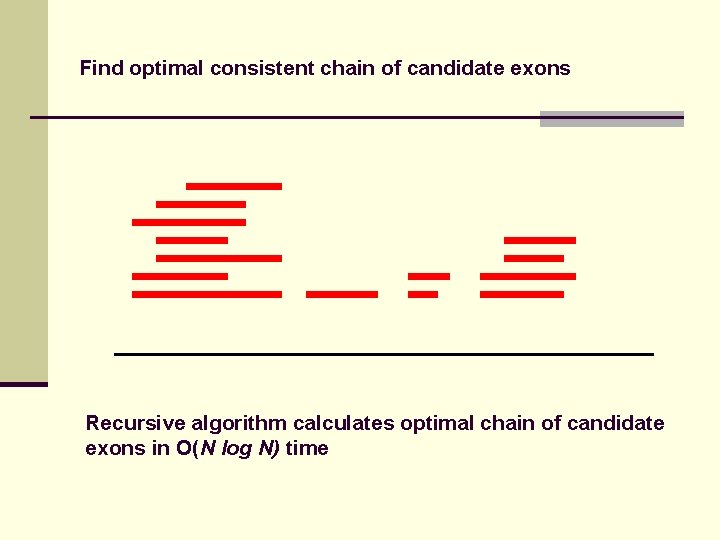
Find optimal consistent chain of candidate exons Recursive algorithm calculates optimal chain of candidate exons in O(N log N) time
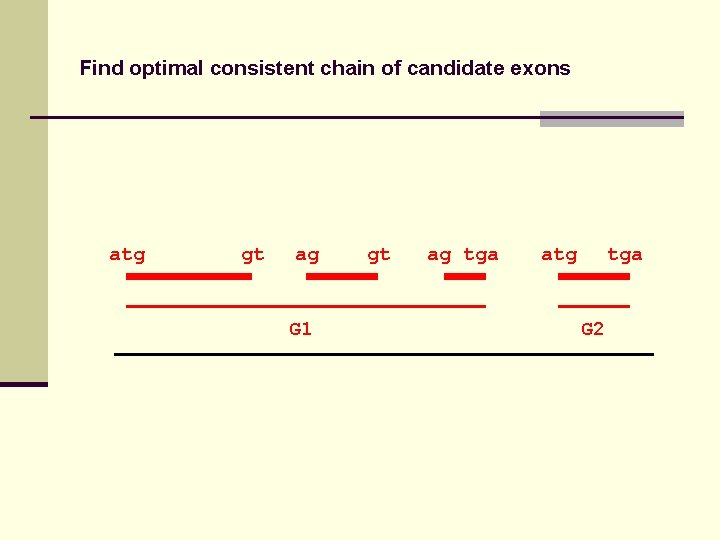
Find optimal consistent chain of candidate exons atg gt ag G 1 gt ag tga atg tga G 2
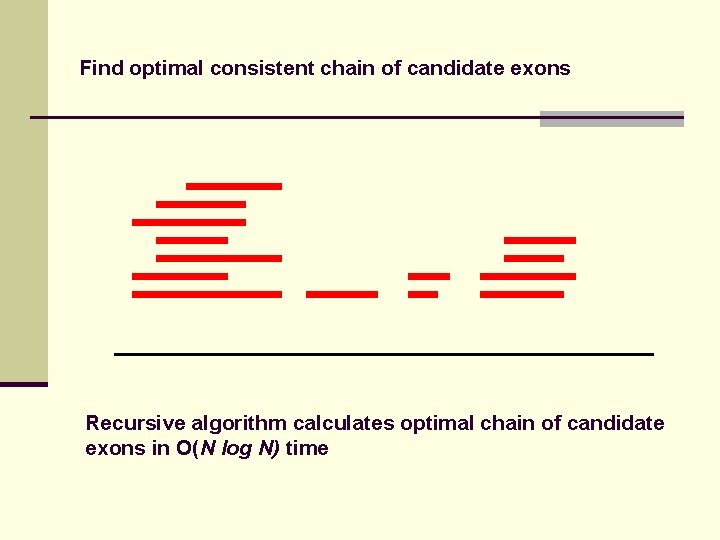
Find optimal consistent chain of candidate exons Recursive algorithm calculates optimal chain of candidate exons in O(N log N) time
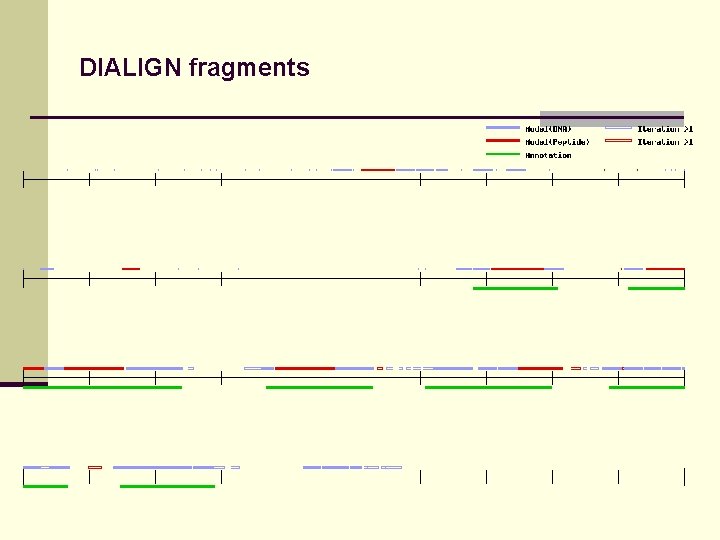
DIALIGN fragments
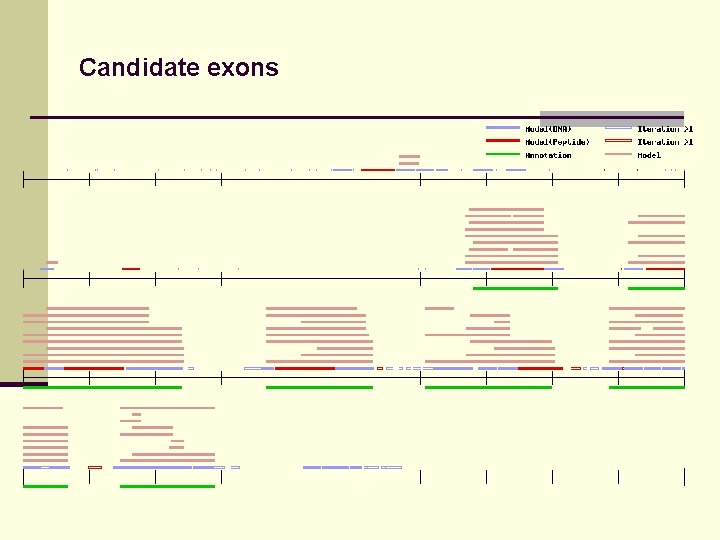
Candidate exons
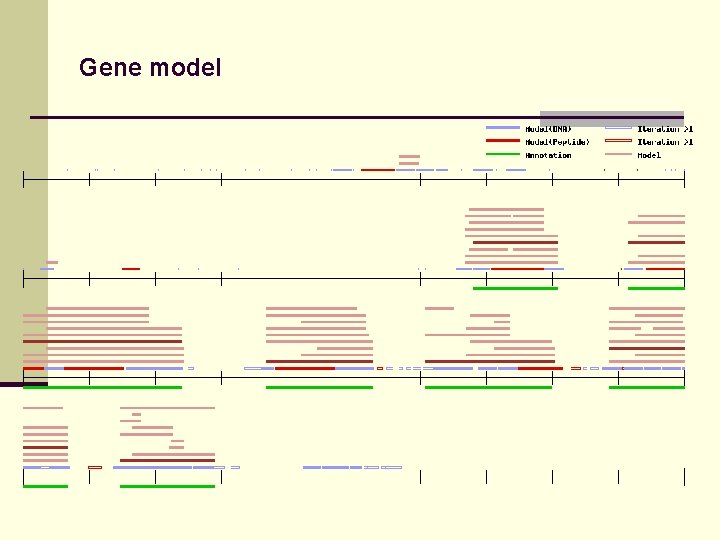
Gene model

Result: 105 pairs of genomic sequences from human and mouse (Batzoglou et al. , 2000)

Result: 105 pairs of genomic sequences from human and mouse (Batzoglou et al. , 2000)

Result: 105 pairs of genomic sequences from human and mouse (Batzoglou et al. , 2000) AGen. DA Gen. Scan 64 % 12 % 17 %

Alignment of large genomic sequences DIALIGN used by tracker for phylogenetic footprinting (Prohaska et al. , 2004)

Alignment of large genomic sequences DIALIGN used by tracker for phylogenetic footprinting (Prohaska et al. , 2004) Alignment of Hox gene cluster:

Alignment of large genomic sequences DIALIGN used by tracker for phylogenetic footprinting (Prohaska et al. , 2004) Alignment of Hox gene cluster: n DIALIGN able to identify small regulatory elements, but

Alignment of large genomic sequences DIALIGN used by tracker for phylogenetic footprinting (Prohaska et al. , 2004) Alignment of Hox gene cluster: n DIALIGN able to identify small regulatory elements, but n Entire genes totally mis-aligned

Alignment of large genomic sequences DIALIGN used by tracker for phylogenetic footprinting (Prohaska et al. , 2004) Alignment of Hox gene cluster: n DIALIGN able to identify small regulatory elements, but n Entire genes totally mis-aligned n Reason for mis-alignment: duplications !
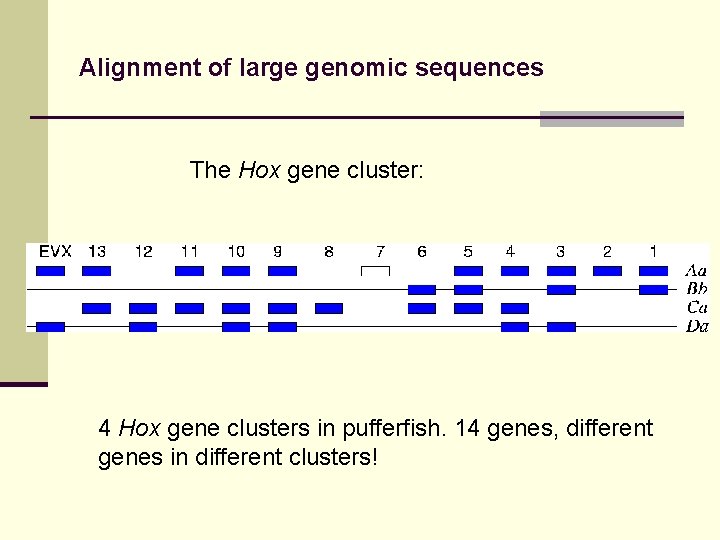
Alignment of large genomic sequences The Hox gene cluster: 4 Hox gene clusters in pufferfish. 14 genes, different genes in different clusters!
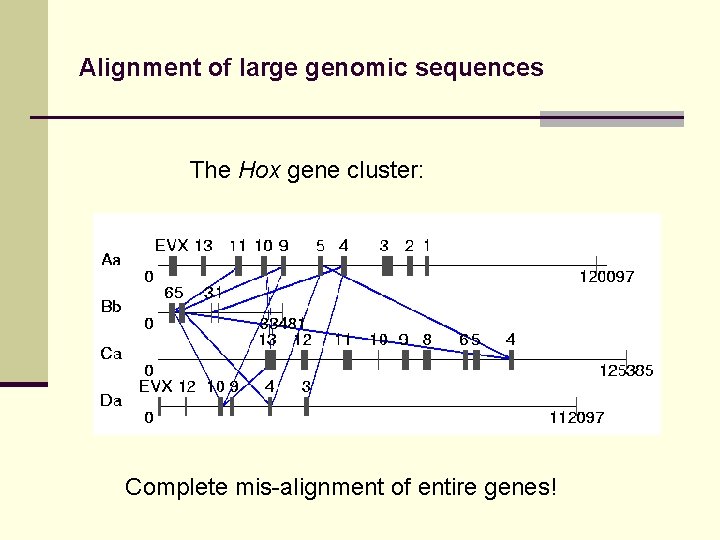
Alignment of large genomic sequences The Hox gene cluster: Complete mis-alignment of entire genes!
Alignment of sequence duplications S 1 S 2
Alignment of sequence duplications S 1 S 2 Conserved motivs; no similarity outside motifs
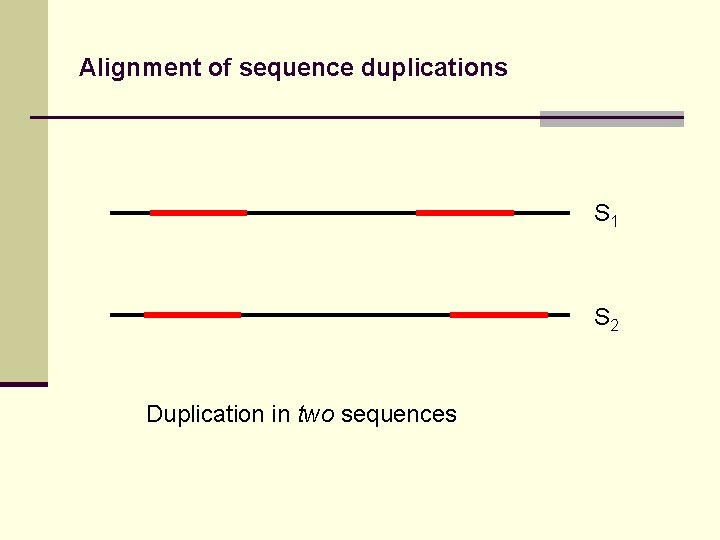
Alignment of sequence duplications S 1 S 2 Duplication in two sequences
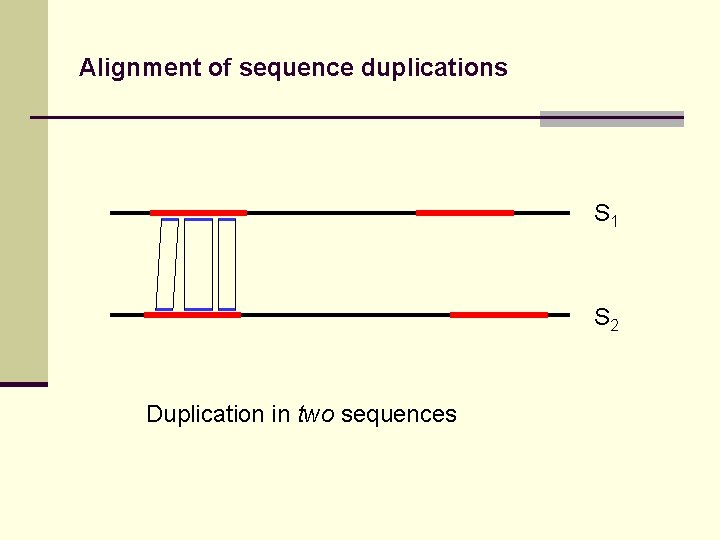
Alignment of sequence duplications S 1 S 2 Duplication in two sequences
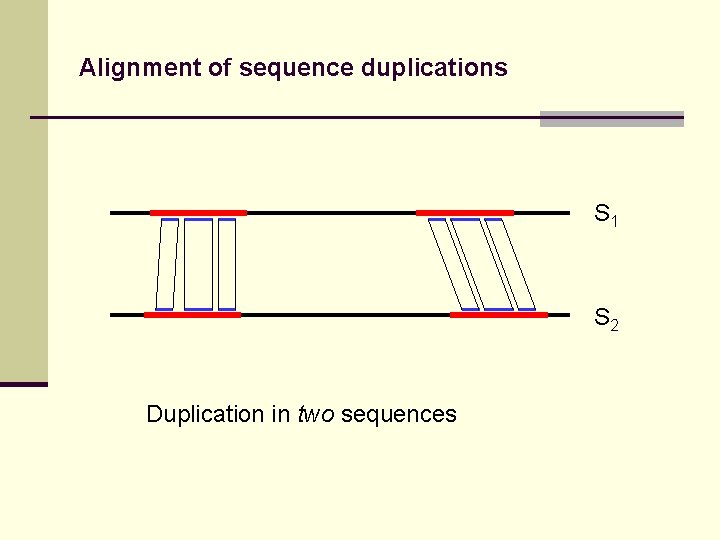
Alignment of sequence duplications S 1 S 2 Duplication in two sequences
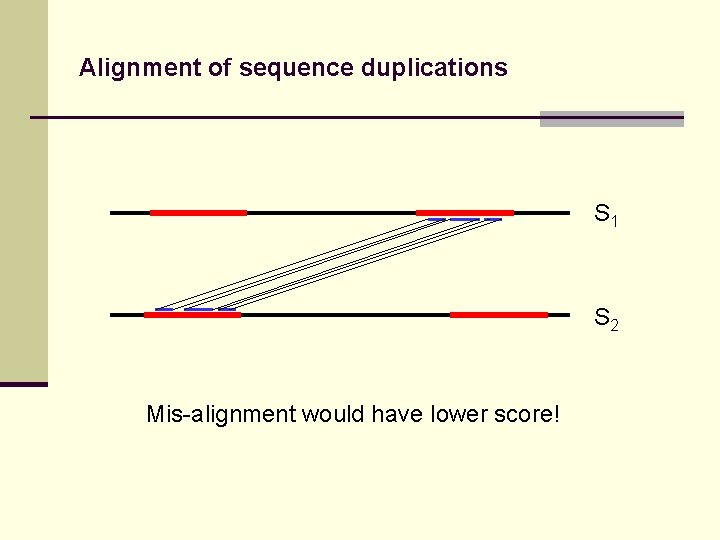
Alignment of sequence duplications S 1 S 2 Mis-alignment would have lower score!
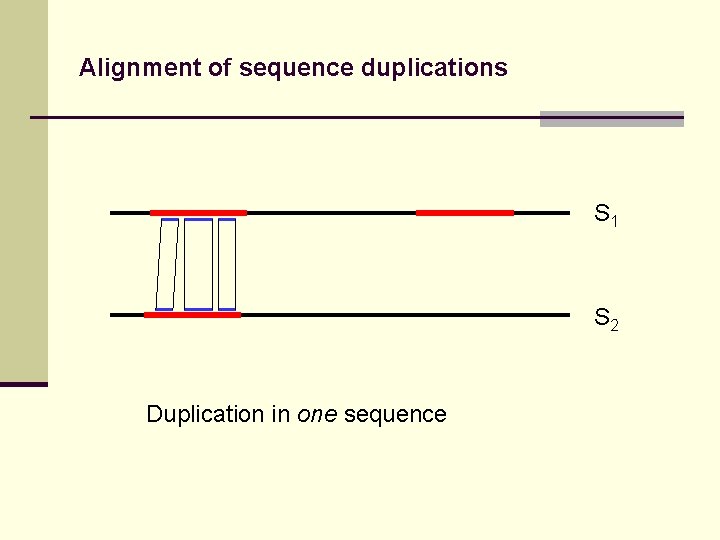
Alignment of sequence duplications S 1 S 2 Duplication in one sequence
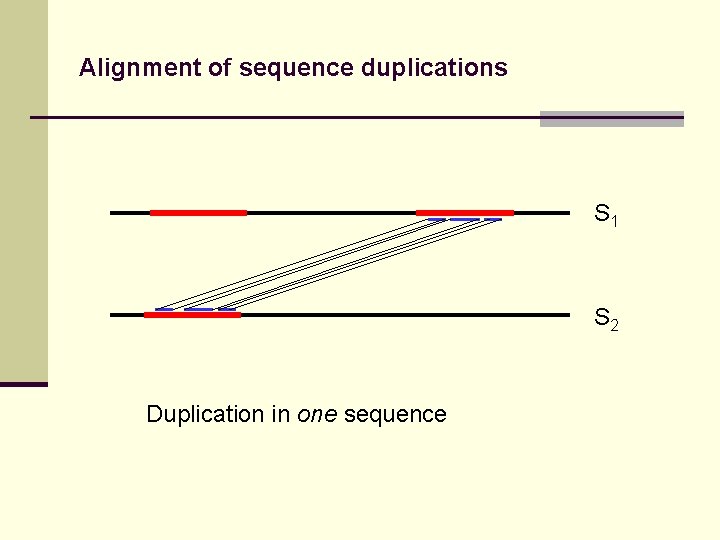
Alignment of sequence duplications S 1 S 2 Duplication in one sequence
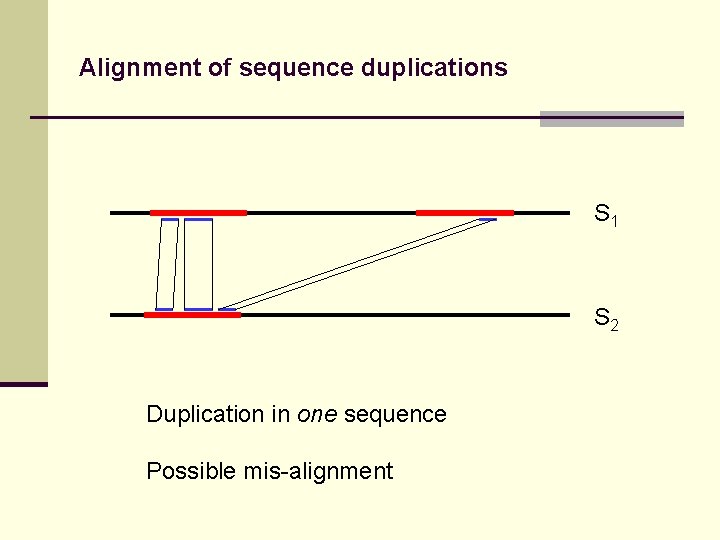
Alignment of sequence duplications S 1 S 2 Duplication in one sequence Possible mis-alignment
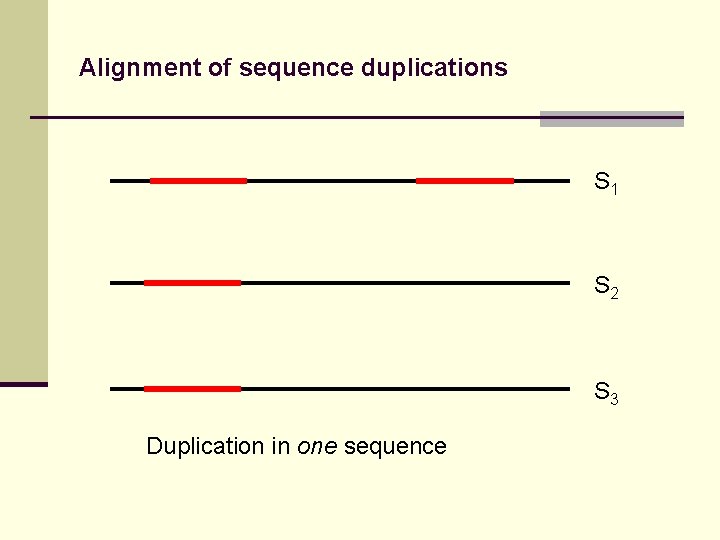
Alignment of sequence duplications S 1 S 2 S 3 Duplication in one sequence
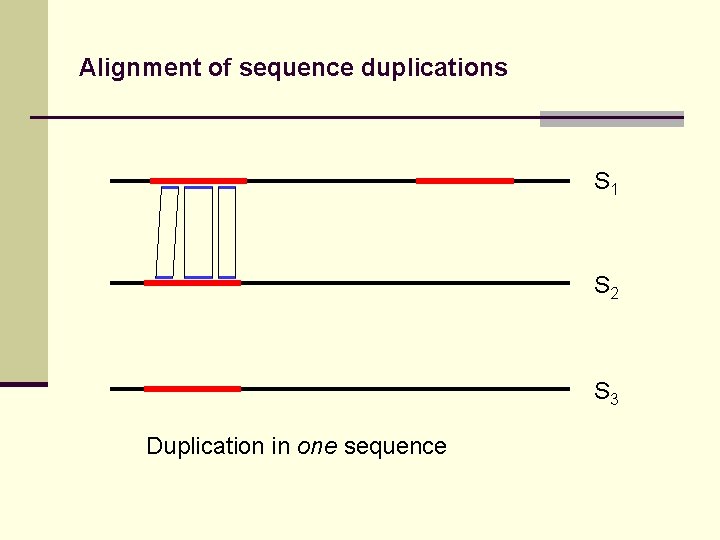
Alignment of sequence duplications S 1 S 2 S 3 Duplication in one sequence

Alignment of sequence duplications S 1 S 2 S 3 Duplication in one sequence
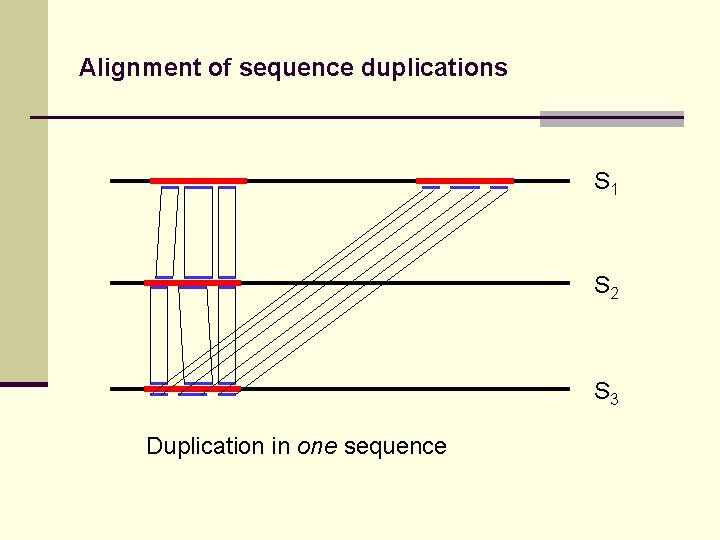
Alignment of sequence duplications S 1 S 2 S 3 Duplication in one sequence
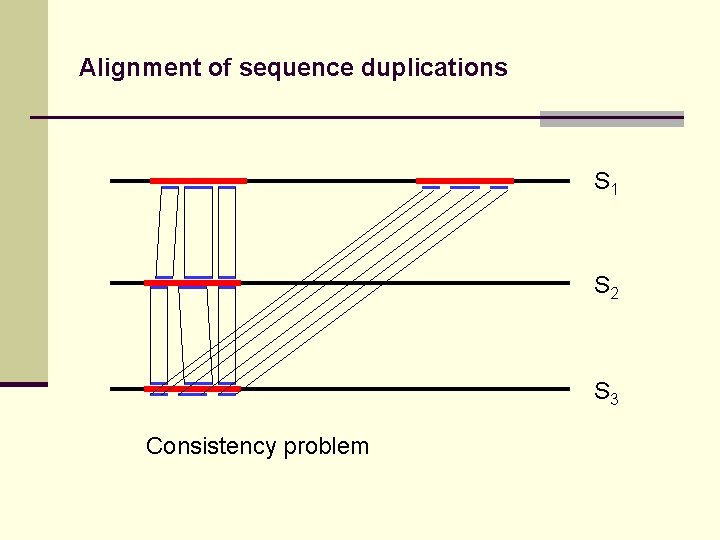
Alignment of sequence duplications S 1 S 2 S 3 Consistency problem
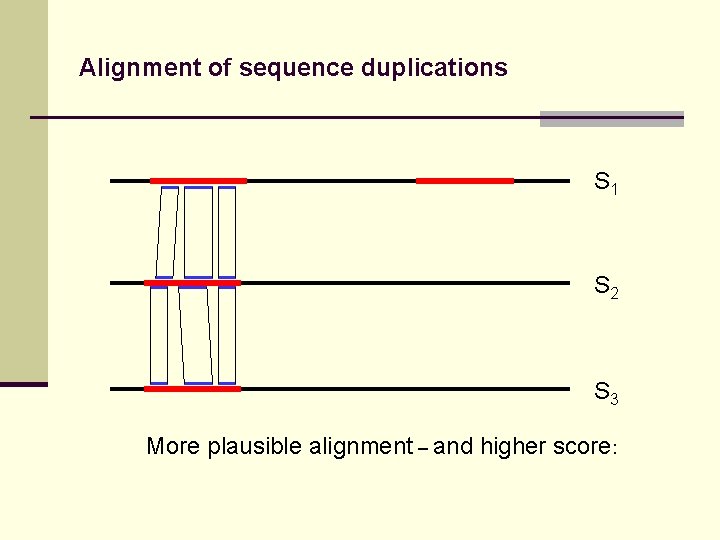
Alignment of sequence duplications S 1 S 2 S 3 More plausible alignment – and higher score:
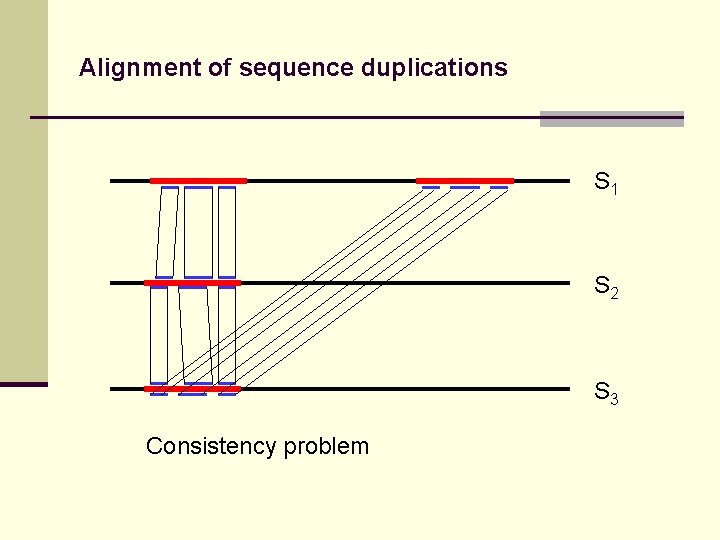
Alignment of sequence duplications S 1 S 2 S 3 Consistency problem
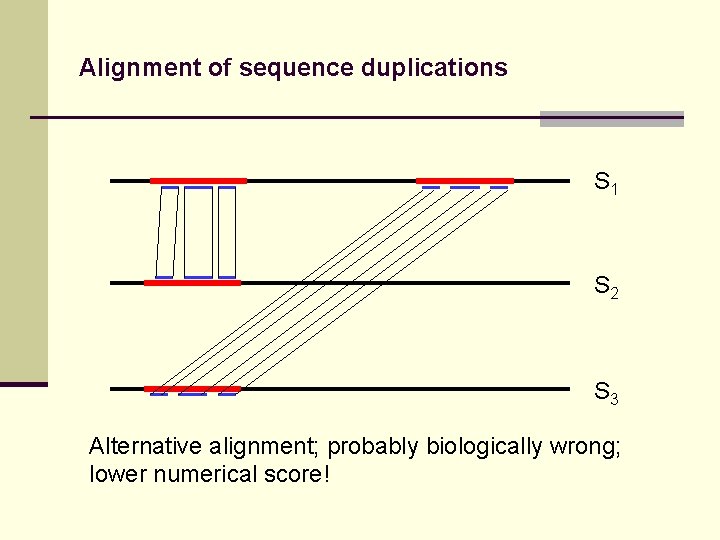
Alignment of sequence duplications S 1 S 2 S 3 Alternative alignment; probably biologically wrong; lower numerical score!

Anchored sequence alignment Biologically meaningful alignment often not possible by automated approaches.

Anchored sequence alignment Biologically meaningful alignment not possible by automated approaches. Idea: use expert knowledge to guide alignment procedure

Anchored sequence alignment Biologically meaningful alignment not possible by automated approaches. Idea: use expert knowledge to guide alignment procedure User defines a set anchor points that are to be „respected“ by the alignment procedure

Anchored sequence alignment NLFVALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT

Anchored sequence alignment NLFVALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT

Anchored sequence alignment NLFVALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT Use known homology as anchor point

Anchored sequence alignment NLFV ALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT Use known homology as anchor point

Anchored sequence alignment NLFV ALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT Use known homology as anchor point Anchor point = anchored fragment (gap-free pair of segments)

Anchored sequence alignment NLFV ALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT Use known homology as anchor point Anchor point = anchored fragment (gap-free pair of segments) Remainder of sequences aligned automatically

Anchored sequence alignment -------NLF VALYDFVASG DNTLSITKGE klrvlgynhn iihredk. GVI YALWDYEPQN DDELPMKEGD cmt------- Anchored alignment

Anchored sequence alignment NLFVALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT GYQYRALYDYKKEREEDIDLHLGDILTVNKGSLVALGFS Anchor points in multiple alignment

Anchored sequence alignment NLFV ALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQND DELPMKEGDCMT GYQYRALYDYKKEREEDIDLHLGDILTVNKGSLVALGFS Anchor points in multiple alignment

Anchored sequence alignment -------NLF V-ALYDFVAS GD---- NTLSITKGEk lrv. LGYNhn iihredk. GVI Y-ALWDYEPQ ND---- DELPMKEGDC MT-------GYQ Yr. ALYDYKKE REedidlhlg DILTVNKGSL VA-LGFS-- Anchored multiple alignment

Algorithmic questions Goal: n Find optimal alignment (=consistent set of fragments) under costraints given by userspecified anchor points!

Algorithmic questions Additional input file with anchor points: 1 2 1 3 3 4 215 34 317 231 78 402 5 23 8 4. 5 1. 23 8. 5

Algorithmic questions NLFVALYDFVASGDNTLSITKGEKLRVLGYNHN IIHREDKGVIYALWDYEPQNDDELPMKEGDCMT GYQYRALYDYKKEREEDIDLHLGDILTVNKGSLVALGFS

Algorithmic questions Additional input file with anchor points: 1 2 1 3 3 4 215 34 317 231 78 402 5 23 8 4. 5 1. 23 8. 5

Algorithmic questions Additional input file with anchor points: 1 2 1 3 3 4 Sequences 215 34 317 231 78 402 5 23 8 4. 5 1. 23 8. 5

Algorithmic questions Additional input file with anchor points: 1 2 1 3 3 4 Sequences 215 34 317 231 78 402 start positions 5 23 8 4. 5 1. 23 8. 5
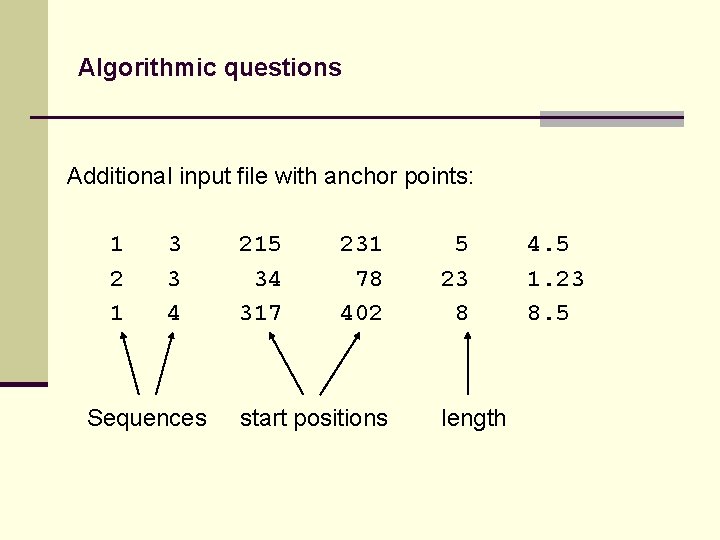
Algorithmic questions Additional input file with anchor points: 1 2 1 3 3 4 Sequences 215 34 317 231 78 402 start positions 5 23 8 length 4. 5 1. 23 8. 5

Algorithmic questions Additional input file with anchor points: 1 2 1 3 3 4 Sequences 215 34 317 231 78 402 start positions 5 23 8 4. 5 1. 23 8. 5 length score

Algorithmic questions Requirements: n Anchor points need to be consistent! – if necessary: select consistent subset from user -specified anchor points

Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa

Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Inconsistent anchor points!

Algorithmic questions atctaat---agttaaactcccccgtgcttag Cagtgcgtgtattac-taacggttcaatcgcg caaagagtatcacccctgaataa Inconsistent anchor points!

Algorithmic questions Requirements: n Anchor points need to be consistent! – if necessary: select consistent subset from user -specified anchor points

Algorithmic questions Requirements: n Anchor points need to be consistent! – if necessary: select consistent subset from user -specified anchor points n Find alignment under constraints given by anchor points!

Algorithmic questions Use data structures from multiple alignment
Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa
Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Greedy procedure for multiple alignment
Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Greedy procedure for multiple alignment
Algorithmic questions atctaatagttaaactcccccgtgcttag cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa Question: which positions are still alignable ?

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x For each position x and each sequence Si exist an upper bound ub(x, i) and a lower bound lb(x, i) for residues y in Si that are alignable with x

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x For each position x and each sequence Si exist an upper bound ub(x, i) and a lower bound lb(x, i) for residues y in Si that are alignable with x

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x ub(x, i) and lb(x, i) updated during greedy procedure

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x Initial values of lb(x, i), ub(x, i)

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x ub(x, i) and lb(x, i) updated during greedy procedure

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x ub(x, i) and lb(x, i) updated during greedy procedure

Algorithmic questions Anchor points treated like fragments in greedy algorithm:
Algorithmic questions Anchor points treated like fragments in greedy algorithm: n Sorted according to user-defined scores

Algorithmic questions Anchor points treated like fragments in greedy algorithm: n n Sorted according to user-defined scores Accepted if consistent with previously accepted anchors

Algorithmic questions Anchor points treated like fragments in greedy algorithm: n n n Sorted according to user-defined scores Accepted if consistent with previously accepted anchors ub(x, i) and lb(x, i) updated during greedy procedure

Algorithmic questions Anchor points treated like fragments in greedy algorithm: n n n Sorted according to user-defined scores Accepted if consistent with previously accepted anchors ub(x, i) and lb(x, i) updated during greedy procedure Resulting values of ub(x, i) and lb(x, i) used as initial values for alignment procedure

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x Initial values of lb(x, i), ub(x, i)

Algorithmic questions atctaatagttaaactcccccgtgcttag Si cagtgcgtgtattactaacggttcaatcgcg caaagagtatcacccctgaataa x Initial values of lb(x, i), ub(x, i) calculated using anchor points

Algorithmic questions Ranking of anchor points to prioritize anchor points, e. g. n anchor points from verified homologies -higher priority n automatically created anchor points (using CHAOS, BLAST, … ) -- lower priority
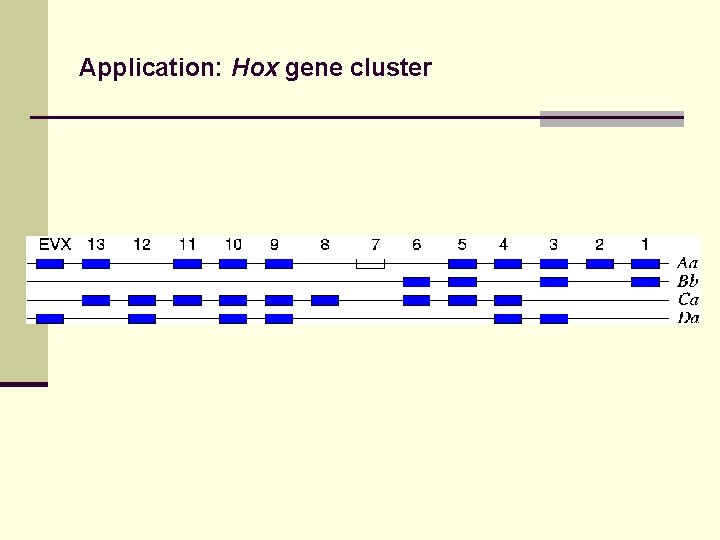
Application: Hox gene cluster
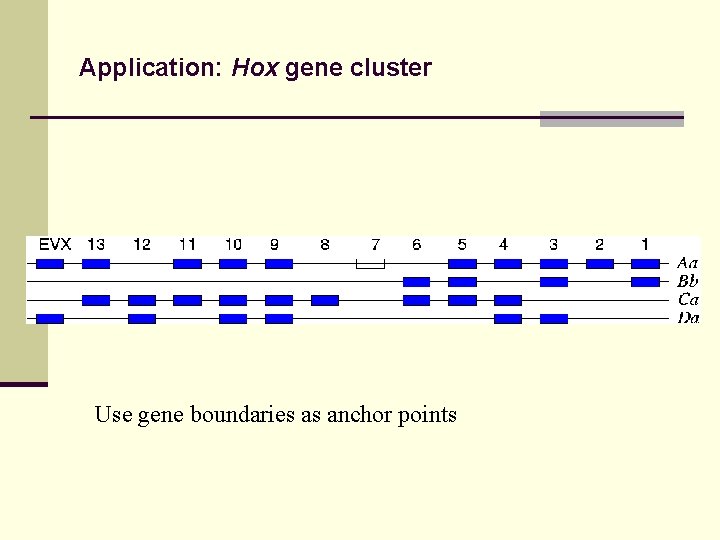
Application: Hox gene cluster Use gene boundaries as anchor points

Application: Hox gene cluster Use gene boundaries as anchor points + CHAOS / BLAST hits
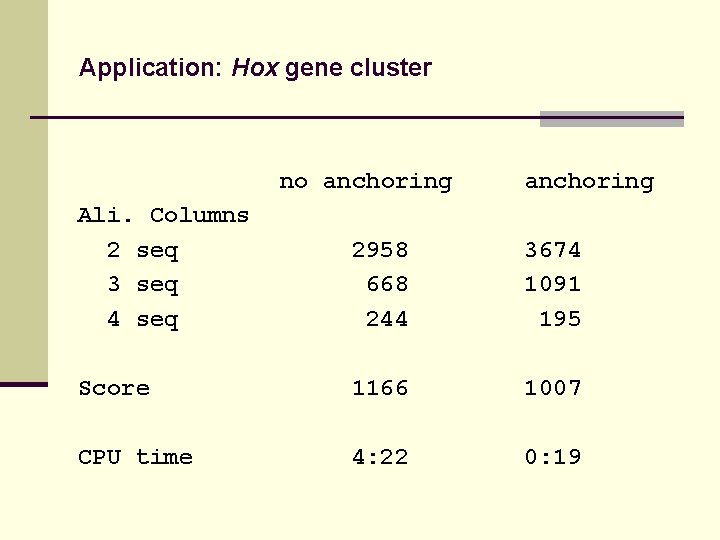
Application: Hox gene cluster no anchoring Ali. Columns 2 seq 3 seq 4 seq 2958 668 244 3674 1091 195 Score 1166 1007 CPU time 4: 22 0: 19

Application: Hox gene cluster Example: Teleost Hox gene cluster:

Application: Hox gene cluster Example: Teleost Hox gene cluster: Score of anchored alignment 15 % higher than score of non-anchored alignment !

Application: Hox gene cluster Example: Teleost Hox gene cluster: Score of anchored alignment 15 % higher than score of non-anchored alignment ! Conclusion: Greedy optimization algorithm does a bad job!

Application: Improvement of Alignment programs Two possible reasons for mis-alignments:

Application: Improvement of Alignment programs Two possible reasons for mis-alignments: n Wrong objective function: Biologically correct alignment gets bad numerical score

Application: Improvement of Alignment programs Two possible reasons for mis-alignments: n Wrong objective function: Biologically correct alignment gets bad numerical score n Bad optimization algorithms: Biologically correct alignment gets best numerical score, but algorithm fails to find this alignment

Application: Improvement of Alignment programs Two possible reasons for mis-alignments: n Anchored alignments can help to decide
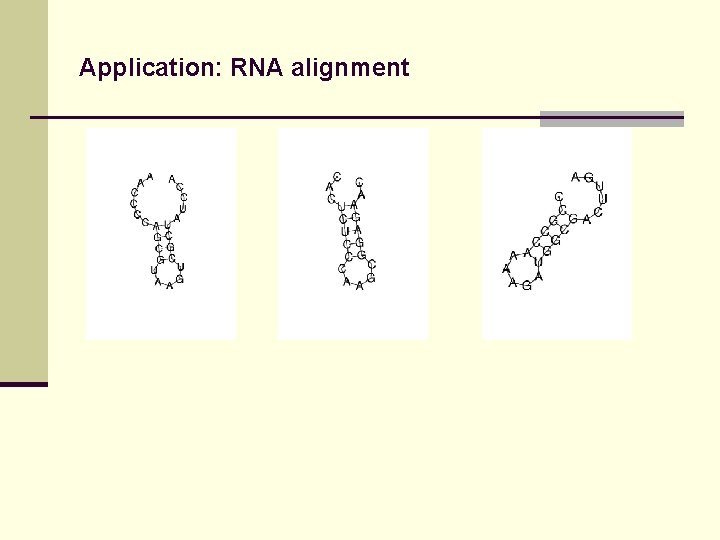
Application: RNA alignment
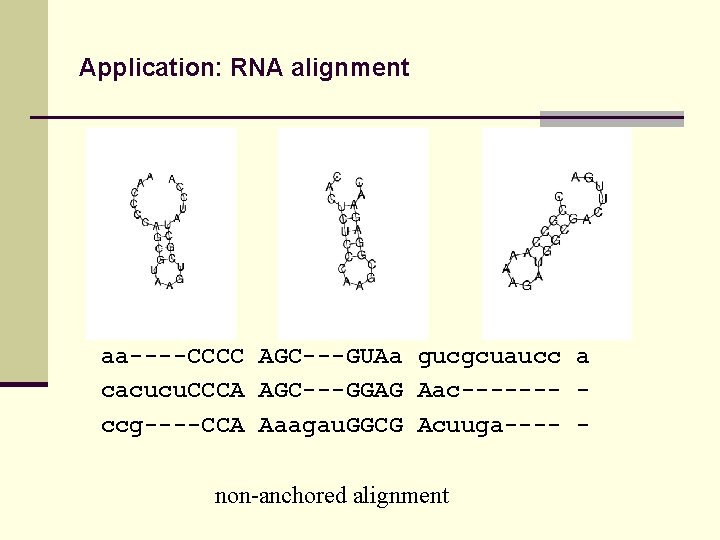
Application: RNA alignment aa----CCCC AGC---GUAa gucgcuaucc a cacucu. CCCA AGC---GGAG Aac------- ccg----CCA Aaagau. GGCG Acuuga---- non-anchored alignment
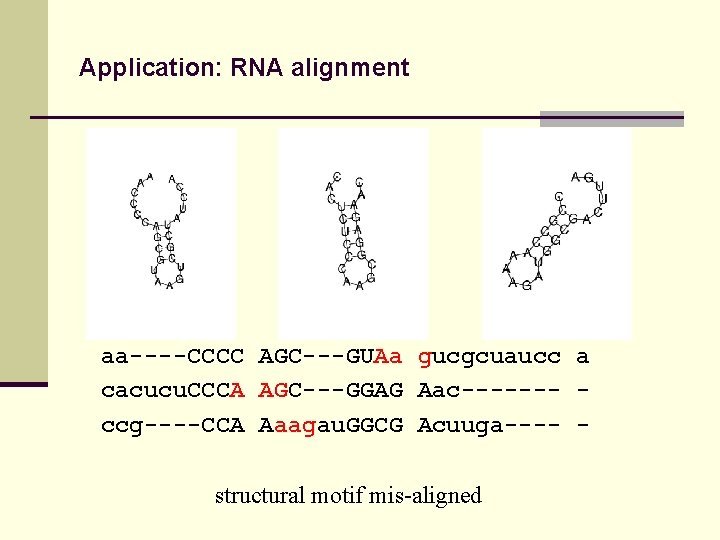
Application: RNA alignment aa----CCCC AGC---GUAa gucgcuaucc a cacucu. CCCA AGC---GGAG Aac------- ccg----CCA Aaagau. GGCG Acuuga---- structural motif mis-aligned
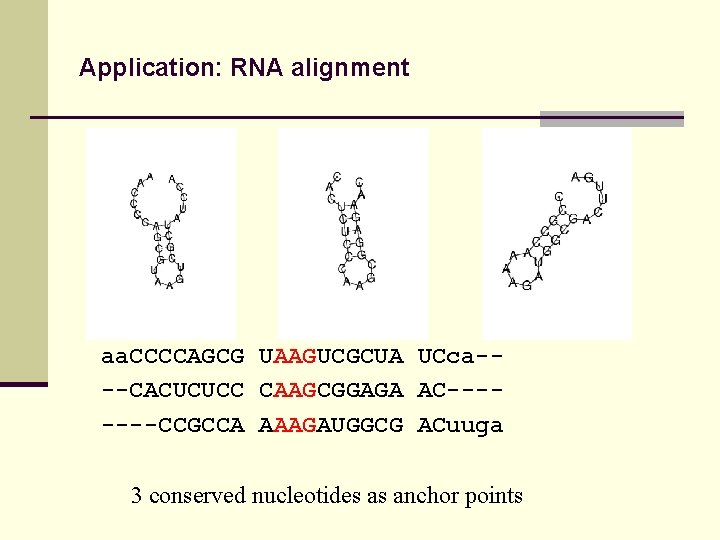
Application: RNA alignment aa. CCCCAGCG UAAGUCGCUA UCca---CACUCUCC CAAGCGGAGA AC-------CCGCCA AAAGAUGGCG ACuuga 3 conserved nucleotides as anchor points

WWW interface at GOBICS (Göttingen Bioinformatics Compute Server)

WWW interface at GOBICS (Göttingen Bioinformatics Compute Server)
- Slides: 158