Algorithms in Bioinformatics A Practical Introduction Project Motif

Algorithms in Bioinformatics: A Practical Introduction Project: Motif finding using Ch. IP-seq peak data
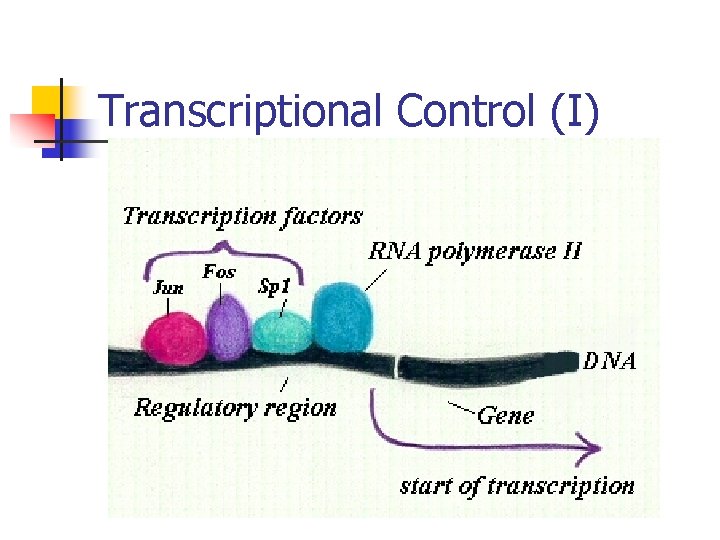
Transcriptional Control (I)
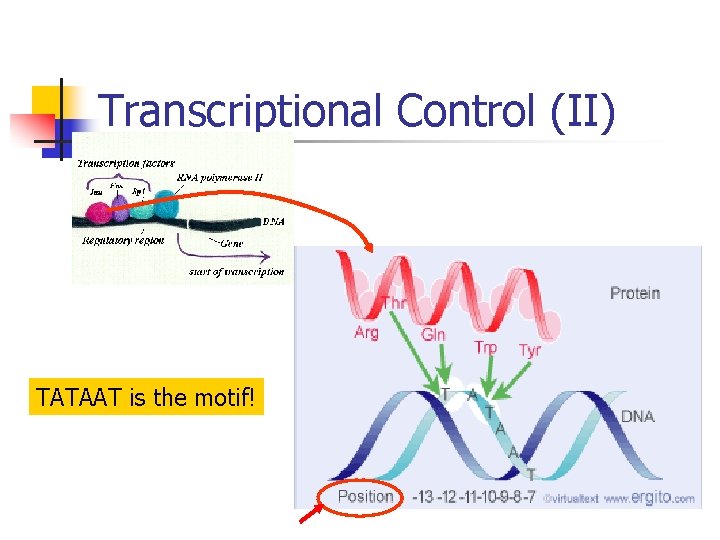
Transcriptional Control (II) TATAAT is the motif!
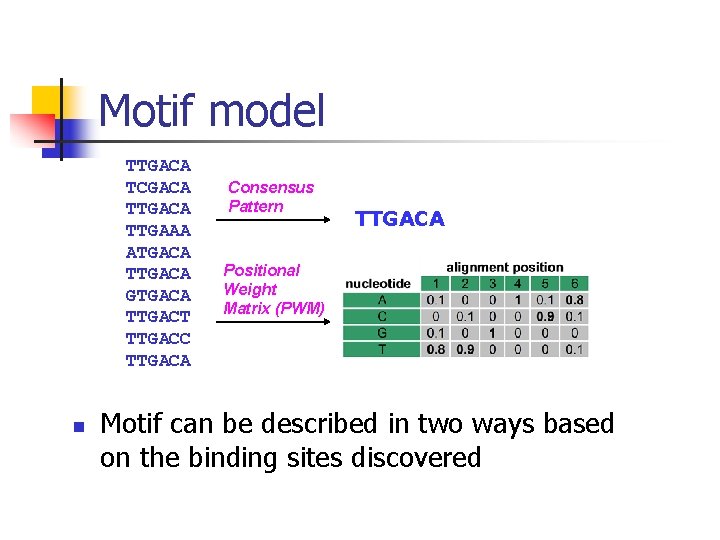
Motif model TTGACA TCGACA TTGAAA ATGACA TTGACA GTGACA TTGACT TTGACC TTGACA n Consensus Pattern TTGACA Positional Weight Matrix (PWM) Motif can be described in two ways based on the binding sites discovered
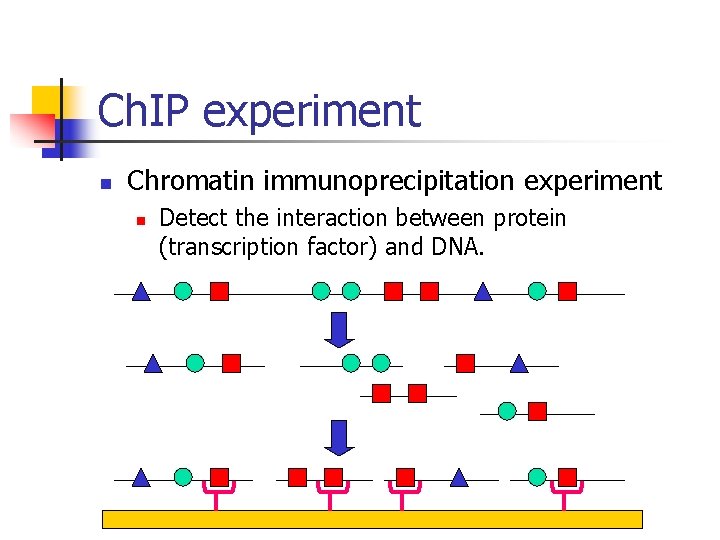
Ch. IP experiment n Chromatin immunoprecipitation experiment n Detect the interaction between protein (transcription factor) and DNA.

Peak data n n Peak data represents the locations where a particular TF binding. The data tells us the locations and intensities. n (Note that due to experimental error, peaks of low intensity may be noise. ) Ch. IP-seq data for Human (MCF 7) E 2 treatment at 45 min chr 1: 883, 686 -958, 485

Our aim n n Given the DNA sequences of those peaks, find motifs which occur in those peak regions. For the example below, we have two motifs: TTGACA and GCATC. n Note that each instance has at most 1 mutation. GCACGCGGTATCGTTAGCTTGACAATGAAGAATCCCCCCGCTCGACAGT GCATACTTTGACACTGACTTCGCTTCTTTAATGAAACATGCG CCCTCTGGAAATTAGTGCGGCATCTCACAACCCGAGGAATGACCAAATG GTATTGAAAGTAAGGCAACGGTGATCCCCATGACACCAAAGATGCTAAG CAACGCTCAGGCAACGTTGACAGGTGACACGTTGACTGCGGCCTCCTGC GTCTCTTGACCGCTTAATCCTAAAGGCCTCCTATTAGTATCCGCAATGT GAACAGGAGCGCGAGCCATCAATTGAAGCGAAGTTGACACCTAATAACT

Input (I) n From every peak, we get approximately +/-200 DNA sequence >cmyc_1_chr 1_4842133_4842148_range_chr 1_4841934_4842348_intensity_20 CCTCCATACCAGCCCCAATGTTCTGCGTTCCCGAATGAAAGACACACAGCCTTTATATTTTGATATGCCTAAAACTG CTCAATGGCTGGGCCACTTCCTAGTATCCACGTGGCTATCCCACCTCTGATATTCCCAAGTCATTACTTA CTAAAATCTGTAATTACATCTTTGCTGCCCTAGGCCCAATCTGGCAGCCCTCCTGTGGCCCCTCAGGCTACTACATG GCAGCTAAGCTCTCTGACCCACATCTTCTCAGGCACCGTGCCTCCTCTTCTCCACCTTATTCAAACATGGTGGCTCTC CTTCCTCCTTCTTCCTGTCCCCAGCCTGGGAATTCTAAAAGTCCCACCTCTGCCCTGTTCAGCCATTGGC TGTCGGCATCTTTACGAG >cmyc_2_chr 1_5073201_5073215_range_chr 1_5073002_5073415_intensity_15 GGTCATAAACCAAGCTTCTTCAAAGATTTTTGGCACCAGTGGCCTGCAGGGTGGCGAGCTCTGCCAGTTTGAA GTGACCAAGTTAAGTGGCCTGGGAAAGGCCATTTGGTGCGCGGTCCAGCAGTTTTGGGCGCTCTCGGCTTCCGCCC TCAGCTGCGGTCACGTGCGGCTGCTCACGTGCCAGACGCTGCTGTCACTTCGTAGCTGTTCCGGCTTCCTCTGAGTG AGGCTCGCAACGTCTCCCACGGAGTCGCCTTCGTTCTGCTCTGGGTCTCCCGTGGCCACTGAGACCTCGGAGCTCGA CCGGCGCCTGCCCGTGCGGCCCTCACTCCCCGAGGCTATCCAGGTGAGGCCGCCTGGGGTCCCCGGCT CCGGAGAGCCGACTGGTTTCCCTGCCG >cmyc_3_chr 1_9530642_9530652_range_chr 1_9530443_9530852_intensity_36 GTAGTCCCAACCAGGTCCTGAGCTGGTTAGCCAACCCTCAGCGCCAGTCGGGCCAACATCCGGTGACGAATCCAAGTCCC GCCTCTAAGCCCATCTGCTGTCCAATGCCGCCCTCTGCCGGTCTTTACCTCCCCGCCTAGCTGTGAGCCGCTTCCAG ACAACCCGGAAGTGATCTTTCCTCTTCCGGATTACGGGTCCGGACGTCCGCACGTGGTTGCCGGTTTAGGGTGCTG CTGTAGTGGCGATACGTCCCGCCGCTGTCCCGAAGTGAGGGATCCGAGCCGCAGCGAGAGCCATGGAGGGCCAGCG CGTGGAGGAGCTGCTGGCCAAGGCAGAGCAGGAGGAGGCGGAGAAGCTGCAGCGCATCACGGTGCACAAGGAGCTGGAGTTCGACCTGGGCAACC ……………

Input (II) n A set of sequences which are likely containing no motif. >SEQ_1 AACAAGGGAAAGAGTAGTGCTTCTATTCAGAGGGGAAGTTGCTGTTAGCTAAGACAGTCAGGACTGAG AAGGGGGGTTTAACTCTCCTGGAGCTGAGAGGTAAAGGGGCGTGAGGTAGAACAAGCCGAG AACACAGGGCAGGTTGGTCTGACTCCAGAGCACAGTGCAGGAGCCCGGAAGTTGACTCAGTTAGCAAGTAT TTTCACACAAGGCGTGAACACTGAAGACAAAAGCAAGAGACACAGCTCTATCTCTAAGAAGATTTTCAGAGCCAAGA TCGATGGGGCACACCTGTTAATCCCAGCACTTAGGAGGCTGAGGCAGGAGGATCCCAAGTTCAAAACCAGCCTGGA CTTGTTTTAAGGAAAA >SEQ_2 AAAAAAAGACTTCCAGTTTAATAAATGACCAATTCAGGAATGGAGATTAGGGCTGGATGACAAGTTTTTAATTG TCAAGGACTCAATTCTGTTTATCAGTTGGTATGGAATTATGTAAGCTTTTAGCGATATGACCGCACGGAGCAGTGTA GAGAGTGATCTGAGAGACGCTTGGGGGTCAGGATGGAGATAGAACTCCCTCTCTATTAGAAGGTGTTTGGTGGTAG GTAACCCTGGGCTAGCATGGTGGGTCTCTTCTTAGGCTTCCATCTTTGTGGTTCAAATCCAAGAAGGACCTGC GTTCCCTCCTTGTGATCAGCTGATTGCTAGAGCATAACTCATCTTAACTTCTCATGTACTCTCCGGGTACAGGA AGGGGGC >SEQ_3 CCACTGCTGACAGTGGAGCATGAAACGACCGGCTTCCTGACTATGTTGGTACCCTTTCAGGAGCCTAAAACAGTGCTTTCA ATACTTGTGTCTATGTCTGTTAGCCACAACTTTCTAGTTTCCCAGAGAGATTTTGAAGTGTAGTTTTGTATTTGCTCA AATATTCATATGGTGAGGTGCACATTTTTTATATTTTTATTCATTTTTGGTGCTTGGGAATTATA CTCTAGGAATAAAGCGCCTGGTAGAAAGTGGCACACATCTTTAATCCCAGCACTCAGGAAGCAGAGGCAGACAAATC TCTGCGTTCCAGGACAGCCTGGTCTATAGAGCAAGGTCCAAGCCAGGTTTACACAAAGAAACCTAGTGTGGAA AAGACAAAA ……………

Output n n You need to output a list of candidate (ranked) motifs. You can model the motif as PWM or consensus sequence. If you model the motif as a PWM, one of the answer for the previous dataset is You may also return other significant motifs.

Aim of the project n Given a sample file and a background file, n n you need to implement a method which output a list of motifs. You need to take advantage of the fact that this is a Ch. IP-seq dataset n Hint: Read papers on Ch. IP-seq and understand its properties.
- Slides: 11